User login
FDA’s new app provides info on drug shortages

a Nokia smart phone
Photo by Halvard Lundgaard
The US Food and Drug Administration (FDA) has launched the agency’s first mobile application (app) designed to speed public access to information on drug shortages.
The app provides details regarding current drug shortages, resolved shortages, and discontinued drug products.
It works just like the FDA’s drug shortages website. App users can search for a drug by its generic name or active ingredient, or they can browse by therapeutic category.
The app can also be used to report a suspected drug shortage or supply issue to the FDA.
The app is available for free download via iTunes (for Apple devices) and the Google Play store (for Android devices). It can be found by searching “FDA Drug Shortages.”
The FDA developed the app to improve access to information about drug shortages, as part of the agency’s efforts outlined in the Strategic Plan for Preventing and Mitigating Drug Shortages.
“The FDA understands that healthcare professionals and pharmacists need real-time information about drug shortages to make treatment decisions,” said Valerie Jensen, associate director of the Drug Shortage Staff in the FDA’s Center for Drug Evaluation and Research.
“The new mobile app is an innovative tool that will offer easier and faster access to important drug shortage information.” ![]()

a Nokia smart phone
Photo by Halvard Lundgaard
The US Food and Drug Administration (FDA) has launched the agency’s first mobile application (app) designed to speed public access to information on drug shortages.
The app provides details regarding current drug shortages, resolved shortages, and discontinued drug products.
It works just like the FDA’s drug shortages website. App users can search for a drug by its generic name or active ingredient, or they can browse by therapeutic category.
The app can also be used to report a suspected drug shortage or supply issue to the FDA.
The app is available for free download via iTunes (for Apple devices) and the Google Play store (for Android devices). It can be found by searching “FDA Drug Shortages.”
The FDA developed the app to improve access to information about drug shortages, as part of the agency’s efforts outlined in the Strategic Plan for Preventing and Mitigating Drug Shortages.
“The FDA understands that healthcare professionals and pharmacists need real-time information about drug shortages to make treatment decisions,” said Valerie Jensen, associate director of the Drug Shortage Staff in the FDA’s Center for Drug Evaluation and Research.
“The new mobile app is an innovative tool that will offer easier and faster access to important drug shortage information.” ![]()

a Nokia smart phone
Photo by Halvard Lundgaard
The US Food and Drug Administration (FDA) has launched the agency’s first mobile application (app) designed to speed public access to information on drug shortages.
The app provides details regarding current drug shortages, resolved shortages, and discontinued drug products.
It works just like the FDA’s drug shortages website. App users can search for a drug by its generic name or active ingredient, or they can browse by therapeutic category.
The app can also be used to report a suspected drug shortage or supply issue to the FDA.
The app is available for free download via iTunes (for Apple devices) and the Google Play store (for Android devices). It can be found by searching “FDA Drug Shortages.”
The FDA developed the app to improve access to information about drug shortages, as part of the agency’s efforts outlined in the Strategic Plan for Preventing and Mitigating Drug Shortages.
“The FDA understands that healthcare professionals and pharmacists need real-time information about drug shortages to make treatment decisions,” said Valerie Jensen, associate director of the Drug Shortage Staff in the FDA’s Center for Drug Evaluation and Research.
“The new mobile app is an innovative tool that will offer easier and faster access to important drug shortage information.” ![]()
Parasite discovery could aid malaria treatment
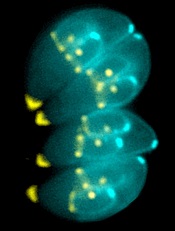
Image by Ke Hu & John Murray
Researchers say they have gained new insight into how malaria-related parasites spread inside humans and other animals.
The team discovered how the malaria relative Toxoplasma gondii manages to replicate its chromosomes up to thousands of times before spinning off into daughter cells—all while avoiding cell death.
The findings, published in PLOS Biology, may have implications for malaria treatment, according to the researchers.
Once transmitted into an animal or human, malaria-related parasites can hide out in a single cell in many different tissues, replicating thousands of times before the host’s immune system can detect them.
Then, they burst forth as daughter cells, which are unleashed in massive quantities, quickly overwhelming the body’s immune response.
The researchers found that Toxoplasma parasites pull this off thanks to the centrosome, which imposes order on the replication chaos.
“Unlike the comparatively simple centrosome present in human cells, the parasite [centrosome] has 2 distinct operating machines,” said study author Michael White, PhD, of the University of South Florida in Tampa.
“One machine controls chromosome copying, while the other machine regulates when to form daughter cell bodies. Working together, but with independent responsibilities, parasite centrosome machines can dictate the scale and timing of pathogen replication.”
This discovery of the centrosome’s function leads to a critical conclusion, Dr White said. Disrupting the centrosome machines kills the parasite. Breaking any part of the highly efficient but highly fragile replication function shuts everything down.
With these findings and the new knowledge of the parasites’ vulnerabilities, Dr White and his fellow researchers are planning to explore drug development for malaria. Whether the team is able to find an already-approved drug or must develop one from scratch, they said the drug will need to be used in conjunction with other therapies.
Dr White noted that current drugs used to treat malaria target the pathogen’s metabolism. But the goal of the new drug will be to undermine the parasite’s foundation in enough of the spreading cells to allow the immune system to fight back and not become overwhelmed. ![]()

