User login
Reporting requirements may affect trial outcomes

Photo by Esther Dyson
The reporting requirements developed to increase transparency in US medical research may lead to fewer positive trial outcomes, according to a study published in PLOS ONE.
Researchers analyzed data from large-budget trials funded by the National Heart, Lung and Blood Institute (NHLBI).
And they found evidence suggesting the reporting requirements may have contributed to a significant reduction in studies with positive findings.
The reporting standards were phased in around 2000. They require researchers conducting drug or dietary supplement trials using human subjects to identify
projected outcomes and register their trials on ClinicalTrials.gov before they begin to collect data.
When entering their trial into the database, researchers are required to state the specific outcome on which they will focus. In the past, a researcher might have published an aspect of a study that was successful, even if the study overall did not produce the expected results.
But the new requirements mean researchers are less likely to change their analysis plan to consider another outcome that may have shown a positive result, said Veronica L. Irvin, PhD, of Oregon State University in Corvallis.
Dr Irvin began working on this project with the study’s lead author, Robert M. Kaplan, PhD, of the Agency for Healthcare Research and Quality in Rockville,
Maryland, while the two worked together in the National Institutes of Health’s Office of Behavior and Social Science Research.
The pair reviewed all large-budget clinical trials evaluating drugs or dietary supplements for the treatment or prevention of cardiovascular disease that had received funding from the NHLBI between 1970 and 2012.
They chose large-budget, NHLBI-funded trials in part because outcomes from the trials were more likely to be published, even if they did not produce the expected result.
Fifty-five studies were included in the research. Thirty were published prior to the reporting changes in 2000 (1970 to 1999), and 25 were published after the changes (2000 to 2012).
Of the studies published after 2000, only 2 (8%) showed positive outcomes, while 17 (57%) of the studies published before 2000 showed positive results.
Drs Kaplan and Irvin acknowledged that factors other than the reporting requirements may be contributing to the decline in positive outcomes, but they were unable to identify other compelling explanations.
For example, one suggestion was that older trials were more likely to compare new treatments to placebos, while newer trials were more likely to compare new
treatments to established treatments.
But when Drs Kaplan and Irvin examined the data, they found that 60% of trials published before 2000 used placebo comparators and nearly the same amount, 64%, of trials published after 2000 used placebos.
The researchers noted that although this work focused on clinical trials related to cardiovascular health, it would be reasonable to see similar changes in results
across other disease types.
“We don’t know if this decrease in positive outcomes also affects drug trials for prevention and treatment of cancer, diabetes, or other diseases,” Dr Irvin said. “But it would not be surprising because they have the same reporting requirements.” ![]()

Photo by Esther Dyson
The reporting requirements developed to increase transparency in US medical research may lead to fewer positive trial outcomes, according to a study published in PLOS ONE.
Researchers analyzed data from large-budget trials funded by the National Heart, Lung and Blood Institute (NHLBI).
And they found evidence suggesting the reporting requirements may have contributed to a significant reduction in studies with positive findings.
The reporting standards were phased in around 2000. They require researchers conducting drug or dietary supplement trials using human subjects to identify
projected outcomes and register their trials on ClinicalTrials.gov before they begin to collect data.
When entering their trial into the database, researchers are required to state the specific outcome on which they will focus. In the past, a researcher might have published an aspect of a study that was successful, even if the study overall did not produce the expected results.
But the new requirements mean researchers are less likely to change their analysis plan to consider another outcome that may have shown a positive result, said Veronica L. Irvin, PhD, of Oregon State University in Corvallis.
Dr Irvin began working on this project with the study’s lead author, Robert M. Kaplan, PhD, of the Agency for Healthcare Research and Quality in Rockville,
Maryland, while the two worked together in the National Institutes of Health’s Office of Behavior and Social Science Research.
The pair reviewed all large-budget clinical trials evaluating drugs or dietary supplements for the treatment or prevention of cardiovascular disease that had received funding from the NHLBI between 1970 and 2012.
They chose large-budget, NHLBI-funded trials in part because outcomes from the trials were more likely to be published, even if they did not produce the expected result.
Fifty-five studies were included in the research. Thirty were published prior to the reporting changes in 2000 (1970 to 1999), and 25 were published after the changes (2000 to 2012).
Of the studies published after 2000, only 2 (8%) showed positive outcomes, while 17 (57%) of the studies published before 2000 showed positive results.
Drs Kaplan and Irvin acknowledged that factors other than the reporting requirements may be contributing to the decline in positive outcomes, but they were unable to identify other compelling explanations.
For example, one suggestion was that older trials were more likely to compare new treatments to placebos, while newer trials were more likely to compare new
treatments to established treatments.
But when Drs Kaplan and Irvin examined the data, they found that 60% of trials published before 2000 used placebo comparators and nearly the same amount, 64%, of trials published after 2000 used placebos.
The researchers noted that although this work focused on clinical trials related to cardiovascular health, it would be reasonable to see similar changes in results
across other disease types.
“We don’t know if this decrease in positive outcomes also affects drug trials for prevention and treatment of cancer, diabetes, or other diseases,” Dr Irvin said. “But it would not be surprising because they have the same reporting requirements.” ![]()

Photo by Esther Dyson
The reporting requirements developed to increase transparency in US medical research may lead to fewer positive trial outcomes, according to a study published in PLOS ONE.
Researchers analyzed data from large-budget trials funded by the National Heart, Lung and Blood Institute (NHLBI).
And they found evidence suggesting the reporting requirements may have contributed to a significant reduction in studies with positive findings.
The reporting standards were phased in around 2000. They require researchers conducting drug or dietary supplement trials using human subjects to identify
projected outcomes and register their trials on ClinicalTrials.gov before they begin to collect data.
When entering their trial into the database, researchers are required to state the specific outcome on which they will focus. In the past, a researcher might have published an aspect of a study that was successful, even if the study overall did not produce the expected results.
But the new requirements mean researchers are less likely to change their analysis plan to consider another outcome that may have shown a positive result, said Veronica L. Irvin, PhD, of Oregon State University in Corvallis.
Dr Irvin began working on this project with the study’s lead author, Robert M. Kaplan, PhD, of the Agency for Healthcare Research and Quality in Rockville,
Maryland, while the two worked together in the National Institutes of Health’s Office of Behavior and Social Science Research.
The pair reviewed all large-budget clinical trials evaluating drugs or dietary supplements for the treatment or prevention of cardiovascular disease that had received funding from the NHLBI between 1970 and 2012.
They chose large-budget, NHLBI-funded trials in part because outcomes from the trials were more likely to be published, even if they did not produce the expected result.
Fifty-five studies were included in the research. Thirty were published prior to the reporting changes in 2000 (1970 to 1999), and 25 were published after the changes (2000 to 2012).
Of the studies published after 2000, only 2 (8%) showed positive outcomes, while 17 (57%) of the studies published before 2000 showed positive results.
Drs Kaplan and Irvin acknowledged that factors other than the reporting requirements may be contributing to the decline in positive outcomes, but they were unable to identify other compelling explanations.
For example, one suggestion was that older trials were more likely to compare new treatments to placebos, while newer trials were more likely to compare new
treatments to established treatments.
But when Drs Kaplan and Irvin examined the data, they found that 60% of trials published before 2000 used placebo comparators and nearly the same amount, 64%, of trials published after 2000 used placebos.
The researchers noted that although this work focused on clinical trials related to cardiovascular health, it would be reasonable to see similar changes in results
across other disease types.
“We don’t know if this decrease in positive outcomes also affects drug trials for prevention and treatment of cancer, diabetes, or other diseases,” Dr Irvin said. “But it would not be surprising because they have the same reporting requirements.” ![]()
T-cell activation is a dynamic process, study shows
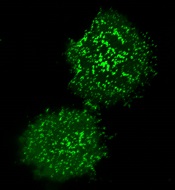
plasma membrane upon T-cell
activation. Image courtesy
of the Salk Institute
New research suggests that, contrary to previous assumptions, T-cell activation is a fluid process that relies on a dynamic protein network at the cell surface.
Previously, scientists thought T-cell activation was a static process in which molecules assemble within a T cell.
Instead, investigators discovered that molecules rapidly come and go, and the times at which these molecules arrive and depart affect the immune response.
“This is a completely new principle for how T-cell activity is controlled—whether it ignores or responds to a threat,” said Björn Lillemeier, PhD, of the Salk Institute in La Jolla, California.
He and his colleagues described this discovery in Nature Immunology.
They noted that the protein kinase ZAP-70 plays a central role in T-cell activation. Until now, scientists assumed that a silent form of ZAP-70 floats around inside the T cell until a threat is detected. The threat recruits ZAP-70 to the cell surface and activates it.
By analyzing mutant forms of ZAP-70, Dr Lillemeier’s group discovered that, instead of ZAP-70 binding the T-cell receptor firmly, it comes in contact with the receptor sporadically. Each time this happens, ZAP-70 has to adopt an unfavorable shape that forces it back inside the cell.
This cycle continues until a second molecule, Lck, helps ZAP-70 remain with the T-cell receptor. The prolonged stay at the cell surface activates ZAP-70 and prompts the T cell to attack invaders and diseased cells.
The investigators said this study suggests the steps underlying T-cell activation are more dynamic than the less mobile modes scientists had suspected before. And the research highlights how ZAP-70 and other molecules communicate in space and time, which is crucial for controlling the ultimate activity of a T cell.
By understanding this process, Dr Lillemeier said, “We might be able to encourage the immune system to be a little more sensitive in order to recognize and eliminate diseases.”
His team is now working to identify new principles that determine if T cells respond to a threat or stay quiet. They are also testing whether their findings could be applied across additional processes in T cells and other immune cells.
Because proteins have many of the same modular building blocks, Dr Lillemeier said that, in principle, any protein with structural characteristics comparable to those of ZAP-70 could be controlled by similar mechanisms. ![]()

