User login
Study reveals potential target for T-ALL

Photo by Rhoda Baer
Targeting CXCL12/CXCR4 signaling could be a “powerful” approach to treating T-cell acute lymphoblastic leukemia (T-ALL), according to researchers.
Results of their preclinical experiments indicate that deleting CXCL12 from vascular endothelial cells can stall T-ALL progression.
And inhibiting or deleting CXCR4 can halt tumor growth and induce remission in mice with T-ALL.
The researchers reported these findings in Cancer Cell.
“Our experiments showed that blocking CXCR4 decimated leukemia cells,” said study author Susan Schwab, PhD, of the New York University School of Medicine in New York, New York.
This suggests CXCR4 antagonists could treat T-ALL, she added, noting that such drugs are already in preliminary testing for myeloid leukemia and have proven well-tolerated thus far.
Dr Schwab and her colleagues conducted this research to build upon previous work, which showed that leukemia-initiating cells concentrate in the bone marrow near CXCL12-producing blood vessels.
This finding inspired the researchers to investigate the expression and function of CXCR4 because it binds to CXCL12, as well as the role CXCR4-CXCL12 molecular signaling plays in disease growth.
With this study, the team found that T-ALL cells are in direct, stable contact with CXCL12-producing bone marrow stroma.
They showed that deleting CXCL12 in vascular endothelial cells suppressed T-ALL. But they did not observe the same benefit when they deleted CXCL12 from perivascular cells.
The researchers also found that T-ALL cells express high surface levels of CXCR4, and deleting CXCR4 can produce sustained remission in mice with T-ALL.
Similarly, a small-molecule inhibitor of CXCR4, AMD3465, exhibited antileukemic activity in murine T-ALL and human xenografts.
Experiments with CXCR4-deficient T-ALL cells showed that CXCR4 expression and signaling influences leukemic cell localization and survival. And further investigation revealed that the loss of CXCR4 expression is associated with decreased levels of MYC.
The researchers said additional work is needed to determine exactly how CXCR4 is able to promote and sustain T-ALL. ![]()

Photo by Rhoda Baer
Targeting CXCL12/CXCR4 signaling could be a “powerful” approach to treating T-cell acute lymphoblastic leukemia (T-ALL), according to researchers.
Results of their preclinical experiments indicate that deleting CXCL12 from vascular endothelial cells can stall T-ALL progression.
And inhibiting or deleting CXCR4 can halt tumor growth and induce remission in mice with T-ALL.
The researchers reported these findings in Cancer Cell.
“Our experiments showed that blocking CXCR4 decimated leukemia cells,” said study author Susan Schwab, PhD, of the New York University School of Medicine in New York, New York.
This suggests CXCR4 antagonists could treat T-ALL, she added, noting that such drugs are already in preliminary testing for myeloid leukemia and have proven well-tolerated thus far.
Dr Schwab and her colleagues conducted this research to build upon previous work, which showed that leukemia-initiating cells concentrate in the bone marrow near CXCL12-producing blood vessels.
This finding inspired the researchers to investigate the expression and function of CXCR4 because it binds to CXCL12, as well as the role CXCR4-CXCL12 molecular signaling plays in disease growth.
With this study, the team found that T-ALL cells are in direct, stable contact with CXCL12-producing bone marrow stroma.
They showed that deleting CXCL12 in vascular endothelial cells suppressed T-ALL. But they did not observe the same benefit when they deleted CXCL12 from perivascular cells.
The researchers also found that T-ALL cells express high surface levels of CXCR4, and deleting CXCR4 can produce sustained remission in mice with T-ALL.
Similarly, a small-molecule inhibitor of CXCR4, AMD3465, exhibited antileukemic activity in murine T-ALL and human xenografts.
Experiments with CXCR4-deficient T-ALL cells showed that CXCR4 expression and signaling influences leukemic cell localization and survival. And further investigation revealed that the loss of CXCR4 expression is associated with decreased levels of MYC.
The researchers said additional work is needed to determine exactly how CXCR4 is able to promote and sustain T-ALL. ![]()

Photo by Rhoda Baer
Targeting CXCL12/CXCR4 signaling could be a “powerful” approach to treating T-cell acute lymphoblastic leukemia (T-ALL), according to researchers.
Results of their preclinical experiments indicate that deleting CXCL12 from vascular endothelial cells can stall T-ALL progression.
And inhibiting or deleting CXCR4 can halt tumor growth and induce remission in mice with T-ALL.
The researchers reported these findings in Cancer Cell.
“Our experiments showed that blocking CXCR4 decimated leukemia cells,” said study author Susan Schwab, PhD, of the New York University School of Medicine in New York, New York.
This suggests CXCR4 antagonists could treat T-ALL, she added, noting that such drugs are already in preliminary testing for myeloid leukemia and have proven well-tolerated thus far.
Dr Schwab and her colleagues conducted this research to build upon previous work, which showed that leukemia-initiating cells concentrate in the bone marrow near CXCL12-producing blood vessels.
This finding inspired the researchers to investigate the expression and function of CXCR4 because it binds to CXCL12, as well as the role CXCR4-CXCL12 molecular signaling plays in disease growth.
With this study, the team found that T-ALL cells are in direct, stable contact with CXCL12-producing bone marrow stroma.
They showed that deleting CXCL12 in vascular endothelial cells suppressed T-ALL. But they did not observe the same benefit when they deleted CXCL12 from perivascular cells.
The researchers also found that T-ALL cells express high surface levels of CXCR4, and deleting CXCR4 can produce sustained remission in mice with T-ALL.
Similarly, a small-molecule inhibitor of CXCR4, AMD3465, exhibited antileukemic activity in murine T-ALL and human xenografts.
Experiments with CXCR4-deficient T-ALL cells showed that CXCR4 expression and signaling influences leukemic cell localization and survival. And further investigation revealed that the loss of CXCR4 expression is associated with decreased levels of MYC.
The researchers said additional work is needed to determine exactly how CXCR4 is able to promote and sustain T-ALL. ![]()
Genome mapping provides insight into MM