Image by Ke Hu & John Murray
Researchers say they have gained new insight into how malaria-related parasites spread inside humans and other animals.
The team discovered how the malaria relative Toxoplasma gondii manages to replicate its chromosomes up to thousands of times before spinning off into daughter cells—all while avoiding cell death.
The findings, published in PLOS Biology, may have implications for malaria treatment, according to the researchers.
Once transmitted into an animal or human, malaria-related parasites can hide out in a single cell in many different tissues, replicating thousands of times before the host’s immune system can detect them.
Then, they burst forth as daughter cells, which are unleashed in massive quantities, quickly overwhelming the body’s immune response.
The researchers found that Toxoplasma parasites pull this off thanks to the centrosome, which imposes order on the replication chaos.
“Unlike the comparatively simple centrosome present in human cells, the parasite [centrosome] has 2 distinct operating machines,” said study author Michael White, PhD, of the University of South Florida in Tampa.
“One machine controls chromosome copying, while the other machine regulates when to form daughter cell bodies. Working together, but with independent responsibilities, parasite centrosome machines can dictate the scale and timing of pathogen replication.”
This discovery of the centrosome’s function leads to a critical conclusion, Dr White said. Disrupting the centrosome machines kills the parasite. Breaking any part of the highly efficient but highly fragile replication function shuts everything down.
With these findings and the new knowledge of the parasites’ vulnerabilities, Dr White and his fellow researchers are planning to explore drug development for malaria. Whether the team is able to find an already-approved drug or must develop one from scratch, they said the drug will need to be used in conjunction with other therapies.
Dr White noted that current drugs used to treat malaria target the pathogen’s metabolism. But the goal of the new drug will be to undermine the parasite’s foundation in enough of the spreading cells to allow the immune system to fight back and not become overwhelmed. ![]()