plasma membrane upon T-cell
activation. Image courtesy
of the Salk Institute
New research suggests that, contrary to previous assumptions, T-cell activation is a fluid process that relies on a dynamic protein network at the cell surface.
Previously, scientists thought T-cell activation was a static process in which molecules assemble within a T cell.
Instead, investigators discovered that molecules rapidly come and go, and the times at which these molecules arrive and depart affect the immune response.
“This is a completely new principle for how T-cell activity is controlled—whether it ignores or responds to a threat,” said Björn Lillemeier, PhD, of the Salk Institute in La Jolla, California.
He and his colleagues described this discovery in Nature Immunology.
They noted that the protein kinase ZAP-70 plays a central role in T-cell activation. Until now, scientists assumed that a silent form of ZAP-70 floats around inside the T cell until a threat is detected. The threat recruits ZAP-70 to the cell surface and activates it.
By analyzing mutant forms of ZAP-70, Dr Lillemeier’s group discovered that, instead of ZAP-70 binding the T-cell receptor firmly, it comes in contact with the receptor sporadically. Each time this happens, ZAP-70 has to adopt an unfavorable shape that forces it back inside the cell.
This cycle continues until a second molecule, Lck, helps ZAP-70 remain with the T-cell receptor. The prolonged stay at the cell surface activates ZAP-70 and prompts the T cell to attack invaders and diseased cells.
The investigators said this study suggests the steps underlying T-cell activation are more dynamic than the less mobile modes scientists had suspected before. And the research highlights how ZAP-70 and other molecules communicate in space and time, which is crucial for controlling the ultimate activity of a T cell.
By understanding this process, Dr Lillemeier said, “We might be able to encourage the immune system to be a little more sensitive in order to recognize and eliminate diseases.”
His team is now working to identify new principles that determine if T cells respond to a threat or stay quiet. They are also testing whether their findings could be applied across additional processes in T cells and other immune cells.
Because proteins have many of the same modular building blocks, Dr Lillemeier said that, in principle, any protein with structural characteristics comparable to those of ZAP-70 could be controlled by similar mechanisms. ![]()

plasma membrane upon T-cell
activation. Image courtesy
of the Salk Institute
New research suggests that, contrary to previous assumptions, T-cell activation is a fluid process that relies on a dynamic protein network at the cell surface.
Previously, scientists thought T-cell activation was a static process in which molecules assemble within a T cell.
Instead, investigators discovered that molecules rapidly come and go, and the times at which these molecules arrive and depart affect the immune response.
“This is a completely new principle for how T-cell activity is controlled—whether it ignores or responds to a threat,” said Björn Lillemeier, PhD, of the Salk Institute in La Jolla, California.
He and his colleagues described this discovery in Nature Immunology.
They noted that the protein kinase ZAP-70 plays a central role in T-cell activation. Until now, scientists assumed that a silent form of ZAP-70 floats around inside the T cell until a threat is detected. The threat recruits ZAP-70 to the cell surface and activates it.
By analyzing mutant forms of ZAP-70, Dr Lillemeier’s group discovered that, instead of ZAP-70 binding the T-cell receptor firmly, it comes in contact with the receptor sporadically. Each time this happens, ZAP-70 has to adopt an unfavorable shape that forces it back inside the cell.
This cycle continues until a second molecule, Lck, helps ZAP-70 remain with the T-cell receptor. The prolonged stay at the cell surface activates ZAP-70 and prompts the T cell to attack invaders and diseased cells.
The investigators said this study suggests the steps underlying T-cell activation are more dynamic than the less mobile modes scientists had suspected before. And the research highlights how ZAP-70 and other molecules communicate in space and time, which is crucial for controlling the ultimate activity of a T cell.
By understanding this process, Dr Lillemeier said, “We might be able to encourage the immune system to be a little more sensitive in order to recognize and eliminate diseases.”
His team is now working to identify new principles that determine if T cells respond to a threat or stay quiet. They are also testing whether their findings could be applied across additional processes in T cells and other immune cells.
Because proteins have many of the same modular building blocks, Dr Lillemeier said that, in principle, any protein with structural characteristics comparable to those of ZAP-70 could be controlled by similar mechanisms. ![]()
How a molecule turns B cells into macrophages
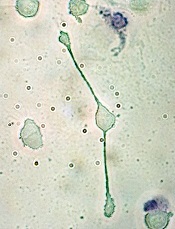
pseudopodia to engulf particles
The transcription factor C/EBPα reprograms B cells into macrophages by “short-circuiting” the cells so they re-express genes reserved for embryonic development, according to research published in Stem Cell Reports.
Over the past 28 years, researchers have shown that a number of specialized cell types can be forcibly converted into other cell types, but the science of how this change takes place is still emerging.
“For a long time, it was unclear whether forcing cell fate decisions by expressing transcription factors in the wrong cell type could teach us something about what happens normally during physiological differentiation,” said Thomas Graf, PhD, of the Center for Genomic Regulation in Barcelona, Spain.
“What we have now found is that the two processes are actually surprisingly similar.”
The researchers found that B-cell transdifferentiation takes place when C/EBPα binds to two regions of DNA that act as gene expression enhancers. One of these regions is normally active in immune cells, and the other is only turned on when macrophage precursors are ready to differentiate.
This indicates that the convergence of these two enhancer pathways can cause the B cell to act like a macrophage precursor, thus triggering the unnatural transdifferentiation.
“This has taught us a great deal about how a transcription factor can activate a new gene expression program (in our case, that of macrophages) but has left us in the dark about the other part of the equation; namely, how the factor silences the B-cell program, something that must happen if transdifferentiation is to work,” Dr Graf said. “This is one of the questions we are focusing on now.”
Dr Graf is interested in this pathway because C/EBPα-induced, B cell-to-macrophage transdifferentiation can convert both human B-cell lymphoma and leukemia cells into functional, non-cancerous macrophages.
He believes that induced transdifferentiation could become therapeutically relevant if a drug could be found that can replace the transcription factor. And understanding the mechanisms of the process could help labs that use this transdifferentiation approach to generate cells for regenerative purposes. ![]()

pseudopodia to engulf particles
The transcription factor C/EBPα reprograms B cells into macrophages by “short-circuiting” the cells so they re-express genes reserved for embryonic development, according to research published in Stem Cell Reports.
Over the past 28 years, researchers have shown that a number of specialized cell types can be forcibly converted into other cell types, but the science of how this change takes place is still emerging.
“For a long time, it was unclear whether forcing cell fate decisions by expressing transcription factors in the wrong cell type could teach us something about what happens normally during physiological differentiation,” said Thomas Graf, PhD, of the Center for Genomic Regulation in Barcelona, Spain.
“What we have now found is that the two processes are actually surprisingly similar.”
The researchers found that B-cell transdifferentiation takes place when C/EBPα binds to two regions of DNA that act as gene expression enhancers. One of these regions is normally active in immune cells, and the other is only turned on when macrophage precursors are ready to differentiate.
This indicates that the convergence of these two enhancer pathways can cause the B cell to act like a macrophage precursor, thus triggering the unnatural transdifferentiation.
“This has taught us a great deal about how a transcription factor can activate a new gene expression program (in our case, that of macrophages) but has left us in the dark about the other part of the equation; namely, how the factor silences the B-cell program, something that must happen if transdifferentiation is to work,” Dr Graf said. “This is one of the questions we are focusing on now.”
Dr Graf is interested in this pathway because C/EBPα-induced, B cell-to-macrophage transdifferentiation can convert both human B-cell lymphoma and leukemia cells into functional, non-cancerous macrophages.
He believes that induced transdifferentiation could become therapeutically relevant if a drug could be found that can replace the transcription factor. And understanding the mechanisms of the process could help labs that use this transdifferentiation approach to generate cells for regenerative purposes. ![]()

pseudopodia to engulf particles
The transcription factor C/EBPα reprograms B cells into macrophages by “short-circuiting” the cells so they re-express genes reserved for embryonic development, according to research published in Stem Cell Reports.
Over the past 28 years, researchers have shown that a number of specialized cell types can be forcibly converted into other cell types, but the science of how this change takes place is still emerging.
“For a long time, it was unclear whether forcing cell fate decisions by expressing transcription factors in the wrong cell type could teach us something about what happens normally during physiological differentiation,” said Thomas Graf, PhD, of the Center for Genomic Regulation in Barcelona, Spain.
“What we have now found is that the two processes are actually surprisingly similar.”
The researchers found that B-cell transdifferentiation takes place when C/EBPα binds to two regions of DNA that act as gene expression enhancers. One of these regions is normally active in immune cells, and the other is only turned on when macrophage precursors are ready to differentiate.
This indicates that the convergence of these two enhancer pathways can cause the B cell to act like a macrophage precursor, thus triggering the unnatural transdifferentiation.
“This has taught us a great deal about how a transcription factor can activate a new gene expression program (in our case, that of macrophages) but has left us in the dark about the other part of the equation; namely, how the factor silences the B-cell program, something that must happen if transdifferentiation is to work,” Dr Graf said. “This is one of the questions we are focusing on now.”
Dr Graf is interested in this pathway because C/EBPα-induced, B cell-to-macrophage transdifferentiation can convert both human B-cell lymphoma and leukemia cells into functional, non-cancerous macrophages.
He believes that induced transdifferentiation could become therapeutically relevant if a drug could be found that can replace the transcription factor. And understanding the mechanisms of the process could help labs that use this transdifferentiation approach to generate cells for regenerative purposes. ![]()
Microfluidics can improve epigenomic analysis, team says