Zhou, PhD, (left) and David
Schwartz, PhD, in the lab
Photo by Jeff Miller/University
of Wisconsin-Madison
A novel approach to genome analysis can provide a clearer picture of cancer genomes, according to research published in PNAS.
Investigators used this approach, which combines optical mapping and DNA sequencing, to analyze samples from a patient with multiple myeloma (MM).
They found “widespread structural variation” in the tumor genome and observed an increase in mutational burden that correlated with disease
progression.
“Cancer genomes are complicated, but we found that, using an approach like this, you can begin to understand them at every level,” said study author David Schwartz, PhD, of the University of Wisconsin-Madison.
“The approach allows an intimate view of a cancer genome. You get to see it, you get to measure it, and you get to see it evolve at many levels. This is what we should be doing with every cancer genome, and the goal here is to make the system fast enough so this becomes a routine tool.”
To test the approach, the investigators obtained cancerous and noncancerous samples from a patient with MM at two different time points: while the patient was still responding to treatment and after the patient’s disease had progressed and become resistant to chemotherapy.
The team performed standard DNA sequencing to read each of the letters spelling out the genomic code of the disease. Then, isolating the individual DNA molecules, they performed optical mapping.
This involves stretching single strands of DNA and placing them in a device. The strands are given specific landmarks and marked with a fluorescent dye. An automated system takes images of each of these marked segments, cataloging the molecules into large datasets that are then pieced together to provide a larger view of the genome.
The investigators said the information provided by DNA sequencing and the “bigger picture” provided by optical mapping allowed for a comprehensive view of the patient’s MM genome.
“It’s a rare, near-complete characterization of the complexity of a myeloma genome, from the smallest variance all the way to big chunks of chromosomal material that differ between the tumor DNA and the normal DNA of the patient,” said study author Fotis Asimakopoulos, MD, PhD, of the University of Wisconsin Carbone Cancer Center.
The approach allowed the investigators to see that, compared to the patient’s noncancerous genome, and across the two time points, the MM genome was marked by an increase in notable mutations and larger-scale changes.
The team highlighted the changes they believe are most worthy of further exploration and that yield the greatest potential for future therapies. They hope this approach could help prevent drug resistance or at least help scientists and physicians develop ways to work around it.
It could allow them to examine changes in a patient’s cancer over the progression of the illness, monitor for signs of resistance, and fine-tune treatments, Dr Schwartz said.
“To cure myeloma, we need to understand how genomes evolve with progression and treatment,” Dr Asimakopoulos added. “The more we can understand the drivers in cancer in significant depth, and in each individual, the better we can tailor treatment to each patient’s disease biology.”
“Instead of [calling the disease] grade 1, we can say, ‘This is Joe’s myeloma, and, given this list of mutations and other info, this is the treatment.’ No two myeloma are alike.”
The investigators are now working toward advancing the system, making it higher-resolution, more cost-effective, and scalable. Ultimately, they would like to build a system capable of analyzing 1000 genomes in 24 hours. ![]()

Zhou, PhD, (left) and David
Schwartz, PhD, in the lab
Photo by Jeff Miller/University
of Wisconsin-Madison
A novel approach to genome analysis can provide a clearer picture of cancer genomes, according to research published in PNAS.
Investigators used this approach, which combines optical mapping and DNA sequencing, to analyze samples from a patient with multiple myeloma (MM).
They found “widespread structural variation” in the tumor genome and observed an increase in mutational burden that correlated with disease
progression.
“Cancer genomes are complicated, but we found that, using an approach like this, you can begin to understand them at every level,” said study author David Schwartz, PhD, of the University of Wisconsin-Madison.
“The approach allows an intimate view of a cancer genome. You get to see it, you get to measure it, and you get to see it evolve at many levels. This is what we should be doing with every cancer genome, and the goal here is to make the system fast enough so this becomes a routine tool.”
To test the approach, the investigators obtained cancerous and noncancerous samples from a patient with MM at two different time points: while the patient was still responding to treatment and after the patient’s disease had progressed and become resistant to chemotherapy.
The team performed standard DNA sequencing to read each of the letters spelling out the genomic code of the disease. Then, isolating the individual DNA molecules, they performed optical mapping.
This involves stretching single strands of DNA and placing them in a device. The strands are given specific landmarks and marked with a fluorescent dye. An automated system takes images of each of these marked segments, cataloging the molecules into large datasets that are then pieced together to provide a larger view of the genome.
The investigators said the information provided by DNA sequencing and the “bigger picture” provided by optical mapping allowed for a comprehensive view of the patient’s MM genome.
“It’s a rare, near-complete characterization of the complexity of a myeloma genome, from the smallest variance all the way to big chunks of chromosomal material that differ between the tumor DNA and the normal DNA of the patient,” said study author Fotis Asimakopoulos, MD, PhD, of the University of Wisconsin Carbone Cancer Center.
The approach allowed the investigators to see that, compared to the patient’s noncancerous genome, and across the two time points, the MM genome was marked by an increase in notable mutations and larger-scale changes.
The team highlighted the changes they believe are most worthy of further exploration and that yield the greatest potential for future therapies. They hope this approach could help prevent drug resistance or at least help scientists and physicians develop ways to work around it.
It could allow them to examine changes in a patient’s cancer over the progression of the illness, monitor for signs of resistance, and fine-tune treatments, Dr Schwartz said.
“To cure myeloma, we need to understand how genomes evolve with progression and treatment,” Dr Asimakopoulos added. “The more we can understand the drivers in cancer in significant depth, and in each individual, the better we can tailor treatment to each patient’s disease biology.”
“Instead of [calling the disease] grade 1, we can say, ‘This is Joe’s myeloma, and, given this list of mutations and other info, this is the treatment.’ No two myeloma are alike.”
The investigators are now working toward advancing the system, making it higher-resolution, more cost-effective, and scalable. Ultimately, they would like to build a system capable of analyzing 1000 genomes in 24 hours. ![]()

Zhou, PhD, (left) and David
Schwartz, PhD, in the lab
Photo by Jeff Miller/University
of Wisconsin-Madison
A novel approach to genome analysis can provide a clearer picture of cancer genomes, according to research published in PNAS.
Investigators used this approach, which combines optical mapping and DNA sequencing, to analyze samples from a patient with multiple myeloma (MM).
They found “widespread structural variation” in the tumor genome and observed an increase in mutational burden that correlated with disease
progression.
“Cancer genomes are complicated, but we found that, using an approach like this, you can begin to understand them at every level,” said study author David Schwartz, PhD, of the University of Wisconsin-Madison.
“The approach allows an intimate view of a cancer genome. You get to see it, you get to measure it, and you get to see it evolve at many levels. This is what we should be doing with every cancer genome, and the goal here is to make the system fast enough so this becomes a routine tool.”
To test the approach, the investigators obtained cancerous and noncancerous samples from a patient with MM at two different time points: while the patient was still responding to treatment and after the patient’s disease had progressed and become resistant to chemotherapy.
The team performed standard DNA sequencing to read each of the letters spelling out the genomic code of the disease. Then, isolating the individual DNA molecules, they performed optical mapping.
This involves stretching single strands of DNA and placing them in a device. The strands are given specific landmarks and marked with a fluorescent dye. An automated system takes images of each of these marked segments, cataloging the molecules into large datasets that are then pieced together to provide a larger view of the genome.
The investigators said the information provided by DNA sequencing and the “bigger picture” provided by optical mapping allowed for a comprehensive view of the patient’s MM genome.
“It’s a rare, near-complete characterization of the complexity of a myeloma genome, from the smallest variance all the way to big chunks of chromosomal material that differ between the tumor DNA and the normal DNA of the patient,” said study author Fotis Asimakopoulos, MD, PhD, of the University of Wisconsin Carbone Cancer Center.
The approach allowed the investigators to see that, compared to the patient’s noncancerous genome, and across the two time points, the MM genome was marked by an increase in notable mutations and larger-scale changes.
The team highlighted the changes they believe are most worthy of further exploration and that yield the greatest potential for future therapies. They hope this approach could help prevent drug resistance or at least help scientists and physicians develop ways to work around it.
It could allow them to examine changes in a patient’s cancer over the progression of the illness, monitor for signs of resistance, and fine-tune treatments, Dr Schwartz said.
“To cure myeloma, we need to understand how genomes evolve with progression and treatment,” Dr Asimakopoulos added. “The more we can understand the drivers in cancer in significant depth, and in each individual, the better we can tailor treatment to each patient’s disease biology.”
“Instead of [calling the disease] grade 1, we can say, ‘This is Joe’s myeloma, and, given this list of mutations and other info, this is the treatment.’ No two myeloma are alike.”
The investigators are now working toward advancing the system, making it higher-resolution, more cost-effective, and scalable. Ultimately, they would like to build a system capable of analyzing 1000 genomes in 24 hours. ![]()
PET tracer allows for whole-body thrombus detection
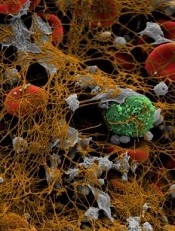
Image by Andre E.X. Brown
BALTIMORE—Scientists say they have synthesized an imaging agent that allows them to detect blood clots throughout the body and estimate the fibrin content of those clots.
In murine experiments, the fibrin-specific PET probe FBP8 detected arterial and venous thrombi with high accuracy.
The tracer also proved sensitive to changes in the fibrin content of clots, which correlated with their age.
Francesco Blasi, PhD, of Massachusetts General Hospital in Charlestown, presented these findings at the 2015 SNMMI Annual Meeting (abstract 78).
Dr Blasi and his colleagues synthesized FBP8 by conjugating a short cyclic peptide with high affinity for fibrin to a macrocyclic chelator (NODAGA) and labeling it with 64Cu.
The researchers then performed FBP8-PET 1, 3, or 7 days after inducing thrombosis in 30 Sprague-Dawley rats (by applying ferric chloride on the carotid artery and femoral vein). The team also performed FBP8-PET in an animal model of deep vein thrombosis and pulmonary embolism.
FBP8-PET was more than 97% accurate for pinpointing arterial and venous thrombi in the rats. It also detected lung and venous thrombi in the model of deep vein thrombosis and pulmonary embolism. And probe uptake was significantly greater in fresh blood clots than in older ones (P<0.01).
“If approved, fibrin-specific PET could facilitate diagnosis, guide therapeutic choices, and help physicians monitor their patients’ treatment,” said study investigator Peter Caravan, PhD, of Massachusetts General Hospital.
“This technique also offers full-body detection of thrombi with a single injection of probe, instead of the current imaging standards, which are limited to specific parts of the body. A one-time, whole-body scan could prevent unnecessary procedures and uncover hidden thrombi before they generate a deadly embolism.”
Contingent on approval by the US Food and Drug Administration, the researchers expect to conduct a first-in-human study of FBP8-PET as soon as this fall. ![]()