Image by Ke Hu & John Murray
Researchers say they have gained new insight into how malaria-related parasites spread inside humans and other animals.
The team discovered how the malaria relative Toxoplasma gondii manages to replicate its chromosomes up to thousands of times before spinning off into daughter cells—all while avoiding cell death.
The findings, published in PLOS Biology, may have implications for malaria treatment, according to the researchers.
Once transmitted into an animal or human, malaria-related parasites can hide out in a single cell in many different tissues, replicating thousands of times before the host’s immune system can detect them.
Then, they burst forth as daughter cells, which are unleashed in massive quantities, quickly overwhelming the body’s immune response.
The researchers found that Toxoplasma parasites pull this off thanks to the centrosome, which imposes order on the replication chaos.
“Unlike the comparatively simple centrosome present in human cells, the parasite [centrosome] has 2 distinct operating machines,” said study author Michael White, PhD, of the University of South Florida in Tampa.
“One machine controls chromosome copying, while the other machine regulates when to form daughter cell bodies. Working together, but with independent responsibilities, parasite centrosome machines can dictate the scale and timing of pathogen replication.”
This discovery of the centrosome’s function leads to a critical conclusion, Dr White said. Disrupting the centrosome machines kills the parasite. Breaking any part of the highly efficient but highly fragile replication function shuts everything down.
With these findings and the new knowledge of the parasites’ vulnerabilities, Dr White and his fellow researchers are planning to explore drug development for malaria. Whether the team is able to find an already-approved drug or must develop one from scratch, they said the drug will need to be used in conjunction with other therapies.
Dr White noted that current drugs used to treat malaria target the pathogen’s metabolism. But the goal of the new drug will be to undermine the parasite’s foundation in enough of the spreading cells to allow the immune system to fight back and not become overwhelmed. ![]()
Cancer care spending doesn’t correlate to lives saved
Photo by Rhoda Baer
A new analysis suggests that although US spending on cancer treatment has increased greatly in recent years, cancer mortality rates have decreased only modestly.
The study showed that care in the US often failed to prevent cancer-related deaths as well as care in Western Europe.
And when deaths were averted in the US, there was a substantial cost attached, said study author Samir Soneji, PhD, of the Norris Cotton Cancer Center in Lebanon, New Hampshire.
He and JaeWon Yang, a former undergraduate at Dartmouth College in Hanover, New Hampshire, reported these findings in Health Affairs.
The researchers compared cancer deaths and money spent on cancer care in the US and Western Europe between 1982 and 2010. They found that costs were higher in the US than in Europe for all cancers analyzed.
And compared to Western Europe, the US had 64,560 excess leukemia deaths; 164,429 excess non-Hodgkin lymphoma (NHL) deaths; 1,119,599 excess lung cancer deaths; and 39,144 excess melanoma deaths.
On the other hand, the US averted 4859 Hodgkin lymphoma deaths; 66,797 breast cancer deaths; 4354 cervical/uterine cancer deaths; 264,632 colorectal cancer deaths; 59,882 prostate cancer deaths; 621,820 stomach cancer deaths; 3372 testicular cancer deaths; and 18,320 thyroid cancer deaths.
“The greatest number of deaths averted occurred in cancers for which decreasing mortality rates were more likely to be the result of successful prevention and screening rather than advancements in treatment,” Dr Soneji noted.
He and Yang also found that the ratio of incremental cost to quality-adjusted life years (QALYs) saved in the US was $156,045 for Hodgkin lymphoma; $402,369 for breast cancer; $110,009 for colorectal cancer; $1,978,542 for prostate cancer; $4635 for stomach cancer; $222,839 for testicular cancer; and $139,681 for thyroid cancer.
But the US lost QALYs despite additional spending for leukemia, NHL, and a few other cancers. The incremental cost divided by QALYs saved was -$30,790 for leukemia; -$41,362 for NHL; -$855,019 for cervical/uterine cancer; -$18,815 for lung cancer, and -$136,592 for melanoma.
Dr Soneji described these results as, “substantially contrary to previous findings, especially for breast and prostate cancer, despite using the same data” as a previous study published in Health Affairs.
Non-replicability is a serious problem throughout academia, Dr Soneji noted. So to promote open discussion, he makes his data and procedures available to all scholars on an open-access repository called Dataverse. ![]()

Photo by Rhoda Baer
A new analysis suggests that although US spending on cancer treatment has increased greatly in recent years, cancer mortality rates have decreased only modestly.
The study showed that care in the US often failed to prevent cancer-related deaths as well as care in Western Europe.
And when deaths were averted in the US, there was a substantial cost attached, said study author Samir Soneji, PhD, of the Norris Cotton Cancer Center in Lebanon, New Hampshire.
He and JaeWon Yang, a former undergraduate at Dartmouth College in Hanover, New Hampshire, reported these findings in Health Affairs.
The researchers compared cancer deaths and money spent on cancer care in the US and Western Europe between 1982 and 2010. They found that costs were higher in the US than in Europe for all cancers analyzed.
And compared to Western Europe, the US had 64,560 excess leukemia deaths; 164,429 excess non-Hodgkin lymphoma (NHL) deaths; 1,119,599 excess lung cancer deaths; and 39,144 excess melanoma deaths.
On the other hand, the US averted 4859 Hodgkin lymphoma deaths; 66,797 breast cancer deaths; 4354 cervical/uterine cancer deaths; 264,632 colorectal cancer deaths; 59,882 prostate cancer deaths; 621,820 stomach cancer deaths; 3372 testicular cancer deaths; and 18,320 thyroid cancer deaths.
“The greatest number of deaths averted occurred in cancers for which decreasing mortality rates were more likely to be the result of successful prevention and screening rather than advancements in treatment,” Dr Soneji noted.
He and Yang also found that the ratio of incremental cost to quality-adjusted life years (QALYs) saved in the US was $156,045 for Hodgkin lymphoma; $402,369 for breast cancer; $110,009 for colorectal cancer; $1,978,542 for prostate cancer; $4635 for stomach cancer; $222,839 for testicular cancer; and $139,681 for thyroid cancer.
But the US lost QALYs despite additional spending for leukemia, NHL, and a few other cancers. The incremental cost divided by QALYs saved was -$30,790 for leukemia; -$41,362 for NHL; -$855,019 for cervical/uterine cancer; -$18,815 for lung cancer, and -$136,592 for melanoma.
Dr Soneji described these results as, “substantially contrary to previous findings, especially for breast and prostate cancer, despite using the same data” as a previous study published in Health Affairs.
Non-replicability is a serious problem throughout academia, Dr Soneji noted. So to promote open discussion, he makes his data and procedures available to all scholars on an open-access repository called Dataverse. ![]()