his student, Zhenning Cao.
Dr Lu is holding the chip
used in the study.
Photo from Virginia Tech
A new technique enables epigenomic analysis using fewer cells than other methods involving chromatin immunoprecipitation and deep sequencing (ChIP-seq), according to researchers.
The technique, microfluidic oscillatory washing-based ChIP-seq (MOWChIP-seq), allows for genome-wide analysis of histone modifications using as few as 100
cells.
Using MOWChIP-seq, researchers uncovered new information regarding early hematopoiesis.
The team described this work in Nature Methods.
They used multilayer soft lithography to create a poly (dimethylsiloxane) device with a microfluidic chamber for ChIP. The researchers flowed magnetic beads coated with a ChIP antibody into the chamber, which formed a packed bed.
They then flowed sonicated chromatin fragments through the chamber, which were adsorbed onto the bead surface. The team said the gaps between the immunoprecipitation beads are smaller than 2 μm and facilitate rapid, high-efficiency adsorption of target chromatin fragments under the small diffusion length.
The researchers washed the beads by oscillatory washing in two different buffers to remove nonspecifically adsorbed chromatin fragments. Then, they flowed the beads out of the chamber and collected them for off-chip processing.
“The use of a packed bed of beads for ChIP allowed us to collect the chromatin fragments with a very high efficiency,” said study author Chang Lu, PhD, of Virginia Tech in Blacksburg.
“At the same time, effective washing for removing undesired molecules and debris guarantees the purity of the collected molecules. These two factors constitute a successful strategy for epigenomic analysis with extremely high sensitivity.”
In addition, the entire MOWChIP-seq process takes about 90 minutes.
To test MOWChIP-seq, Dr Lu and his colleagues used the technique to study the epigenomes of hematopoietic stem and progenitor cells (HSPCs) isolated from the fetal liver of a mouse.
“Little is known about the dynamics of the epigenome during embryonic hematopoiesis, largely due to the difficulty in isolating sufficient quantities of these cells from developing embryos,” said study author Kai Tan, PhD, of the University of Iowa in Iowa City. “This technology is the perfect tool for tackling this problem.”
MOWChIP-seq revealed new enhancers and super enhancers in the HSPCs.
By comparing all of the enhancers they identified to an enhancer catalog covering 16 blood cell types, the researchers found that 2561 (58%) of the enhancers they found were unique to fetal liver HSPCs. They said this suggests enhancer activity is highly dynamic during early hematopoiesis.
Now, the researchers plan to use MOWChIP-seq to study other epigenomic changes involved in inflammation and cancer.
“Our technology paves the way for studies of epigenomes with extremely low numbers of cells from animals and from patients,” Dr Lu said.
Virginia Tech Intellectual Properties has filed a utility patent on MOWChIP-seq on behalf of Dr Lu. ![]()

his student, Zhenning Cao.
Dr Lu is holding the chip
used in the study.
Photo from Virginia Tech
A new technique enables epigenomic analysis using fewer cells than other methods involving chromatin immunoprecipitation and deep sequencing (ChIP-seq), according to researchers.
The technique, microfluidic oscillatory washing-based ChIP-seq (MOWChIP-seq), allows for genome-wide analysis of histone modifications using as few as 100
cells.
Using MOWChIP-seq, researchers uncovered new information regarding early hematopoiesis.
The team described this work in Nature Methods.
They used multilayer soft lithography to create a poly (dimethylsiloxane) device with a microfluidic chamber for ChIP. The researchers flowed magnetic beads coated with a ChIP antibody into the chamber, which formed a packed bed.
They then flowed sonicated chromatin fragments through the chamber, which were adsorbed onto the bead surface. The team said the gaps between the immunoprecipitation beads are smaller than 2 μm and facilitate rapid, high-efficiency adsorption of target chromatin fragments under the small diffusion length.
The researchers washed the beads by oscillatory washing in two different buffers to remove nonspecifically adsorbed chromatin fragments. Then, they flowed the beads out of the chamber and collected them for off-chip processing.
“The use of a packed bed of beads for ChIP allowed us to collect the chromatin fragments with a very high efficiency,” said study author Chang Lu, PhD, of Virginia Tech in Blacksburg.
“At the same time, effective washing for removing undesired molecules and debris guarantees the purity of the collected molecules. These two factors constitute a successful strategy for epigenomic analysis with extremely high sensitivity.”
In addition, the entire MOWChIP-seq process takes about 90 minutes.
To test MOWChIP-seq, Dr Lu and his colleagues used the technique to study the epigenomes of hematopoietic stem and progenitor cells (HSPCs) isolated from the fetal liver of a mouse.
“Little is known about the dynamics of the epigenome during embryonic hematopoiesis, largely due to the difficulty in isolating sufficient quantities of these cells from developing embryos,” said study author Kai Tan, PhD, of the University of Iowa in Iowa City. “This technology is the perfect tool for tackling this problem.”
MOWChIP-seq revealed new enhancers and super enhancers in the HSPCs.
By comparing all of the enhancers they identified to an enhancer catalog covering 16 blood cell types, the researchers found that 2561 (58%) of the enhancers they found were unique to fetal liver HSPCs. They said this suggests enhancer activity is highly dynamic during early hematopoiesis.
Now, the researchers plan to use MOWChIP-seq to study other epigenomic changes involved in inflammation and cancer.
“Our technology paves the way for studies of epigenomes with extremely low numbers of cells from animals and from patients,” Dr Lu said.
Virginia Tech Intellectual Properties has filed a utility patent on MOWChIP-seq on behalf of Dr Lu. ![]()

his student, Zhenning Cao.
Dr Lu is holding the chip
used in the study.
Photo from Virginia Tech
A new technique enables epigenomic analysis using fewer cells than other methods involving chromatin immunoprecipitation and deep sequencing (ChIP-seq), according to researchers.
The technique, microfluidic oscillatory washing-based ChIP-seq (MOWChIP-seq), allows for genome-wide analysis of histone modifications using as few as 100
cells.
Using MOWChIP-seq, researchers uncovered new information regarding early hematopoiesis.
The team described this work in Nature Methods.
They used multilayer soft lithography to create a poly (dimethylsiloxane) device with a microfluidic chamber for ChIP. The researchers flowed magnetic beads coated with a ChIP antibody into the chamber, which formed a packed bed.
They then flowed sonicated chromatin fragments through the chamber, which were adsorbed onto the bead surface. The team said the gaps between the immunoprecipitation beads are smaller than 2 μm and facilitate rapid, high-efficiency adsorption of target chromatin fragments under the small diffusion length.
The researchers washed the beads by oscillatory washing in two different buffers to remove nonspecifically adsorbed chromatin fragments. Then, they flowed the beads out of the chamber and collected them for off-chip processing.
“The use of a packed bed of beads for ChIP allowed us to collect the chromatin fragments with a very high efficiency,” said study author Chang Lu, PhD, of Virginia Tech in Blacksburg.
“At the same time, effective washing for removing undesired molecules and debris guarantees the purity of the collected molecules. These two factors constitute a successful strategy for epigenomic analysis with extremely high sensitivity.”
In addition, the entire MOWChIP-seq process takes about 90 minutes.
To test MOWChIP-seq, Dr Lu and his colleagues used the technique to study the epigenomes of hematopoietic stem and progenitor cells (HSPCs) isolated from the fetal liver of a mouse.
“Little is known about the dynamics of the epigenome during embryonic hematopoiesis, largely due to the difficulty in isolating sufficient quantities of these cells from developing embryos,” said study author Kai Tan, PhD, of the University of Iowa in Iowa City. “This technology is the perfect tool for tackling this problem.”
MOWChIP-seq revealed new enhancers and super enhancers in the HSPCs.
By comparing all of the enhancers they identified to an enhancer catalog covering 16 blood cell types, the researchers found that 2561 (58%) of the enhancers they found were unique to fetal liver HSPCs. They said this suggests enhancer activity is highly dynamic during early hematopoiesis.
Now, the researchers plan to use MOWChIP-seq to study other epigenomic changes involved in inflammation and cancer.
“Our technology paves the way for studies of epigenomes with extremely low numbers of cells from animals and from patients,” Dr Lu said.
Virginia Tech Intellectual Properties has filed a utility patent on MOWChIP-seq on behalf of Dr Lu. ![]()
Newfound mechanism could be used to fight cancers
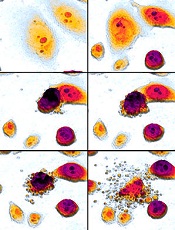
apoptosis in cancer cells
Researchers say they have identified a new mechanism by which the tumor suppressor protein p53 triggers apoptosis, and they believe this process could be harnessed to kill cancer cells.
The team discovered how p53 acts in the cytoplasm to trigger cell death by binding to and activating the BAX protein.
The process involves a shape change in one of p53’s amino acids that serves as the “switch” to activate BAX and trigger the apoptotic pathway.
The team also identified the enzyme in the cytoplasm that promotes the change that controls the “switch.”
Richard Kriwacki, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and his colleagues described these findings in Molecular Cell.
Like up to half of all proteins, p53 includes both structured and disordered regions. A disordered region is a segment that does not adopt a single shape but
remains flexible and constantly switches between different shapes until
it encounters a partner.
Dr Kriwacki and his colleagues showed that both structured and disordered regions of p53 play a role in BAX activation in the cytoplasm.
The process starts when a structured region of p53 known as the DNA-binding domain binds to BAX. That sets the stage for the unstructured region of p53 to form a second bond, which activates BAX and triggers apoptosis.
“There were no previous reports of this disordered region of p53 binding to BAX, so the finding that this region was the key to BAX activation was a total surprise,” Dr Kriwacki said.
The disordered p53 segment included the amino acid proline, which can change between two shapes, particularly in the presence of the enzyme Pin1.
Using NMR spectroscopy, the researchers showed that the proline shape change promotes p53 binding and activation of BAX.
“These results expand our understanding of the different ways p53 modulates cell behavior,” Dr Kriwacki said. “The findings also raise the possibility of killing tumor cells using small molecules to trigger BAX-dependent apoptosis.” ![]()