Image by Andre E.X. Brown
BALTIMORE—Scientists say they have synthesized an imaging agent that allows them to detect blood clots throughout the body and estimate the fibrin content of those clots.
In murine experiments, the fibrin-specific PET probe FBP8 detected arterial and venous thrombi with high accuracy.
The tracer also proved sensitive to changes in the fibrin content of clots, which correlated with their age.
Francesco Blasi, PhD, of Massachusetts General Hospital in Charlestown, presented these findings at the 2015 SNMMI Annual Meeting (abstract 78).
Dr Blasi and his colleagues synthesized FBP8 by conjugating a short cyclic peptide with high affinity for fibrin to a macrocyclic chelator (NODAGA) and labeling it with 64Cu.
The researchers then performed FBP8-PET 1, 3, or 7 days after inducing thrombosis in 30 Sprague-Dawley rats (by applying ferric chloride on the carotid artery and femoral vein). The team also performed FBP8-PET in an animal model of deep vein thrombosis and pulmonary embolism.
FBP8-PET was more than 97% accurate for pinpointing arterial and venous thrombi in the rats. It also detected lung and venous thrombi in the model of deep vein thrombosis and pulmonary embolism. And probe uptake was significantly greater in fresh blood clots than in older ones (P<0.01).
“If approved, fibrin-specific PET could facilitate diagnosis, guide therapeutic choices, and help physicians monitor their patients’ treatment,” said study investigator Peter Caravan, PhD, of Massachusetts General Hospital.
“This technique also offers full-body detection of thrombi with a single injection of probe, instead of the current imaging standards, which are limited to specific parts of the body. A one-time, whole-body scan could prevent unnecessary procedures and uncover hidden thrombi before they generate a deadly embolism.”
Contingent on approval by the US Food and Drug Administration, the researchers expect to conduct a first-in-human study of FBP8-PET as soon as this fall. ![]()

Image by Andre E.X. Brown
BALTIMORE—Scientists say they have synthesized an imaging agent that allows them to detect blood clots throughout the body and estimate the fibrin content of those clots.
In murine experiments, the fibrin-specific PET probe FBP8 detected arterial and venous thrombi with high accuracy.
The tracer also proved sensitive to changes in the fibrin content of clots, which correlated with their age.
Francesco Blasi, PhD, of Massachusetts General Hospital in Charlestown, presented these findings at the 2015 SNMMI Annual Meeting (abstract 78).
Dr Blasi and his colleagues synthesized FBP8 by conjugating a short cyclic peptide with high affinity for fibrin to a macrocyclic chelator (NODAGA) and labeling it with 64Cu.
The researchers then performed FBP8-PET 1, 3, or 7 days after inducing thrombosis in 30 Sprague-Dawley rats (by applying ferric chloride on the carotid artery and femoral vein). The team also performed FBP8-PET in an animal model of deep vein thrombosis and pulmonary embolism.
FBP8-PET was more than 97% accurate for pinpointing arterial and venous thrombi in the rats. It also detected lung and venous thrombi in the model of deep vein thrombosis and pulmonary embolism. And probe uptake was significantly greater in fresh blood clots than in older ones (P<0.01).
“If approved, fibrin-specific PET could facilitate diagnosis, guide therapeutic choices, and help physicians monitor their patients’ treatment,” said study investigator Peter Caravan, PhD, of Massachusetts General Hospital.
“This technique also offers full-body detection of thrombi with a single injection of probe, instead of the current imaging standards, which are limited to specific parts of the body. A one-time, whole-body scan could prevent unnecessary procedures and uncover hidden thrombi before they generate a deadly embolism.”
Contingent on approval by the US Food and Drug Administration, the researchers expect to conduct a first-in-human study of FBP8-PET as soon as this fall. ![]()
Mylan recalls gemcitabine, methotrexate

Photo by Bill Branson
Mylan N.V. is conducting a US-wide recall of injectable products due to the presence of visible foreign particulate matter observed in retention samples.
This voluntary recall includes select lots of gemcitabine and a single lot of methotrexate.
Although administration of a sterile injectable containing foreign particulates can cause severe health consequences, Mylan has not received any reports of adverse events related to this recall.
The recall includes the following products:
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801084; Expiration date: 07/2015
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801110; Expiration date: 08/2015
Gemcitabine for Injection, USP 2 g; Package size: 100 mL; NDC number: 67457-463-02; Lot number: 7801221; Expiration date: 03/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801398; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801406; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801427; Expiration date: 09/2016
Gemcitabine for Injection, USP 1 g; Package size: 50 mL; NDC number: 67457-462-01; Lot number: 7801284; Expiration date: 05/2016
Methotrexate Injection, USP 50 mg/2 mL (25 mg/mL); Package size: 5 x 2 mL; NDC number: 67457-467-99; Lot number: 7801421; Expiration date: 09/2016
Gemcitabine for Injection, USP is an intravenously administered product indicated for the treatment of ovarian cancer, breast cancer, non-small cell lung cancer, and pancreatic cancer. These lots were distributed in the US between January 8, 2014, and February 10, 2015. They were manufactured and packaged by Agila Onco Therapies Limited, a Mylan company. Lot 7801084 and 7801110 are packaged with a Pfizer Injectable label.
Methotrexate Injection, USP 25 mg/mL can be administered intramuscularly, intravenously, intra-arterially, or intrathecally and is indicated for certain neoplastic diseases, severe psoriasis, and adult rheumatoid arthritis. The lot was distributed in the US between December 8, 2014, and December 19, 2014, and was packaged by Agila Onco Therapies Limited, a Mylan company.
Mylan is notifying its distributors and customers by letter and is arranging for the return of all recalled products. Distributors, retailers, hospitals, clinics, and physicians with products included in this recall should stop use and return the products to the place of purchase.
Consumers with questions regarding this recall can contact Mylan Customer Relations at 800-796-9526 or [email protected], Monday through Friday from 8 am to 5 pm EST. Consumers should contact their physician or healthcare provider if they have experienced any problems that may be related to using these products.
Adverse reactions or quality problems experienced with the use of this product may be reported to the US Food and Drug Administration’s MedWatch Adverse Event Reporting Program. ![]()