Photo by Rhoda Baer
A new analysis suggests that although US spending on cancer treatment has increased greatly in recent years, cancer mortality rates have decreased only modestly.
The study showed that care in the US often failed to prevent cancer-related deaths as well as care in Western Europe.
And when deaths were averted in the US, there was a substantial cost attached, said study author Samir Soneji, PhD, of the Norris Cotton Cancer Center in Lebanon, New Hampshire.
He and JaeWon Yang, a former undergraduate at Dartmouth College in Hanover, New Hampshire, reported these findings in Health Affairs.
The researchers compared cancer deaths and money spent on cancer care in the US and Western Europe between 1982 and 2010. They found that costs were higher in the US than in Europe for all cancers analyzed.
And compared to Western Europe, the US had 64,560 excess leukemia deaths; 164,429 excess non-Hodgkin lymphoma (NHL) deaths; 1,119,599 excess lung cancer deaths; and 39,144 excess melanoma deaths.
On the other hand, the US averted 4859 Hodgkin lymphoma deaths; 66,797 breast cancer deaths; 4354 cervical/uterine cancer deaths; 264,632 colorectal cancer deaths; 59,882 prostate cancer deaths; 621,820 stomach cancer deaths; 3372 testicular cancer deaths; and 18,320 thyroid cancer deaths.
“The greatest number of deaths averted occurred in cancers for which decreasing mortality rates were more likely to be the result of successful prevention and screening rather than advancements in treatment,” Dr Soneji noted.
He and Yang also found that the ratio of incremental cost to quality-adjusted life years (QALYs) saved in the US was $156,045 for Hodgkin lymphoma; $402,369 for breast cancer; $110,009 for colorectal cancer; $1,978,542 for prostate cancer; $4635 for stomach cancer; $222,839 for testicular cancer; and $139,681 for thyroid cancer.
But the US lost QALYs despite additional spending for leukemia, NHL, and a few other cancers. The incremental cost divided by QALYs saved was -$30,790 for leukemia; -$41,362 for NHL; -$855,019 for cervical/uterine cancer; -$18,815 for lung cancer, and -$136,592 for melanoma.
Dr Soneji described these results as, “substantially contrary to previous findings, especially for breast and prostate cancer, despite using the same data” as a previous study published in Health Affairs.
Non-replicability is a serious problem throughout academia, Dr Soneji noted. So to promote open discussion, he makes his data and procedures available to all scholars on an open-access repository called Dataverse. ![]()
3D-printed devices can deliver drugs in vitro
ATLANTA—Interventional radiologists say they’ve successfully used 3D printers to develop personalized medical devices that can deliver antibiotics and chemotherapy in a targeted manner in vitro.
Researchers and engineers collaborated to print catheters, stents, and filaments that were bioactive, giving these devices the ability to deliver antibiotics and chemotherapeutic medications to a targeted area in cell cultures.
Horacio R. D’Agostino, MD, of Louisiana State University Health Sciences Center in Shreveport, discussed this work at the 2015 Society of Interventional Radiology’s Annual Scientific Meeting (abstract 13).
“3D printing allows for tailor-made materials for personalized medicine,” Dr D’Agostino said. “It gives us the ability to construct devices that meet patients’ needs, from their unique anatomy to specific medicine requirements. And as tools in interventional radiology, these devices are part of treatment options that are less invasive than traditional surgery.”
Using 3D printing technology and resorbable bioplastics, Dr D’Agostino and his colleagues developed bioactive filaments, chemotherapy beads, and catheters and stents containing antibiotics or chemotherapeutic agents.
The team then tested these devices in cell cultures. They found the antibiotic-containing catheters inhibited E coli growth, and filaments carrying chemotherapeutic agents inhibited the growth of osteosarcoma cells.
“We treat a wide variety of patients and, with some patients, the current one-size-fits-all devices are not an option,” Dr D’Agostino noted. “3D printing gives us the ability to craft devices that are better suited for certain patient populations that are traditionally tough to treat, such as children and the obese, who have different anatomy. There’s limitless potential to be explored with this technology.”
The researchers were also able to print biodegradable filaments, catheters, and stents that contain antibiotics and chemotherapeutic agents. These devices might help patients avoid the need to undergo a second procedure or treatment to remove or destroy the delivery vehicle.
Dr D’Agostino said this early success with 3D-printed instruments in the lab warrants further studies, with the goal of receiving approval to use these devices in humans. He also sees an opportunity to collaborate with other medical specialties to deliver higher-quality, personalized care to all types of patients. ![]()