apoptosis in cancer cells
Researchers say they have identified a new mechanism by which the tumor suppressor protein p53 triggers apoptosis, and they believe this process could be harnessed to kill cancer cells.
The team discovered how p53 acts in the cytoplasm to trigger cell death by binding to and activating the BAX protein.
The process involves a shape change in one of p53’s amino acids that serves as the “switch” to activate BAX and trigger the apoptotic pathway.
The team also identified the enzyme in the cytoplasm that promotes the change that controls the “switch.”
Richard Kriwacki, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and his colleagues described these findings in Molecular Cell.
Like up to half of all proteins, p53 includes both structured and disordered regions. A disordered region is a segment that does not adopt a single shape but
remains flexible and constantly switches between different shapes until
it encounters a partner.
Dr Kriwacki and his colleagues showed that both structured and disordered regions of p53 play a role in BAX activation in the cytoplasm.
The process starts when a structured region of p53 known as the DNA-binding domain binds to BAX. That sets the stage for the unstructured region of p53 to form a second bond, which activates BAX and triggers apoptosis.
“There were no previous reports of this disordered region of p53 binding to BAX, so the finding that this region was the key to BAX activation was a total surprise,” Dr Kriwacki said.
The disordered p53 segment included the amino acid proline, which can change between two shapes, particularly in the presence of the enzyme Pin1.
Using NMR spectroscopy, the researchers showed that the proline shape change promotes p53 binding and activation of BAX.
“These results expand our understanding of the different ways p53 modulates cell behavior,” Dr Kriwacki said. “The findings also raise the possibility of killing tumor cells using small molecules to trigger BAX-dependent apoptosis.” ![]()

apoptosis in cancer cells
Researchers say they have identified a new mechanism by which the tumor suppressor protein p53 triggers apoptosis, and they believe this process could be harnessed to kill cancer cells.
The team discovered how p53 acts in the cytoplasm to trigger cell death by binding to and activating the BAX protein.
The process involves a shape change in one of p53’s amino acids that serves as the “switch” to activate BAX and trigger the apoptotic pathway.
The team also identified the enzyme in the cytoplasm that promotes the change that controls the “switch.”
Richard Kriwacki, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and his colleagues described these findings in Molecular Cell.
Like up to half of all proteins, p53 includes both structured and disordered regions. A disordered region is a segment that does not adopt a single shape but
remains flexible and constantly switches between different shapes until
it encounters a partner.
Dr Kriwacki and his colleagues showed that both structured and disordered regions of p53 play a role in BAX activation in the cytoplasm.
The process starts when a structured region of p53 known as the DNA-binding domain binds to BAX. That sets the stage for the unstructured region of p53 to form a second bond, which activates BAX and triggers apoptosis.
“There were no previous reports of this disordered region of p53 binding to BAX, so the finding that this region was the key to BAX activation was a total surprise,” Dr Kriwacki said.
The disordered p53 segment included the amino acid proline, which can change between two shapes, particularly in the presence of the enzyme Pin1.
Using NMR spectroscopy, the researchers showed that the proline shape change promotes p53 binding and activation of BAX.
“These results expand our understanding of the different ways p53 modulates cell behavior,” Dr Kriwacki said. “The findings also raise the possibility of killing tumor cells using small molecules to trigger BAX-dependent apoptosis.” ![]()
Delayed cancer diagnosis tied to care dissatisfaction

Photo courtesy of NIH
Patients who make 3 or more trips to the general practitioner (GP) before they are referred for cancer tests are more likely to be dissatisfied with subsequent care, according to research published in the European Journal of Cancer Care.
Researchers analyzed survey responses from nearly 60,000 cancer patients and found that about 23% had visited their GP 3 or more times before they were referred for cancer tests.
These patients were more likely than patients with 1 or 2 GP visits to report negative experiences with regard to many different aspects of their care.
“This research shows that first impressions go a long way in determining how cancer patients view their experience of cancer treatment,” said study author Georgios Lyratzopoulos, MD, of University College London in the UK.
“A negative experience of diagnosis can trigger loss of confidence in their care throughout the cancer journey. When they occur, diagnostic delays are largely due to cancer symptoms being extremely hard to distinguish from other diseases, combined with a lack of accurate and easy-to-use tests. New diagnostic tools to help doctors decide which patients need referring are vital to improve the care experience for even more cancer patients.”
Dr Lyratzopoulos and his colleagues initially analyzed survey data from 73,462 cancer patients, but 15,355 of those patients (21%) had not seen a GP prior to their cancer diagnosis.
Of the 58,107 patients who had at least 1 primary care consultation, 44,827 (77%) had seen their GP once or twice before referral for cancer tests, and 13,280 (23%) had seen the GP 3 or more times.
The researchers analyzed patients’ responses to 12 survey questions that assessed satisfaction with various aspects of their cancer care.
And results showed that patients who had seen their GP 3 or more times before referral were significantly more likely than patients with 1 to 2 GP visits to report negative experiences across 10 of the 12 aspects of care.
The 12 aspects of care and patient responses (for 3+ GP visits and 1-2 visits, respectively) were as follows:
- Dissatisfied with overall care: 16% vs 9% (adjusted* odds ratio [aOR]=1.44, P<0.001)
- Dissatisfied with the way they were informed of their cancer: 22% vs 14% (aOR=1.38, P<0.001)
- Dissatisfied with their involvement in decision-making: 32% vs 25% (aOR=1.13, P<0.001)
- Lack of confidence/trust in their doctors: 19% vs 12% (aOR=1.22, P<0.001)
- Lack of confidence/trust in their nurses: 36% vs 27% (aOR=1.22, P<0.001)
- Thought doctors/nurses deliberately withheld information: 16% vs 10% (aOR=1.27, P<0.001)
- Dissatisfied with staff support: 41% vs 27% (aOR=1.68, P<0.001)
- Dissatisfied with integration of various caregivers (GP, hospital doctors/nurses, specialist nurses, etc): 45% vs 32% (aOR=1.48, P<0.001)
- Dissatisfied with the amount of time they had to wait at appointments with their hospital doctor: 31% vs 28% (aOR=1.10, P=0.007)
- Said their GP did not receive enough information about their condition or hospital treatment: 8% vs 5% (aOR=1.36, P<0.001)
- Said they did not receive clear written information about what they should or should not do after leaving the hospital: 20% vs 14% (aOR=1.06, P=0.100)
- Said they did not receive the name of a Clinical Nurse Specialist who would be in charge of their care: 11% vs 9% (aOR=1.00, P=0.894).
“This is the first time we’ve had direct feedback from patients on such a large scale to show how the timeliness of their diagnosis colors their experience of the care they later receive,” said Sara Hiom, of Cancer Research UK, which supported this study.
“It’s another good reason to highlight the importance of diagnosing cancer as quickly as possible, not just to give patients the best chances of survival, but also to improve their experience of the care they receive throughout their cancer journey.” ![]()
*The odds ratio was adjusted for patient age, sex, ethnicity, cancer diagnosis, and response tendency (likelihood of giving a positive or negative response to questions).

Photo courtesy of NIH
Patients who make 3 or more trips to the general practitioner (GP) before they are referred for cancer tests are more likely to be dissatisfied with subsequent care, according to research published in the European Journal of Cancer Care.
Researchers analyzed survey responses from nearly 60,000 cancer patients and found that about 23% had visited their GP 3 or more times before they were referred for cancer tests.
These patients were more likely than patients with 1 or 2 GP visits to report negative experiences with regard to many different aspects of their care.
“This research shows that first impressions go a long way in determining how cancer patients view their experience of cancer treatment,” said study author Georgios Lyratzopoulos, MD, of University College London in the UK.
“A negative experience of diagnosis can trigger loss of confidence in their care throughout the cancer journey. When they occur, diagnostic delays are largely due to cancer symptoms being extremely hard to distinguish from other diseases, combined with a lack of accurate and easy-to-use tests. New diagnostic tools to help doctors decide which patients need referring are vital to improve the care experience for even more cancer patients.”
Dr Lyratzopoulos and his colleagues initially analyzed survey data from 73,462 cancer patients, but 15,355 of those patients (21%) had not seen a GP prior to their cancer diagnosis.
Of the 58,107 patients who had at least 1 primary care consultation, 44,827 (77%) had seen their GP once or twice before referral for cancer tests, and 13,280 (23%) had seen the GP 3 or more times.
The researchers analyzed patients’ responses to 12 survey questions that assessed satisfaction with various aspects of their cancer care.
And results showed that patients who had seen their GP 3 or more times before referral were significantly more likely than patients with 1 to 2 GP visits to report negative experiences across 10 of the 12 aspects of care.
The 12 aspects of care and patient responses (for 3+ GP visits and 1-2 visits, respectively) were as follows:
- Dissatisfied with overall care: 16% vs 9% (adjusted* odds ratio [aOR]=1.44, P<0.001)
- Dissatisfied with the way they were informed of their cancer: 22% vs 14% (aOR=1.38, P<0.001)
- Dissatisfied with their involvement in decision-making: 32% vs 25% (aOR=1.13, P<0.001)
- Lack of confidence/trust in their doctors: 19% vs 12% (aOR=1.22, P<0.001)
- Lack of confidence/trust in their nurses: 36% vs 27% (aOR=1.22, P<0.001)
- Thought doctors/nurses deliberately withheld information: 16% vs 10% (aOR=1.27, P<0.001)
- Dissatisfied with staff support: 41% vs 27% (aOR=1.68, P<0.001)
- Dissatisfied with integration of various caregivers (GP, hospital doctors/nurses, specialist nurses, etc): 45% vs 32% (aOR=1.48, P<0.001)
- Dissatisfied with the amount of time they had to wait at appointments with their hospital doctor: 31% vs 28% (aOR=1.10, P=0.007)
- Said their GP did not receive enough information about their condition or hospital treatment: 8% vs 5% (aOR=1.36, P<0.001)
- Said they did not receive clear written information about what they should or should not do after leaving the hospital: 20% vs 14% (aOR=1.06, P=0.100)
- Said they did not receive the name of a Clinical Nurse Specialist who would be in charge of their care: 11% vs 9% (aOR=1.00, P=0.894).
“This is the first time we’ve had direct feedback from patients on such a large scale to show how the timeliness of their diagnosis colors their experience of the care they later receive,” said Sara Hiom, of Cancer Research UK, which supported this study.
“It’s another good reason to highlight the importance of diagnosing cancer as quickly as possible, not just to give patients the best chances of survival, but also to improve their experience of the care they receive throughout their cancer journey.” ![]()
*The odds ratio was adjusted for patient age, sex, ethnicity, cancer diagnosis, and response tendency (likelihood of giving a positive or negative response to questions).