Photo by Bill Branson
Mylan N.V. is conducting a US-wide recall of injectable products due to the presence of visible foreign particulate matter observed in retention samples.
This voluntary recall includes select lots of gemcitabine and a single lot of methotrexate.
Although administration of a sterile injectable containing foreign particulates can cause severe health consequences, Mylan has not received any reports of adverse events related to this recall.
The recall includes the following products:
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801084; Expiration date: 07/2015
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801110; Expiration date: 08/2015
Gemcitabine for Injection, USP 2 g; Package size: 100 mL; NDC number: 67457-463-02; Lot number: 7801221; Expiration date: 03/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801398; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801406; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801427; Expiration date: 09/2016
Gemcitabine for Injection, USP 1 g; Package size: 50 mL; NDC number: 67457-462-01; Lot number: 7801284; Expiration date: 05/2016
Methotrexate Injection, USP 50 mg/2 mL (25 mg/mL); Package size: 5 x 2 mL; NDC number: 67457-467-99; Lot number: 7801421; Expiration date: 09/2016
Gemcitabine for Injection, USP is an intravenously administered product indicated for the treatment of ovarian cancer, breast cancer, non-small cell lung cancer, and pancreatic cancer. These lots were distributed in the US between January 8, 2014, and February 10, 2015. They were manufactured and packaged by Agila Onco Therapies Limited, a Mylan company. Lot 7801084 and 7801110 are packaged with a Pfizer Injectable label.
Methotrexate Injection, USP 25 mg/mL can be administered intramuscularly, intravenously, intra-arterially, or intrathecally and is indicated for certain neoplastic diseases, severe psoriasis, and adult rheumatoid arthritis. The lot was distributed in the US between December 8, 2014, and December 19, 2014, and was packaged by Agila Onco Therapies Limited, a Mylan company.
Mylan is notifying its distributors and customers by letter and is arranging for the return of all recalled products. Distributors, retailers, hospitals, clinics, and physicians with products included in this recall should stop use and return the products to the place of purchase.
Consumers with questions regarding this recall can contact Mylan Customer Relations at 800-796-9526 or [email protected], Monday through Friday from 8 am to 5 pm EST. Consumers should contact their physician or healthcare provider if they have experienced any problems that may be related to using these products.
Adverse reactions or quality problems experienced with the use of this product may be reported to the US Food and Drug Administration’s MedWatch Adverse Event Reporting Program. ![]()

Photo by Bill Branson
Mylan N.V. is conducting a US-wide recall of injectable products due to the presence of visible foreign particulate matter observed in retention samples.
This voluntary recall includes select lots of gemcitabine and a single lot of methotrexate.
Although administration of a sterile injectable containing foreign particulates can cause severe health consequences, Mylan has not received any reports of adverse events related to this recall.
The recall includes the following products:
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801084; Expiration date: 07/2015
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 0069-3857-10; Lot number: 7801110; Expiration date: 08/2015
Gemcitabine for Injection, USP 2 g; Package size: 100 mL; NDC number: 67457-463-02; Lot number: 7801221; Expiration date: 03/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801398; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801406; Expiration date: 08/2016
Gemcitabine for Injection, USP 200 mg; Package size: 10 mL; NDC number: 67457-464-20; Lot number: 7801427; Expiration date: 09/2016
Gemcitabine for Injection, USP 1 g; Package size: 50 mL; NDC number: 67457-462-01; Lot number: 7801284; Expiration date: 05/2016
Methotrexate Injection, USP 50 mg/2 mL (25 mg/mL); Package size: 5 x 2 mL; NDC number: 67457-467-99; Lot number: 7801421; Expiration date: 09/2016
Gemcitabine for Injection, USP is an intravenously administered product indicated for the treatment of ovarian cancer, breast cancer, non-small cell lung cancer, and pancreatic cancer. These lots were distributed in the US between January 8, 2014, and February 10, 2015. They were manufactured and packaged by Agila Onco Therapies Limited, a Mylan company. Lot 7801084 and 7801110 are packaged with a Pfizer Injectable label.
Methotrexate Injection, USP 25 mg/mL can be administered intramuscularly, intravenously, intra-arterially, or intrathecally and is indicated for certain neoplastic diseases, severe psoriasis, and adult rheumatoid arthritis. The lot was distributed in the US between December 8, 2014, and December 19, 2014, and was packaged by Agila Onco Therapies Limited, a Mylan company.
Mylan is notifying its distributors and customers by letter and is arranging for the return of all recalled products. Distributors, retailers, hospitals, clinics, and physicians with products included in this recall should stop use and return the products to the place of purchase.
Consumers with questions regarding this recall can contact Mylan Customer Relations at 800-796-9526 or [email protected], Monday through Friday from 8 am to 5 pm EST. Consumers should contact their physician or healthcare provider if they have experienced any problems that may be related to using these products.
Adverse reactions or quality problems experienced with the use of this product may be reported to the US Food and Drug Administration’s MedWatch Adverse Event Reporting Program. ![]()
Prenatal test can detect lymphoma in mothers
Photo by Graham Colm
GLASGOW—A non-invasive prenatal test (NIPT) used to identify chromosomal fetal disorders can detect maternal cancers before symptoms appear, a new study has shown.
Testing revealed chromosomal abnormalities in 3 women that bore a resemblance to abnormalities observed in cancers. And additional testing confirmed the women had cancer—Hodgkin lymphoma (HL), follicular lymphoma (FL), and ovarian carcinoma.
This research was presented at the European Human Genetics Conference 2015 and published simultaneously in JAMA Oncology.
Nathalie Brison, PhD, of University Hospitals Leuven in Belgium, and her colleagues conducted this research with the goal of improving the NIPT test. They wanted to overcome some of the technical problems that can cause the test to produce false-negative or false-positive results.
“Even though it is very reliable, we believed that we could make it even better, and, in doing so, we could also find other chromosomal abnormalities apart from the traditional trisomy syndromes—Down’s [trisomy 21], Edward’s [trisomy 18], and Patau [trisomy 13],” Dr Brison said.
“Using the new, adapted test in over 6000 pregnancies, and looking at other chromosomes, we identified 3 different genomic abnormalities in 3 women that could not be linked to either the maternal or fetal genomic profile. We realized that the abnormalities bore a resemblance to those found in cancer and referred the women to the oncology unit.”
Further examination, including whole-body MRI scanning and pathological and genetic investigations, revealed the presence of 3 different early stage cancers in the women: ovarian carcinoma, FL, and HL.
The researchers said that, without NIPT, these cancers likely would not have been detected until the women developed symptoms.
“Considering the bad prognosis of some cancers when detected later, and given that we know that it is both possible and safe to treat the disease during pregnancy, this is an important added advantage of NIPT,” said study author Joris Vermeesch, PhD, also of University Hospitals Leuven.
“During pregnancy, cancer-related symptoms may well be masked. Fatigue, nausea, abdominal pain, and vaginal blood loss are easily interpretable as a normal part of being pregnant. NIPT offers an opportunity for the accurate screening of high-risk women for cancer, allowing us to overcome the challenge of early diagnosis in pregnant women.”
Two of the 3 women diagnosed with cancer were treated. The woman with ovarian cancer was treated after delivery.
The woman with HL was treated during pregnancy and ultimately gave birth to a healthy girl. The woman with FL has indolent disease and may not require treatment for years, according to the researchers.
Follow-up investigations in the treated women showed that NIPT had the additional advantage of allowing for treatment monitoring. The researchers were able to see that chromosomal profiles became normal during and after chemotherapy.
Because NIPT involves looking at chromosomes other than 13, 18, and 21, the women taking part in this study were informed about the possibility of incidental findings.
“However, our study feeds into the ethical debate about whether or not to report incidental findings to patients and also has implications for the current political discussions concerning reimbursement and funding of NIPT by national healthcare systems,” Dr Vermeesch said.
The results also suggest that NIPT might enable the detection of pre-symptomatic cancers in the general population.
“We now know that it is possible to offer the accurate detection of chromosomally imbalanced cancers to the general population via minimally invasive screening methods,” Dr Brison said. “The normalization of the NIPT profile in these patients following treatment indicates that we can also measure response to treatment as early as after the first administration of chemotherapy.”
“Of course, larger-scale studies will be required to validate these results further, but we are confident that we have made an important step towards the possibility of wide-scale, effective, non-invasive cancer screening capable of detecting disease at an early stage.” ![]()

Photo by Graham Colm
GLASGOW—A non-invasive prenatal test (NIPT) used to identify chromosomal fetal disorders can detect maternal cancers before symptoms appear, a new study has shown.
Testing revealed chromosomal abnormalities in 3 women that bore a resemblance to abnormalities observed in cancers. And additional testing confirmed the women had cancer—Hodgkin lymphoma (HL), follicular lymphoma (FL), and ovarian carcinoma.
This research was presented at the European Human Genetics Conference 2015 and published simultaneously in JAMA Oncology.
Nathalie Brison, PhD, of University Hospitals Leuven in Belgium, and her colleagues conducted this research with the goal of improving the NIPT test. They wanted to overcome some of the technical problems that can cause the test to produce false-negative or false-positive results.
“Even though it is very reliable, we believed that we could make it even better, and, in doing so, we could also find other chromosomal abnormalities apart from the traditional trisomy syndromes—Down’s [trisomy 21], Edward’s [trisomy 18], and Patau [trisomy 13],” Dr Brison said.
“Using the new, adapted test in over 6000 pregnancies, and looking at other chromosomes, we identified 3 different genomic abnormalities in 3 women that could not be linked to either the maternal or fetal genomic profile. We realized that the abnormalities bore a resemblance to those found in cancer and referred the women to the oncology unit.”
Further examination, including whole-body MRI scanning and pathological and genetic investigations, revealed the presence of 3 different early stage cancers in the women: ovarian carcinoma, FL, and HL.
The researchers said that, without NIPT, these cancers likely would not have been detected until the women developed symptoms.
“Considering the bad prognosis of some cancers when detected later, and given that we know that it is both possible and safe to treat the disease during pregnancy, this is an important added advantage of NIPT,” said study author Joris Vermeesch, PhD, also of University Hospitals Leuven.
“During pregnancy, cancer-related symptoms may well be masked. Fatigue, nausea, abdominal pain, and vaginal blood loss are easily interpretable as a normal part of being pregnant. NIPT offers an opportunity for the accurate screening of high-risk women for cancer, allowing us to overcome the challenge of early diagnosis in pregnant women.”
Two of the 3 women diagnosed with cancer were treated. The woman with ovarian cancer was treated after delivery.
The woman with HL was treated during pregnancy and ultimately gave birth to a healthy girl. The woman with FL has indolent disease and may not require treatment for years, according to the researchers.
Follow-up investigations in the treated women showed that NIPT had the additional advantage of allowing for treatment monitoring. The researchers were able to see that chromosomal profiles became normal during and after chemotherapy.
Because NIPT involves looking at chromosomes other than 13, 18, and 21, the women taking part in this study were informed about the possibility of incidental findings.
“However, our study feeds into the ethical debate about whether or not to report incidental findings to patients and also has implications for the current political discussions concerning reimbursement and funding of NIPT by national healthcare systems,” Dr Vermeesch said.
The results also suggest that NIPT might enable the detection of pre-symptomatic cancers in the general population.
“We now know that it is possible to offer the accurate detection of chromosomally imbalanced cancers to the general population via minimally invasive screening methods,” Dr Brison said. “The normalization of the NIPT profile in these patients following treatment indicates that we can also measure response to treatment as early as after the first administration of chemotherapy.”
“Of course, larger-scale studies will be required to validate these results further, but we are confident that we have made an important step towards the possibility of wide-scale, effective, non-invasive cancer screening capable of detecting disease at an early stage.” ![]()