Photo courtesy of NIH
Patients who make 3 or more trips to the general practitioner (GP) before they are referred for cancer tests are more likely to be dissatisfied with subsequent care, according to research published in the European Journal of Cancer Care.
Researchers analyzed survey responses from nearly 60,000 cancer patients and found that about 23% had visited their GP 3 or more times before they were referred for cancer tests.
These patients were more likely than patients with 1 or 2 GP visits to report negative experiences with regard to many different aspects of their care.
“This research shows that first impressions go a long way in determining how cancer patients view their experience of cancer treatment,” said study author Georgios Lyratzopoulos, MD, of University College London in the UK.
“A negative experience of diagnosis can trigger loss of confidence in their care throughout the cancer journey. When they occur, diagnostic delays are largely due to cancer symptoms being extremely hard to distinguish from other diseases, combined with a lack of accurate and easy-to-use tests. New diagnostic tools to help doctors decide which patients need referring are vital to improve the care experience for even more cancer patients.”
Dr Lyratzopoulos and his colleagues initially analyzed survey data from 73,462 cancer patients, but 15,355 of those patients (21%) had not seen a GP prior to their cancer diagnosis.
Of the 58,107 patients who had at least 1 primary care consultation, 44,827 (77%) had seen their GP once or twice before referral for cancer tests, and 13,280 (23%) had seen the GP 3 or more times.
The researchers analyzed patients’ responses to 12 survey questions that assessed satisfaction with various aspects of their cancer care.
And results showed that patients who had seen their GP 3 or more times before referral were significantly more likely than patients with 1 to 2 GP visits to report negative experiences across 10 of the 12 aspects of care.
The 12 aspects of care and patient responses (for 3+ GP visits and 1-2 visits, respectively) were as follows:
- Dissatisfied with overall care: 16% vs 9% (adjusted* odds ratio [aOR]=1.44, P<0.001)
- Dissatisfied with the way they were informed of their cancer: 22% vs 14% (aOR=1.38, P<0.001)
- Dissatisfied with their involvement in decision-making: 32% vs 25% (aOR=1.13, P<0.001)
- Lack of confidence/trust in their doctors: 19% vs 12% (aOR=1.22, P<0.001)
- Lack of confidence/trust in their nurses: 36% vs 27% (aOR=1.22, P<0.001)
- Thought doctors/nurses deliberately withheld information: 16% vs 10% (aOR=1.27, P<0.001)
- Dissatisfied with staff support: 41% vs 27% (aOR=1.68, P<0.001)
- Dissatisfied with integration of various caregivers (GP, hospital doctors/nurses, specialist nurses, etc): 45% vs 32% (aOR=1.48, P<0.001)
- Dissatisfied with the amount of time they had to wait at appointments with their hospital doctor: 31% vs 28% (aOR=1.10, P=0.007)
- Said their GP did not receive enough information about their condition or hospital treatment: 8% vs 5% (aOR=1.36, P<0.001)
- Said they did not receive clear written information about what they should or should not do after leaving the hospital: 20% vs 14% (aOR=1.06, P=0.100)
- Said they did not receive the name of a Clinical Nurse Specialist who would be in charge of their care: 11% vs 9% (aOR=1.00, P=0.894).
“This is the first time we’ve had direct feedback from patients on such a large scale to show how the timeliness of their diagnosis colors their experience of the care they later receive,” said Sara Hiom, of Cancer Research UK, which supported this study.
“It’s another good reason to highlight the importance of diagnosing cancer as quickly as possible, not just to give patients the best chances of survival, but also to improve their experience of the care they receive throughout their cancer journey.” ![]()
*The odds ratio was adjusted for patient age, sex, ethnicity, cancer diagnosis, and response tendency (likelihood of giving a positive or negative response to questions).
Companies ‘underinvest’ in long-term cancer research

for a clinical trial
Photo by Esther Dyson
Pharmaceutical companies “underinvest” in long-term research to develop new anticancer drugs, according to a study published in American Economic Review.
Investigators used historical data to show that companies are more likely to develop drugs for late-stage cancers than early stage cancers or cancer prevention, and this is likely because late-stage cancer drugs can be brought to market faster.
The team found that late-stage drugs extend patient survival for shorter periods so that clinical trials for these drugs get wrapped up more quickly. This, in turn, gives drug manufacturers more time to control patented drugs in the marketplace.
“There is a pattern where we get more investment in drugs that take a short time to complete and less investment in drugs that take a longer time to complete,” said study author Heidi Williams, PhD, of the Massachusetts Institute of Technology in Cambridge.
To conduct this study, Dr Williams and her colleagues analyzed 4 decades of clinical trial data from a variety of sources, including the National Cancer Institute and the US Food and Drug Administration. The study encompassed more than 200 subcategories of cancers detected at different stages of development.
In analyzing the data, the investigators divided research and development (R&D) into 2 stages: invention (developing the basic idea for a product to the point where it is patentable) and commercialization (bringing an invented product to market).
They defined the “commercialization lag” of an R&D project as the amount of time between invention and commercialization.
The data showed that patient groups with longer commercialization lags (as proxied by longer survival times) tended to have lower levels of R&D investment than groups with shorter commercialization lags (and survival times).
The investigators also found that when surrogate endpoints (endpoints other than survival) were allowed, there were more trials and money poured into research. This supports the idea that companies are more likely to invest in drugs that will have shorter trials and take less time to develop.
Dr Williams and her colleagues used the surrogate endpoint variation to estimate improvements in cancer survival rates that would have been observed if commercialization lags were reduced. The team estimated that, among US cancer patients diagnosed in 2003, longer commercialization lags resulted in around 890,000 lost life-years.
The investigators noted that commercialization lags reduce both public and private R&D investments, but they found the commercialization lag-R&D correlation is significantly more negative for privately financed trials than publicly financed trials.
The team said that, due to either excessive discounting or the fixed patent term, private incentives decline more rapidly than public incentives, which is what gives rise to the distortion.
Based on their findings, Dr Williams and her colleagues devised 3 new policy approaches that could potentially spark the development of more drugs for early stage cancers or cancer prevention.
The first is expanded use of surrogate endpoints or more research to determine if wider use of surrogate endpoints is valid.
A second possible policy change is more public funding of R&D for anticancer drugs, since such funding is free of short-term, private-sector shareholder pressure to produce returns.
A third potential new policy would be changing the terms of drug patents, which typically run from the time of patent filing to when the drug hits the market. ![]()

for a clinical trial
Photo by Esther Dyson
Pharmaceutical companies “underinvest” in long-term research to develop new anticancer drugs, according to a study published in American Economic Review.
Investigators used historical data to show that companies are more likely to develop drugs for late-stage cancers than early stage cancers or cancer prevention, and this is likely because late-stage cancer drugs can be brought to market faster.
The team found that late-stage drugs extend patient survival for shorter periods so that clinical trials for these drugs get wrapped up more quickly. This, in turn, gives drug manufacturers more time to control patented drugs in the marketplace.
“There is a pattern where we get more investment in drugs that take a short time to complete and less investment in drugs that take a longer time to complete,” said study author Heidi Williams, PhD, of the Massachusetts Institute of Technology in Cambridge.
To conduct this study, Dr Williams and her colleagues analyzed 4 decades of clinical trial data from a variety of sources, including the National Cancer Institute and the US Food and Drug Administration. The study encompassed more than 200 subcategories of cancers detected at different stages of development.
In analyzing the data, the investigators divided research and development (R&D) into 2 stages: invention (developing the basic idea for a product to the point where it is patentable) and commercialization (bringing an invented product to market).
They defined the “commercialization lag” of an R&D project as the amount of time between invention and commercialization.
The data showed that patient groups with longer commercialization lags (as proxied by longer survival times) tended to have lower levels of R&D investment than groups with shorter commercialization lags (and survival times).
The investigators also found that when surrogate endpoints (endpoints other than survival) were allowed, there were more trials and money poured into research. This supports the idea that companies are more likely to invest in drugs that will have shorter trials and take less time to develop.
Dr Williams and her colleagues used the surrogate endpoint variation to estimate improvements in cancer survival rates that would have been observed if commercialization lags were reduced. The team estimated that, among US cancer patients diagnosed in 2003, longer commercialization lags resulted in around 890,000 lost life-years.
The investigators noted that commercialization lags reduce both public and private R&D investments, but they found the commercialization lag-R&D correlation is significantly more negative for privately financed trials than publicly financed trials.
The team said that, due to either excessive discounting or the fixed patent term, private incentives decline more rapidly than public incentives, which is what gives rise to the distortion.
Based on their findings, Dr Williams and her colleagues devised 3 new policy approaches that could potentially spark the development of more drugs for early stage cancers or cancer prevention.
The first is expanded use of surrogate endpoints or more research to determine if wider use of surrogate endpoints is valid.
A second possible policy change is more public funding of R&D for anticancer drugs, since such funding is free of short-term, private-sector shareholder pressure to produce returns.
A third potential new policy would be changing the terms of drug patents, which typically run from the time of patent filing to when the drug hits the market. ![]()

for a clinical trial
Photo by Esther Dyson
Pharmaceutical companies “underinvest” in long-term research to develop new anticancer drugs, according to a study published in American Economic Review.
Investigators used historical data to show that companies are more likely to develop drugs for late-stage cancers than early stage cancers or cancer prevention, and this is likely because late-stage cancer drugs can be brought to market faster.
The team found that late-stage drugs extend patient survival for shorter periods so that clinical trials for these drugs get wrapped up more quickly. This, in turn, gives drug manufacturers more time to control patented drugs in the marketplace.
“There is a pattern where we get more investment in drugs that take a short time to complete and less investment in drugs that take a longer time to complete,” said study author Heidi Williams, PhD, of the Massachusetts Institute of Technology in Cambridge.
To conduct this study, Dr Williams and her colleagues analyzed 4 decades of clinical trial data from a variety of sources, including the National Cancer Institute and the US Food and Drug Administration. The study encompassed more than 200 subcategories of cancers detected at different stages of development.
In analyzing the data, the investigators divided research and development (R&D) into 2 stages: invention (developing the basic idea for a product to the point where it is patentable) and commercialization (bringing an invented product to market).
They defined the “commercialization lag” of an R&D project as the amount of time between invention and commercialization.
The data showed that patient groups with longer commercialization lags (as proxied by longer survival times) tended to have lower levels of R&D investment than groups with shorter commercialization lags (and survival times).
The investigators also found that when surrogate endpoints (endpoints other than survival) were allowed, there were more trials and money poured into research. This supports the idea that companies are more likely to invest in drugs that will have shorter trials and take less time to develop.
Dr Williams and her colleagues used the surrogate endpoint variation to estimate improvements in cancer survival rates that would have been observed if commercialization lags were reduced. The team estimated that, among US cancer patients diagnosed in 2003, longer commercialization lags resulted in around 890,000 lost life-years.
The investigators noted that commercialization lags reduce both public and private R&D investments, but they found the commercialization lag-R&D correlation is significantly more negative for privately financed trials than publicly financed trials.
The team said that, due to either excessive discounting or the fixed patent term, private incentives decline more rapidly than public incentives, which is what gives rise to the distortion.
Based on their findings, Dr Williams and her colleagues devised 3 new policy approaches that could potentially spark the development of more drugs for early stage cancers or cancer prevention.
The first is expanded use of surrogate endpoints or more research to determine if wider use of surrogate endpoints is valid.
A second possible policy change is more public funding of R&D for anticancer drugs, since such funding is free of short-term, private-sector shareholder pressure to produce returns.
A third potential new policy would be changing the terms of drug patents, which typically run from the time of patent filing to when the drug hits the market.
Tool identifies optimal TKI for cancers

Photo courtesy of the
University of Colorado
Researchers say they have developed a tool that allows us to determine which tyrosine kinase inhibitor (TKI) will be most effective against a certain type of cancer.
The tool, known as the Kinase Addiction Ranker (KAR), predicts the genetic abnormalities that are driving the cancer in any population of cells and chooses the best TKI or combination of TKIs to target these abnormalities.
The researchers described the tool in Bioinformatics.
“A lot of [TKIs] inhibit a lot more than what they’re supposed to inhibit,” said study author Aik Choon Tan, PhD, of the University of Colorado Anschutz Medical Campus in Aurora.
“Maybe drug A was designed to inhibit kinase B, but it also inhibits kinase C and D as well. Our approach centers on exploiting the promiscuity of these drugs, the ‘drug spillover.’”
For each TKI, there is a signature describing the kinases each drug fully or partially inhibits. Dr Tan and his colleagues combined these kinase inhibition signatures with the results of high-throughput screening. They used the Genomics of Drug Sensitivity in Cancer database to determine which TKIs have already proven active against which cancer cell lines.
The result is KAR, which does 2 things. For any cancer cell line, the program ranks the kinases that are most important to the growth of the disease. Then, the program recommends the combination of existing TKIs that is likely to do the most good against the implicated kinases.
Dr Tan and his colleagues tested KAR using samples from 151 leukemia patients and found that, among the kinases analyzed, FLT3 had the highest variance in sensitivity to TKIs.
But EPHA5, EPHA3, and BTK were the kinases most commonly associated with drug sensitivity. They had significant associations in 72%, 58%, and 54% of the patient samples, respectively.
The researchers said the frequency of BTK dependence they observed is interesting given the fact that the BTK inhibitor ibrutinib produced favorable results in a phase 1b/2 trial of patients with chronic lymphocytic leukemia (CLL). The progression-free survival rate at 26 months was 75% in that trial.
Dr Tan and his colleagues said this was consistent with their findings, which showed that 70% of CLL patient data had a significant association between BTK inhibition and drug sensitivity.
The researchers also found that KAR could predict TKI sensitivity in 21 lung cancer cell lines. In addition, the tool was able to recommend a combination of TKIs that hindered proliferation in the lung cancer cell line H1581. KAR suggested ponatinib and the experimental anticancer agent AZD8055, and experiments showed that these drugs synergistically reduced proliferation in H1581.
KAR is available for download on the Tan lab’s website.

Photo courtesy of the
University of Colorado
Researchers say they have developed a tool that allows us to determine which tyrosine kinase inhibitor (TKI) will be most effective against a certain type of cancer.
The tool, known as the Kinase Addiction Ranker (KAR), predicts the genetic abnormalities that are driving the cancer in any population of cells and chooses the best TKI or combination of TKIs to target these abnormalities.
The researchers described the tool in Bioinformatics.
“A lot of [TKIs] inhibit a lot more than what they’re supposed to inhibit,” said study author Aik Choon Tan, PhD, of the University of Colorado Anschutz Medical Campus in Aurora.
“Maybe drug A was designed to inhibit kinase B, but it also inhibits kinase C and D as well. Our approach centers on exploiting the promiscuity of these drugs, the ‘drug spillover.’”
For each TKI, there is a signature describing the kinases each drug fully or partially inhibits. Dr Tan and his colleagues combined these kinase inhibition signatures with the results of high-throughput screening. They used the Genomics of Drug Sensitivity in Cancer database to determine which TKIs have already proven active against which cancer cell lines.
The result is KAR, which does 2 things. For any cancer cell line, the program ranks the kinases that are most important to the growth of the disease. Then, the program recommends the combination of existing TKIs that is likely to do the most good against the implicated kinases.
Dr Tan and his colleagues tested KAR using samples from 151 leukemia patients and found that, among the kinases analyzed, FLT3 had the highest variance in sensitivity to TKIs.
But EPHA5, EPHA3, and BTK were the kinases most commonly associated with drug sensitivity. They had significant associations in 72%, 58%, and 54% of the patient samples, respectively.
The researchers said the frequency of BTK dependence they observed is interesting given the fact that the BTK inhibitor ibrutinib produced favorable results in a phase 1b/2 trial of patients with chronic lymphocytic leukemia (CLL). The progression-free survival rate at 26 months was 75% in that trial.
Dr Tan and his colleagues said this was consistent with their findings, which showed that 70% of CLL patient data had a significant association between BTK inhibition and drug sensitivity.
The researchers also found that KAR could predict TKI sensitivity in 21 lung cancer cell lines. In addition, the tool was able to recommend a combination of TKIs that hindered proliferation in the lung cancer cell line H1581. KAR suggested ponatinib and the experimental anticancer agent AZD8055, and experiments showed that these drugs synergistically reduced proliferation in H1581.
KAR is available for download on the Tan lab’s website.