Photo by Graham Colm
GLASGOW—A non-invasive prenatal test (NIPT) used to identify chromosomal fetal disorders can detect maternal cancers before symptoms appear, a new study has shown.
Testing revealed chromosomal abnormalities in 3 women that bore a resemblance to abnormalities observed in cancers. And additional testing confirmed the women had cancer—Hodgkin lymphoma (HL), follicular lymphoma (FL), and ovarian carcinoma.
This research was presented at the European Human Genetics Conference 2015 and published simultaneously in JAMA Oncology.
Nathalie Brison, PhD, of University Hospitals Leuven in Belgium, and her colleagues conducted this research with the goal of improving the NIPT test. They wanted to overcome some of the technical problems that can cause the test to produce false-negative or false-positive results.
“Even though it is very reliable, we believed that we could make it even better, and, in doing so, we could also find other chromosomal abnormalities apart from the traditional trisomy syndromes—Down’s [trisomy 21], Edward’s [trisomy 18], and Patau [trisomy 13],” Dr Brison said.
“Using the new, adapted test in over 6000 pregnancies, and looking at other chromosomes, we identified 3 different genomic abnormalities in 3 women that could not be linked to either the maternal or fetal genomic profile. We realized that the abnormalities bore a resemblance to those found in cancer and referred the women to the oncology unit.”
Further examination, including whole-body MRI scanning and pathological and genetic investigations, revealed the presence of 3 different early stage cancers in the women: ovarian carcinoma, FL, and HL.
The researchers said that, without NIPT, these cancers likely would not have been detected until the women developed symptoms.
“Considering the bad prognosis of some cancers when detected later, and given that we know that it is both possible and safe to treat the disease during pregnancy, this is an important added advantage of NIPT,” said study author Joris Vermeesch, PhD, also of University Hospitals Leuven.
“During pregnancy, cancer-related symptoms may well be masked. Fatigue, nausea, abdominal pain, and vaginal blood loss are easily interpretable as a normal part of being pregnant. NIPT offers an opportunity for the accurate screening of high-risk women for cancer, allowing us to overcome the challenge of early diagnosis in pregnant women.”
Two of the 3 women diagnosed with cancer were treated. The woman with ovarian cancer was treated after delivery.
The woman with HL was treated during pregnancy and ultimately gave birth to a healthy girl. The woman with FL has indolent disease and may not require treatment for years, according to the researchers.
Follow-up investigations in the treated women showed that NIPT had the additional advantage of allowing for treatment monitoring. The researchers were able to see that chromosomal profiles became normal during and after chemotherapy.
Because NIPT involves looking at chromosomes other than 13, 18, and 21, the women taking part in this study were informed about the possibility of incidental findings.
“However, our study feeds into the ethical debate about whether or not to report incidental findings to patients and also has implications for the current political discussions concerning reimbursement and funding of NIPT by national healthcare systems,” Dr Vermeesch said.
The results also suggest that NIPT might enable the detection of pre-symptomatic cancers in the general population.
“We now know that it is possible to offer the accurate detection of chromosomally imbalanced cancers to the general population via minimally invasive screening methods,” Dr Brison said. “The normalization of the NIPT profile in these patients following treatment indicates that we can also measure response to treatment as early as after the first administration of chemotherapy.”
“Of course, larger-scale studies will be required to validate these results further, but we are confident that we have made an important step towards the possibility of wide-scale, effective, non-invasive cancer screening capable of detecting disease at an early stage.” ![]()
Host cell type may impact malaria treatment
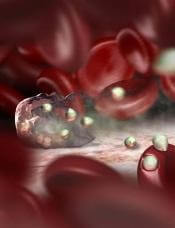
Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism. ![]()

Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism. ![]()

Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism. ![]()
BCL-2 inhibitor shows potential for treating MM

to an attendee at ASCO 2015
© ASCO/Zach Boyden-Holmes
CHICAGO—A BCL-2 inhibitor that has shown activity in patients with chronic and acute leukemias could be a feasible treatment option for multiple myeloma (MM) too, according to research presented at the 2015 ASCO Annual Meeting.
In a pair of phase 1 studies, the drug, venetoclax, prompted responses in certain patients with relapsed or refractory MM, both when given alone and in combination with bortezomib and dexamethasone.
As monotherapy, venetoclax induced complete responses (CRs) in 2 patients with t(11;14), but the overall response rate (ORR) was low, and most patients discontinued treatment, largely due to progression.
Venetoclax in combination with bortezomib and dexamethasone prompted a response in nearly half of the patients studied, but more than half discontinued treatment. The combination produced a high ORR in patients who were bortezomib-naïve and bortezomib-sensitive, but there was no response among bortezomib-refractory patients.
“Multiple myeloma remains a high area of unmet medical need, and additional research to identify new therapies is important,” said investigator Cyrille Touzeau, MD, of the University of Nantes in France.
“The response rates shown in these studies suggest potential of venetoclax in this patient population and warrant further evaluation.”
Dr Touzeau and his colleagues presented these studies at ASCO as abstracts 8580 and 8576. Both studies were funded by AbbVie, the company developing venetoclax.
Venetoclax in combination
In a phase 1b study (abstract 8580*), the researchers evaluated venetoclax in combination with bortezomib and dexamethasone in 38 patients with relapsed or refractory MM.
The patients’ median age was 65 (range, 38-79). Five patients were t(11;14)-positive, 3 were t(4;14)-positive, 9 had del17p, and 19 had del13q. They had received a median of 5 prior lines of therapy (range, 1-15). Ten patients were bortezomib-refractory, 21 were lenalidomide-refractory, and 8 patients were refractory to both drugs.
The patients received venetoclax at doses ranging from 50 mg to 600 mg daily (with a 1-week lead-in period for the 50 mg cohort). On cycles 1-8, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 4, 8, and 11. They received dexamethasone (20 mg) with venetoclax on days 2, 5, 9, and 12.
On cycles 9-11, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 8, 15, and 22. Patients received venetoclax monotherapy for cycles 12 and beyond. They also received prophylaxis for tumor lysis syndrome.
Twenty-five patients (66%) discontinued treatment, 19 due to disease progression (including 4 deaths), 2 due to adverse events (AEs), 3 due to patient decision, and 1 due to withdrawn consent.
The ORR was 47% among the 36 evaluable patients, 83% for bortezomib-naïve patients (5/6), 60% for bortezomib-sensitive patients (12/20), and 0% for bortezomib-refractory patients (0/10).
Two patients had a stringent CR, 1 had a CR, 4 had a very good partial response, 10 had a partial response, and 2 had a minimal response. Six patients had stable disease, and 9 progressed.
The most common AEs—occurring in at least 20% of patients—were constipation (37%), diarrhea (32%), insomnia (32%), thrombocytopenia (29%), peripheral neuropathy (26%), asthenia (26%), dyspnea (26%), peripheral edema (24%), and anemia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (21%), anemia (13%), and dyspnea (11%). Serious AEs—each occurring in 5% of patients—were cardiac failure, pyrexia, sepsis, respiratory failure, pneumonia, and embolism.
Venetoclax as monotherapy
In a phase 1 study (abstract 8576*), researchers evaluated venetoclax as monotherapy in 29 patients with relapsed or refractory MM. Eleven patients were t(11;14)-positive, 2 were t(4;14)-positive, 4 had del17p, and 14 had del13q.
The patients’ median age was 66 (range, 42-79), and they had received a median of 6 prior therapies (range, 1-13). Fifteen patients were bortezomib-refractory, 12 were refractory to lenalidomide, and 10 patients were refractory to both drugs.
After a 2-week lead-in period, patients received venetoclax daily on a 21-day cycle, ranging from 300 mg to 1200 mg on a 3+3 design. They also received prophylaxis for tumor lysis syndrome. Patients who progressed during treatment were allowed to receive dexamethasone as well and continue the study.
Twenty-three patients (79%) discontinued treatment, 18 due to disease progression, 4 due to AEs, and 1 due to patient decision.
The ORR was 7% (n=2). Both patients achieved a CR, and both were t(11;14)-positive. One of the patients is in the 600 mg cohort and is still responding to treatment. The duration of response, thus far, is 2.1 months.
The other patient with a CR is in the 900 mg dose cohort. This patient is still responding as well, and the response duration, thus far, is 2.8 months.
Common AEs—occurring in at least 20% of patients—were diarrhea (41%), nausea (41%), fatigue (24%), vomiting (21%), asthenia (21%), and neutropenia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (24%), neutropenia (14%), and anemia (10%). Serious AEs—each occurring in 7% of patients—were pyrexia, malignant neoplasm progression, and cough.
Two patients had dose-limiting toxicities at the 600 mg dose of venetoclax. However, the researchers said the maximum tolerated dose was not reached, and the recommended phase 2 dose is 1200 mg. ![]()
*Information in the abstract differs from that presented at the meeting.