Photo courtesy of the
University of Colorado
Researchers say they have developed a tool that allows us to determine which tyrosine kinase inhibitor (TKI) will be most effective against a certain type of cancer.
The tool, known as the Kinase Addiction Ranker (KAR), predicts the genetic abnormalities that are driving the cancer in any population of cells and chooses the best TKI or combination of TKIs to target these abnormalities.
The researchers described the tool in Bioinformatics.
“A lot of [TKIs] inhibit a lot more than what they’re supposed to inhibit,” said study author Aik Choon Tan, PhD, of the University of Colorado Anschutz Medical Campus in Aurora.
“Maybe drug A was designed to inhibit kinase B, but it also inhibits kinase C and D as well. Our approach centers on exploiting the promiscuity of these drugs, the ‘drug spillover.’”
For each TKI, there is a signature describing the kinases each drug fully or partially inhibits. Dr Tan and his colleagues combined these kinase inhibition signatures with the results of high-throughput screening. They used the Genomics of Drug Sensitivity in Cancer database to determine which TKIs have already proven active against which cancer cell lines.
The result is KAR, which does 2 things. For any cancer cell line, the program ranks the kinases that are most important to the growth of the disease. Then, the program recommends the combination of existing TKIs that is likely to do the most good against the implicated kinases.
Dr Tan and his colleagues tested KAR using samples from 151 leukemia patients and found that, among the kinases analyzed, FLT3 had the highest variance in sensitivity to TKIs.
But EPHA5, EPHA3, and BTK were the kinases most commonly associated with drug sensitivity. They had significant associations in 72%, 58%, and 54% of the patient samples, respectively.
The researchers said the frequency of BTK dependence they observed is interesting given the fact that the BTK inhibitor ibrutinib produced favorable results in a phase 1b/2 trial of patients with chronic lymphocytic leukemia (CLL). The progression-free survival rate at 26 months was 75% in that trial.
Dr Tan and his colleagues said this was consistent with their findings, which showed that 70% of CLL patient data had a significant association between BTK inhibition and drug sensitivity.
The researchers also found that KAR could predict TKI sensitivity in 21 lung cancer cell lines. In addition, the tool was able to recommend a combination of TKIs that hindered proliferation in the lung cancer cell line H1581. KAR suggested ponatinib and the experimental anticancer agent AZD8055, and experiments showed that these drugs synergistically reduced proliferation in H1581.
KAR is available for download on the Tan lab’s website.
Drones can transport blood samples

Robert Chalmers launch drone
Photo courtesy of
Johns Hopkins Medicine
Small drones can safely transport blood samples, according to a study published in PLOS ONE.
Investigators found that 40 minutes of travel on hobby-sized drones did not affect the results of common and routine blood tests.
The team said that’s promising news for people living in rural and economically impoverished areas that lack passable roads because drones can give healthcare workers quick access to lab tests needed for diagnoses and treatments.
“Biological samples can be very sensitive and fragile,” said study author Timothy Amukele, MD, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
That sensitivity makes even the pneumatic tube systems used by many hospitals, for example, unsuitable for transporting blood for certain purposes.
Of particular concern related to the use of drones, Dr Amukele noted, is the sudden acceleration that marks the launch of the vehicle and the jostling when the drone lands on its belly.
“Such movements could have destroyed blood cells or prompted blood to coagulate, and I thought all kinds of blood tests might be affected, but our study shows they weren’t . . . ,” he said.
For the study, which Dr Amukele believes is the first rigorous examination of the impact of drone transport on biological samples, his team collected 6 blood samples from each of 56 healthy adult volunteers at The Johns Hopkins Hospital.
The samples were then driven to a flight site an hour’s drive from the hospital on days when the temperature was in the 70s. There, half of the samples were packaged for flight, with a view to protecting them for the in-flight environment and preventing leakage.
Those samples were then loaded into a hand-launched, fixed-wing drone and flown around for periods of 6 to 38 minutes. Owing to Federal Aviation Administration rules, the flights were conducted in an unpopulated area, stayed below 100 meters (328 feet) and were in the line of sight of the certified pilot.
The other samples were driven back from the drone flight field to The Johns Hopkins Hospital’s Core Laboratory, where they underwent the 33 most common laboratory tests that account for around 80% of all such tests done. A few of the tests performed were for sodium, glucose, and red blood cell count.
Comparing lab results of the flown versus nonflown blood samples, the investigators found that the flight did not have a significant impact.
Dr Amukele and his team noted that one blood test—the bicarbonate test—did yield different results for some of the flown versus nonflown samples. Dr Amukele said the team isn’t sure why, but the reason could be because the blood sat around for up to 8 hours before being tested.
There were no consistent differences between flown versus nonflown blood, he said, and it’s unknown whether the out-of-range results were due to the time lag or because of the drone transport.
“The ideal way to test that would be to fly the blood around immediately after drawing it, but neither the FAA nor Johns Hopkins would like drones flying around the hospital,” he said.
Given the successful results of this proof-of-concept study, Dr Amukele said the likely next step is a pilot study in a location in Africa where healthcare clinics are sometimes 60 or more miles away from labs.
“A drone could go 100 kilometers in 40 minutes,” he noted. “They’re less expensive than motorcycles and are not subject to traffic delays, and the technology already exists for the drone to be programmed to home to certain GPS coordinates, like a carrier pigeon.”
Drones have already been tested as carriers of medicines to clinics in remote areas, but whether and how drones will be used in flights over populated areas will depend on laws and regulations.

Robert Chalmers launch drone
Photo courtesy of
Johns Hopkins Medicine
Small drones can safely transport blood samples, according to a study published in PLOS ONE.
Investigators found that 40 minutes of travel on hobby-sized drones did not affect the results of common and routine blood tests.
The team said that’s promising news for people living in rural and economically impoverished areas that lack passable roads because drones can give healthcare workers quick access to lab tests needed for diagnoses and treatments.
“Biological samples can be very sensitive and fragile,” said study author Timothy Amukele, MD, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
That sensitivity makes even the pneumatic tube systems used by many hospitals, for example, unsuitable for transporting blood for certain purposes.
Of particular concern related to the use of drones, Dr Amukele noted, is the sudden acceleration that marks the launch of the vehicle and the jostling when the drone lands on its belly.
“Such movements could have destroyed blood cells or prompted blood to coagulate, and I thought all kinds of blood tests might be affected, but our study shows they weren’t . . . ,” he said.
For the study, which Dr Amukele believes is the first rigorous examination of the impact of drone transport on biological samples, his team collected 6 blood samples from each of 56 healthy adult volunteers at The Johns Hopkins Hospital.
The samples were then driven to a flight site an hour’s drive from the hospital on days when the temperature was in the 70s. There, half of the samples were packaged for flight, with a view to protecting them for the in-flight environment and preventing leakage.
Those samples were then loaded into a hand-launched, fixed-wing drone and flown around for periods of 6 to 38 minutes. Owing to Federal Aviation Administration rules, the flights were conducted in an unpopulated area, stayed below 100 meters (328 feet) and were in the line of sight of the certified pilot.
The other samples were driven back from the drone flight field to The Johns Hopkins Hospital’s Core Laboratory, where they underwent the 33 most common laboratory tests that account for around 80% of all such tests done. A few of the tests performed were for sodium, glucose, and red blood cell count.
Comparing lab results of the flown versus nonflown blood samples, the investigators found that the flight did not have a significant impact.
Dr Amukele and his team noted that one blood test—the bicarbonate test—did yield different results for some of the flown versus nonflown samples. Dr Amukele said the team isn’t sure why, but the reason could be because the blood sat around for up to 8 hours before being tested.
There were no consistent differences between flown versus nonflown blood, he said, and it’s unknown whether the out-of-range results were due to the time lag or because of the drone transport.
“The ideal way to test that would be to fly the blood around immediately after drawing it, but neither the FAA nor Johns Hopkins would like drones flying around the hospital,” he said.
Given the successful results of this proof-of-concept study, Dr Amukele said the likely next step is a pilot study in a location in Africa where healthcare clinics are sometimes 60 or more miles away from labs.
“A drone could go 100 kilometers in 40 minutes,” he noted. “They’re less expensive than motorcycles and are not subject to traffic delays, and the technology already exists for the drone to be programmed to home to certain GPS coordinates, like a carrier pigeon.”
Drones have already been tested as carriers of medicines to clinics in remote areas, but whether and how drones will be used in flights over populated areas will depend on laws and regulations.