to an attendee at ASCO 2015
© ASCO/Zach Boyden-Holmes
CHICAGO—A BCL-2 inhibitor that has shown activity in patients with chronic and acute leukemias could be a feasible treatment option for multiple myeloma (MM) too, according to research presented at the 2015 ASCO Annual Meeting.
In a pair of phase 1 studies, the drug, venetoclax, prompted responses in certain patients with relapsed or refractory MM, both when given alone and in combination with bortezomib and dexamethasone.
As monotherapy, venetoclax induced complete responses (CRs) in 2 patients with t(11;14), but the overall response rate (ORR) was low, and most patients discontinued treatment, largely due to progression.
Venetoclax in combination with bortezomib and dexamethasone prompted a response in nearly half of the patients studied, but more than half discontinued treatment. The combination produced a high ORR in patients who were bortezomib-naïve and bortezomib-sensitive, but there was no response among bortezomib-refractory patients.
“Multiple myeloma remains a high area of unmet medical need, and additional research to identify new therapies is important,” said investigator Cyrille Touzeau, MD, of the University of Nantes in France.
“The response rates shown in these studies suggest potential of venetoclax in this patient population and warrant further evaluation.”
Dr Touzeau and his colleagues presented these studies at ASCO as abstracts 8580 and 8576. Both studies were funded by AbbVie, the company developing venetoclax.
Venetoclax in combination
In a phase 1b study (abstract 8580*), the researchers evaluated venetoclax in combination with bortezomib and dexamethasone in 38 patients with relapsed or refractory MM.
The patients’ median age was 65 (range, 38-79). Five patients were t(11;14)-positive, 3 were t(4;14)-positive, 9 had del17p, and 19 had del13q. They had received a median of 5 prior lines of therapy (range, 1-15). Ten patients were bortezomib-refractory, 21 were lenalidomide-refractory, and 8 patients were refractory to both drugs.
The patients received venetoclax at doses ranging from 50 mg to 600 mg daily (with a 1-week lead-in period for the 50 mg cohort). On cycles 1-8, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 4, 8, and 11. They received dexamethasone (20 mg) with venetoclax on days 2, 5, 9, and 12.
On cycles 9-11, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 8, 15, and 22. Patients received venetoclax monotherapy for cycles 12 and beyond. They also received prophylaxis for tumor lysis syndrome.
Twenty-five patients (66%) discontinued treatment, 19 due to disease progression (including 4 deaths), 2 due to adverse events (AEs), 3 due to patient decision, and 1 due to withdrawn consent.
The ORR was 47% among the 36 evaluable patients, 83% for bortezomib-naïve patients (5/6), 60% for bortezomib-sensitive patients (12/20), and 0% for bortezomib-refractory patients (0/10).
Two patients had a stringent CR, 1 had a CR, 4 had a very good partial response, 10 had a partial response, and 2 had a minimal response. Six patients had stable disease, and 9 progressed.
The most common AEs—occurring in at least 20% of patients—were constipation (37%), diarrhea (32%), insomnia (32%), thrombocytopenia (29%), peripheral neuropathy (26%), asthenia (26%), dyspnea (26%), peripheral edema (24%), and anemia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (21%), anemia (13%), and dyspnea (11%). Serious AEs—each occurring in 5% of patients—were cardiac failure, pyrexia, sepsis, respiratory failure, pneumonia, and embolism.
Venetoclax as monotherapy
In a phase 1 study (abstract 8576*), researchers evaluated venetoclax as monotherapy in 29 patients with relapsed or refractory MM. Eleven patients were t(11;14)-positive, 2 were t(4;14)-positive, 4 had del17p, and 14 had del13q.
The patients’ median age was 66 (range, 42-79), and they had received a median of 6 prior therapies (range, 1-13). Fifteen patients were bortezomib-refractory, 12 were refractory to lenalidomide, and 10 patients were refractory to both drugs.
After a 2-week lead-in period, patients received venetoclax daily on a 21-day cycle, ranging from 300 mg to 1200 mg on a 3+3 design. They also received prophylaxis for tumor lysis syndrome. Patients who progressed during treatment were allowed to receive dexamethasone as well and continue the study.
Twenty-three patients (79%) discontinued treatment, 18 due to disease progression, 4 due to AEs, and 1 due to patient decision.
The ORR was 7% (n=2). Both patients achieved a CR, and both were t(11;14)-positive. One of the patients is in the 600 mg cohort and is still responding to treatment. The duration of response, thus far, is 2.1 months.
The other patient with a CR is in the 900 mg dose cohort. This patient is still responding as well, and the response duration, thus far, is 2.8 months.
Common AEs—occurring in at least 20% of patients—were diarrhea (41%), nausea (41%), fatigue (24%), vomiting (21%), asthenia (21%), and neutropenia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (24%), neutropenia (14%), and anemia (10%). Serious AEs—each occurring in 7% of patients—were pyrexia, malignant neoplasm progression, and cough.
Two patients had dose-limiting toxicities at the 600 mg dose of venetoclax. However, the researchers said the maximum tolerated dose was not reached, and the recommended phase 2 dose is 1200 mg. ![]()
*Information in the abstract differs from that presented at the meeting.

to an attendee at ASCO 2015
© ASCO/Zach Boyden-Holmes
CHICAGO—A BCL-2 inhibitor that has shown activity in patients with chronic and acute leukemias could be a feasible treatment option for multiple myeloma (MM) too, according to research presented at the 2015 ASCO Annual Meeting.
In a pair of phase 1 studies, the drug, venetoclax, prompted responses in certain patients with relapsed or refractory MM, both when given alone and in combination with bortezomib and dexamethasone.
As monotherapy, venetoclax induced complete responses (CRs) in 2 patients with t(11;14), but the overall response rate (ORR) was low, and most patients discontinued treatment, largely due to progression.
Venetoclax in combination with bortezomib and dexamethasone prompted a response in nearly half of the patients studied, but more than half discontinued treatment. The combination produced a high ORR in patients who were bortezomib-naïve and bortezomib-sensitive, but there was no response among bortezomib-refractory patients.
“Multiple myeloma remains a high area of unmet medical need, and additional research to identify new therapies is important,” said investigator Cyrille Touzeau, MD, of the University of Nantes in France.
“The response rates shown in these studies suggest potential of venetoclax in this patient population and warrant further evaluation.”
Dr Touzeau and his colleagues presented these studies at ASCO as abstracts 8580 and 8576. Both studies were funded by AbbVie, the company developing venetoclax.
Venetoclax in combination
In a phase 1b study (abstract 8580*), the researchers evaluated venetoclax in combination with bortezomib and dexamethasone in 38 patients with relapsed or refractory MM.
The patients’ median age was 65 (range, 38-79). Five patients were t(11;14)-positive, 3 were t(4;14)-positive, 9 had del17p, and 19 had del13q. They had received a median of 5 prior lines of therapy (range, 1-15). Ten patients were bortezomib-refractory, 21 were lenalidomide-refractory, and 8 patients were refractory to both drugs.
The patients received venetoclax at doses ranging from 50 mg to 600 mg daily (with a 1-week lead-in period for the 50 mg cohort). On cycles 1-8, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 4, 8, and 11. They received dexamethasone (20 mg) with venetoclax on days 2, 5, 9, and 12.
On cycles 9-11, patients received dexamethasone (20 mg) and bortezomib (1.3 mg/m2) with venetoclax on days 1, 8, 15, and 22. Patients received venetoclax monotherapy for cycles 12 and beyond. They also received prophylaxis for tumor lysis syndrome.
Twenty-five patients (66%) discontinued treatment, 19 due to disease progression (including 4 deaths), 2 due to adverse events (AEs), 3 due to patient decision, and 1 due to withdrawn consent.
The ORR was 47% among the 36 evaluable patients, 83% for bortezomib-naïve patients (5/6), 60% for bortezomib-sensitive patients (12/20), and 0% for bortezomib-refractory patients (0/10).
Two patients had a stringent CR, 1 had a CR, 4 had a very good partial response, 10 had a partial response, and 2 had a minimal response. Six patients had stable disease, and 9 progressed.
The most common AEs—occurring in at least 20% of patients—were constipation (37%), diarrhea (32%), insomnia (32%), thrombocytopenia (29%), peripheral neuropathy (26%), asthenia (26%), dyspnea (26%), peripheral edema (24%), and anemia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (21%), anemia (13%), and dyspnea (11%). Serious AEs—each occurring in 5% of patients—were cardiac failure, pyrexia, sepsis, respiratory failure, pneumonia, and embolism.
Venetoclax as monotherapy
In a phase 1 study (abstract 8576*), researchers evaluated venetoclax as monotherapy in 29 patients with relapsed or refractory MM. Eleven patients were t(11;14)-positive, 2 were t(4;14)-positive, 4 had del17p, and 14 had del13q.
The patients’ median age was 66 (range, 42-79), and they had received a median of 6 prior therapies (range, 1-13). Fifteen patients were bortezomib-refractory, 12 were refractory to lenalidomide, and 10 patients were refractory to both drugs.
After a 2-week lead-in period, patients received venetoclax daily on a 21-day cycle, ranging from 300 mg to 1200 mg on a 3+3 design. They also received prophylaxis for tumor lysis syndrome. Patients who progressed during treatment were allowed to receive dexamethasone as well and continue the study.
Twenty-three patients (79%) discontinued treatment, 18 due to disease progression, 4 due to AEs, and 1 due to patient decision.
The ORR was 7% (n=2). Both patients achieved a CR, and both were t(11;14)-positive. One of the patients is in the 600 mg cohort and is still responding to treatment. The duration of response, thus far, is 2.1 months.
The other patient with a CR is in the 900 mg dose cohort. This patient is still responding as well, and the response duration, thus far, is 2.8 months.
Common AEs—occurring in at least 20% of patients—were diarrhea (41%), nausea (41%), fatigue (24%), vomiting (21%), asthenia (21%), and neutropenia (21%).
Grade 3/4 AEs—occurring in at least 10% of patients—were thrombocytopenia (24%), neutropenia (14%), and anemia (10%). Serious AEs—each occurring in 7% of patients—were pyrexia, malignant neoplasm progression, and cough.
Two patients had dose-limiting toxicities at the 600 mg dose of venetoclax. However, the researchers said the maximum tolerated dose was not reached, and the recommended phase 2 dose is 1200 mg.
*Information in the abstract differs from that presented at the meeting.
Resolution draws attention to sickle cell trait

and a normal one
Image by Betty Pace
Two members of the US House of Representatives have introduced a resolution calling for expanded government efforts to increase sickle cell trait (SCT) research and improve access to SCT screening and education.
Barbara Lee (D-CA) and Michael C. Burgess (R-TX) introduced the resolution, which was assigned to a congressional committee that will consider it before possibly sending it on to the House or Senate as a whole.
The resolution, H.Res.296, encourages the medical community as well as state and federal governments to work to ensure that all individuals are made aware of their SCT status by developing a common strategy for disseminating screening results and providing education and counseling to parents and families in collaboration with all 50 states’ newborn screening programs.
The resolution calls on the US Department of Health and Human Services to develop a public awareness campaign stressing the importance of knowing your SCT status and to expand access for screening and appropriate counseling for SCT carriers.
The resolution also says the House commits to ensuring support for research that expands our understanding of SCT and sickle cell disease.
The American Society of Hematology (ASH) said it applauds this bipartisan effort to advance public understanding of SCT, which is estimated to affect more than 300,000 people in the US, many of whom are unaware of their status.
ASH also said it has identified several priorities for research in this area and has collaborated with the US Centers for Disease Control and Prevention and the Sickle Cell Disease Association of America to develop a Sickle Cell Trait Toolkit for SCT carriers and their families.
To follow the progress of the resolution, H.Res.296 – Calling for Sickle Cell Trait Research, visit Congress.gov.

and a normal one
Image by Betty Pace
Two members of the US House of Representatives have introduced a resolution calling for expanded government efforts to increase sickle cell trait (SCT) research and improve access to SCT screening and education.
Barbara Lee (D-CA) and Michael C. Burgess (R-TX) introduced the resolution, which was assigned to a congressional committee that will consider it before possibly sending it on to the House or Senate as a whole.
The resolution, H.Res.296, encourages the medical community as well as state and federal governments to work to ensure that all individuals are made aware of their SCT status by developing a common strategy for disseminating screening results and providing education and counseling to parents and families in collaboration with all 50 states’ newborn screening programs.
The resolution calls on the US Department of Health and Human Services to develop a public awareness campaign stressing the importance of knowing your SCT status and to expand access for screening and appropriate counseling for SCT carriers.
The resolution also says the House commits to ensuring support for research that expands our understanding of SCT and sickle cell disease.
The American Society of Hematology (ASH) said it applauds this bipartisan effort to advance public understanding of SCT, which is estimated to affect more than 300,000 people in the US, many of whom are unaware of their status.
ASH also said it has identified several priorities for research in this area and has collaborated with the US Centers for Disease Control and Prevention and the Sickle Cell Disease Association of America to develop a Sickle Cell Trait Toolkit for SCT carriers and their families.
To follow the progress of the resolution, H.Res.296 – Calling for Sickle Cell Trait Research, visit Congress.gov.

and a normal one
Image by Betty Pace
Two members of the US House of Representatives have introduced a resolution calling for expanded government efforts to increase sickle cell trait (SCT) research and improve access to SCT screening and education.
Barbara Lee (D-CA) and Michael C. Burgess (R-TX) introduced the resolution, which was assigned to a congressional committee that will consider it before possibly sending it on to the House or Senate as a whole.
The resolution, H.Res.296, encourages the medical community as well as state and federal governments to work to ensure that all individuals are made aware of their SCT status by developing a common strategy for disseminating screening results and providing education and counseling to parents and families in collaboration with all 50 states’ newborn screening programs.
The resolution calls on the US Department of Health and Human Services to develop a public awareness campaign stressing the importance of knowing your SCT status and to expand access for screening and appropriate counseling for SCT carriers.
The resolution also says the House commits to ensuring support for research that expands our understanding of SCT and sickle cell disease.
The American Society of Hematology (ASH) said it applauds this bipartisan effort to advance public understanding of SCT, which is estimated to affect more than 300,000 people in the US, many of whom are unaware of their status.
ASH also said it has identified several priorities for research in this area and has collaborated with the US Centers for Disease Control and Prevention and the Sickle Cell Disease Association of America to develop a Sickle Cell Trait Toolkit for SCT carriers and their families.
To follow the progress of the resolution, H.Res.296 – Calling for Sickle Cell Trait Research, visit Congress.gov.
Blood test screens for many viruses simultaneously
Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.

Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.

Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.
Biochemist Irwin Rose dies at 88

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.