Robert Chalmers launch drone
Photo courtesy of
Johns Hopkins Medicine
Small drones can safely transport blood samples, according to a study published in PLOS ONE.
Investigators found that 40 minutes of travel on hobby-sized drones did not affect the results of common and routine blood tests.
The team said that’s promising news for people living in rural and economically impoverished areas that lack passable roads because drones can give healthcare workers quick access to lab tests needed for diagnoses and treatments.
“Biological samples can be very sensitive and fragile,” said study author Timothy Amukele, MD, PhD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
That sensitivity makes even the pneumatic tube systems used by many hospitals, for example, unsuitable for transporting blood for certain purposes.
Of particular concern related to the use of drones, Dr Amukele noted, is the sudden acceleration that marks the launch of the vehicle and the jostling when the drone lands on its belly.
“Such movements could have destroyed blood cells or prompted blood to coagulate, and I thought all kinds of blood tests might be affected, but our study shows they weren’t . . . ,” he said.
For the study, which Dr Amukele believes is the first rigorous examination of the impact of drone transport on biological samples, his team collected 6 blood samples from each of 56 healthy adult volunteers at The Johns Hopkins Hospital.
The samples were then driven to a flight site an hour’s drive from the hospital on days when the temperature was in the 70s. There, half of the samples were packaged for flight, with a view to protecting them for the in-flight environment and preventing leakage.
Those samples were then loaded into a hand-launched, fixed-wing drone and flown around for periods of 6 to 38 minutes. Owing to Federal Aviation Administration rules, the flights were conducted in an unpopulated area, stayed below 100 meters (328 feet) and were in the line of sight of the certified pilot.
The other samples were driven back from the drone flight field to The Johns Hopkins Hospital’s Core Laboratory, where they underwent the 33 most common laboratory tests that account for around 80% of all such tests done. A few of the tests performed were for sodium, glucose, and red blood cell count.
Comparing lab results of the flown versus nonflown blood samples, the investigators found that the flight did not have a significant impact.
Dr Amukele and his team noted that one blood test—the bicarbonate test—did yield different results for some of the flown versus nonflown samples. Dr Amukele said the team isn’t sure why, but the reason could be because the blood sat around for up to 8 hours before being tested.
There were no consistent differences between flown versus nonflown blood, he said, and it’s unknown whether the out-of-range results were due to the time lag or because of the drone transport.
“The ideal way to test that would be to fly the blood around immediately after drawing it, but neither the FAA nor Johns Hopkins would like drones flying around the hospital,” he said.
Given the successful results of this proof-of-concept study, Dr Amukele said the likely next step is a pilot study in a location in Africa where healthcare clinics are sometimes 60 or more miles away from labs.
“A drone could go 100 kilometers in 40 minutes,” he noted. “They’re less expensive than motorcycles and are not subject to traffic delays, and the technology already exists for the drone to be programmed to home to certain GPS coordinates, like a carrier pigeon.”
Drones have already been tested as carriers of medicines to clinics in remote areas, but whether and how drones will be used in flights over populated areas will depend on laws and regulations.
Fertility preservation in young cancer patients
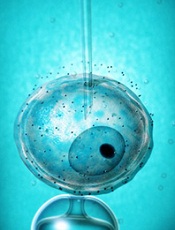
Image courtesy of NHS
Young patients with cancer, particularly females, may be uninformed about their options for preserving fertility, according to a study published in Cancer.
The research showed that males were both more likely to have discussed fertility preservation with their physicians and more likely to have taken steps to preserve fertility.
Other factors such as education and insurance status also appeared to have an impact on fertility preservation.
Margarett Shnorhavorian, MD, of the University of Washington in Seattle, and her colleagues conducted this research.
The team enlisted 459 adolescents and young adults who were diagnosed with cancer in 2007 or 2008, asking them to complete questionnaires on fertility preservation.
Eighty percent of males and 74% of females said they had been told that cancer therapy might affect their fertility. For females, multivariable analysis revealed no significant factors associated with this discussion.
However, multivariable analysis showed that males with an unknown treatment fertility risk were more likely to be uninformed of the potential risk (odds ratio [OR]= 2.73; 95% CI, 1.09-6.86), as were males who did not consult a medical oncologist (OR=2.28; 95% CI, 1.03-5.00).
Twenty-nine percent of males and 56.3% of females said they did not discuss fertility preservation with their doctors before they began cancer treatment. Males raising children younger than 18 were more likely than males without children to miss out on the discussion (OR=2.45; 95% CI, 1.24-4.85).
Males were also more likely to miss the discussion if they had a treatment fertility risk classified as “none/low” rather than “intermediate/high” (OR=3.39; 95% CI, 1.60-7.16) and if they had no insurance or government insurance rather than private insurance (OR=2.91; 95% CI, 1.41-5.97).
Males diagnosed in 2008 were less likely than those diagnosed in 2007 to miss out on the discussion (OR=0.43; 95% CI, 0.20-0.80).
Females raising children under 18 were more likely than females without children to say they did not discuss fertility preservation with their doctors (OR=3.38; 95% CI, 1.43-8.02). Females without private insurance were more likely to miss the discussion as well (OR=5.46; 95% CI, 1.59-18.72).
Females diagnosed in 2008 were less likely to miss the discussion than those diagnosed in 2007 (OR=0.36; 95% CI, 0.15-0.85).
Sixty-nine percent of males and 93.2% of females said they did not make fertility preservation arrangements. Men were more likely to lack arrangements if they were raising children younger than 18 years (OR=3.53; 95% CI, 1.63-7.65) or had less than a college degree (OR, 1.98; 95% CI, 1.00-3.97).
The researchers did not conduct a multivariable analysis for women because so few women made arrangements for fertility preservation.
“The access and health-related reasons for not making arrangements for fertility preservation reported by participants in this study further highlight the need for decreased cost, improved insurance coverage, and partnerships between cancer healthcare providers and fertility experts to develop strategies that increase awareness of fertility preservation options and decrease delays in cancer therapy as fertility preservation for adolescent and young adult cancer patients improves,” Dr Shnorhavorian concluded.

Image courtesy of NHS
Young patients with cancer, particularly females, may be uninformed about their options for preserving fertility, according to a study published in Cancer.
The research showed that males were both more likely to have discussed fertility preservation with their physicians and more likely to have taken steps to preserve fertility.
Other factors such as education and insurance status also appeared to have an impact on fertility preservation.
Margarett Shnorhavorian, MD, of the University of Washington in Seattle, and her colleagues conducted this research.
The team enlisted 459 adolescents and young adults who were diagnosed with cancer in 2007 or 2008, asking them to complete questionnaires on fertility preservation.
Eighty percent of males and 74% of females said they had been told that cancer therapy might affect their fertility. For females, multivariable analysis revealed no significant factors associated with this discussion.
However, multivariable analysis showed that males with an unknown treatment fertility risk were more likely to be uninformed of the potential risk (odds ratio [OR]= 2.73; 95% CI, 1.09-6.86), as were males who did not consult a medical oncologist (OR=2.28; 95% CI, 1.03-5.00).
Twenty-nine percent of males and 56.3% of females said they did not discuss fertility preservation with their doctors before they began cancer treatment. Males raising children younger than 18 were more likely than males without children to miss out on the discussion (OR=2.45; 95% CI, 1.24-4.85).
Males were also more likely to miss the discussion if they had a treatment fertility risk classified as “none/low” rather than “intermediate/high” (OR=3.39; 95% CI, 1.60-7.16) and if they had no insurance or government insurance rather than private insurance (OR=2.91; 95% CI, 1.41-5.97).
Males diagnosed in 2008 were less likely than those diagnosed in 2007 to miss out on the discussion (OR=0.43; 95% CI, 0.20-0.80).
Females raising children under 18 were more likely than females without children to say they did not discuss fertility preservation with their doctors (OR=3.38; 95% CI, 1.43-8.02). Females without private insurance were more likely to miss the discussion as well (OR=5.46; 95% CI, 1.59-18.72).
Females diagnosed in 2008 were less likely to miss the discussion than those diagnosed in 2007 (OR=0.36; 95% CI, 0.15-0.85).
Sixty-nine percent of males and 93.2% of females said they did not make fertility preservation arrangements. Men were more likely to lack arrangements if they were raising children younger than 18 years (OR=3.53; 95% CI, 1.63-7.65) or had less than a college degree (OR, 1.98; 95% CI, 1.00-3.97).
The researchers did not conduct a multivariable analysis for women because so few women made arrangements for fertility preservation.
“The access and health-related reasons for not making arrangements for fertility preservation reported by participants in this study further highlight the need for decreased cost, improved insurance coverage, and partnerships between cancer healthcare providers and fertility experts to develop strategies that increase awareness of fertility preservation options and decrease delays in cancer therapy as fertility preservation for adolescent and young adult cancer patients improves,” Dr Shnorhavorian concluded.

Image courtesy of NHS
Young patients with cancer, particularly females, may be uninformed about their options for preserving fertility, according to a study published in Cancer.
The research showed that males were both more likely to have discussed fertility preservation with their physicians and more likely to have taken steps to preserve fertility.
Other factors such as education and insurance status also appeared to have an impact on fertility preservation.
Margarett Shnorhavorian, MD, of the University of Washington in Seattle, and her colleagues conducted this research.
The team enlisted 459 adolescents and young adults who were diagnosed with cancer in 2007 or 2008, asking them to complete questionnaires on fertility preservation.
Eighty percent of males and 74% of females said they had been told that cancer therapy might affect their fertility. For females, multivariable analysis revealed no significant factors associated with this discussion.
However, multivariable analysis showed that males with an unknown treatment fertility risk were more likely to be uninformed of the potential risk (odds ratio [OR]= 2.73; 95% CI, 1.09-6.86), as were males who did not consult a medical oncologist (OR=2.28; 95% CI, 1.03-5.00).
Twenty-nine percent of males and 56.3% of females said they did not discuss fertility preservation with their doctors before they began cancer treatment. Males raising children younger than 18 were more likely than males without children to miss out on the discussion (OR=2.45; 95% CI, 1.24-4.85).
Males were also more likely to miss the discussion if they had a treatment fertility risk classified as “none/low” rather than “intermediate/high” (OR=3.39; 95% CI, 1.60-7.16) and if they had no insurance or government insurance rather than private insurance (OR=2.91; 95% CI, 1.41-5.97).
Males diagnosed in 2008 were less likely than those diagnosed in 2007 to miss out on the discussion (OR=0.43; 95% CI, 0.20-0.80).
Females raising children under 18 were more likely than females without children to say they did not discuss fertility preservation with their doctors (OR=3.38; 95% CI, 1.43-8.02). Females without private insurance were more likely to miss the discussion as well (OR=5.46; 95% CI, 1.59-18.72).
Females diagnosed in 2008 were less likely to miss the discussion than those diagnosed in 2007 (OR=0.36; 95% CI, 0.15-0.85).
Sixty-nine percent of males and 93.2% of females said they did not make fertility preservation arrangements. Men were more likely to lack arrangements if they were raising children younger than 18 years (OR=3.53; 95% CI, 1.63-7.65) or had less than a college degree (OR, 1.98; 95% CI, 1.00-3.97).
The researchers did not conduct a multivariable analysis for women because so few women made arrangements for fertility preservation.
“The access and health-related reasons for not making arrangements for fertility preservation reported by participants in this study further highlight the need for decreased cost, improved insurance coverage, and partnerships between cancer healthcare providers and fertility experts to develop strategies that increase awareness of fertility preservation options and decrease delays in cancer therapy as fertility preservation for adolescent and young adult cancer patients improves,” Dr Shnorhavorian concluded.