ATLANTA—Interventional radiologists say they’ve successfully used 3D printers to develop personalized medical devices that can deliver antibiotics and chemotherapy in a targeted manner in vitro.
Researchers and engineers collaborated to print catheters, stents, and filaments that were bioactive, giving these devices the ability to deliver antibiotics and chemotherapeutic medications to a targeted area in cell cultures.
Horacio R. D’Agostino, MD, of Louisiana State University Health Sciences Center in Shreveport, discussed this work at the 2015 Society of Interventional Radiology’s Annual Scientific Meeting (abstract 13).
“3D printing allows for tailor-made materials for personalized medicine,” Dr D’Agostino said. “It gives us the ability to construct devices that meet patients’ needs, from their unique anatomy to specific medicine requirements. And as tools in interventional radiology, these devices are part of treatment options that are less invasive than traditional surgery.”
Using 3D printing technology and resorbable bioplastics, Dr D’Agostino and his colleagues developed bioactive filaments, chemotherapy beads, and catheters and stents containing antibiotics or chemotherapeutic agents.
The team then tested these devices in cell cultures. They found the antibiotic-containing catheters inhibited E coli growth, and filaments carrying chemotherapeutic agents inhibited the growth of osteosarcoma cells.
“We treat a wide variety of patients and, with some patients, the current one-size-fits-all devices are not an option,” Dr D’Agostino noted. “3D printing gives us the ability to craft devices that are better suited for certain patient populations that are traditionally tough to treat, such as children and the obese, who have different anatomy. There’s limitless potential to be explored with this technology.”
The researchers were also able to print biodegradable filaments, catheters, and stents that contain antibiotics and chemotherapeutic agents. These devices might help patients avoid the need to undergo a second procedure or treatment to remove or destroy the delivery vehicle.
Dr D’Agostino said this early success with 3D-printed instruments in the lab warrants further studies, with the goal of receiving approval to use these devices in humans. He also sees an opportunity to collaborate with other medical specialties to deliver higher-quality, personalized care to all types of patients. ![]()

ATLANTA—Interventional radiologists say they’ve successfully used 3D printers to develop personalized medical devices that can deliver antibiotics and chemotherapy in a targeted manner in vitro.
Researchers and engineers collaborated to print catheters, stents, and filaments that were bioactive, giving these devices the ability to deliver antibiotics and chemotherapeutic medications to a targeted area in cell cultures.
Horacio R. D’Agostino, MD, of Louisiana State University Health Sciences Center in Shreveport, discussed this work at the 2015 Society of Interventional Radiology’s Annual Scientific Meeting (abstract 13).
“3D printing allows for tailor-made materials for personalized medicine,” Dr D’Agostino said. “It gives us the ability to construct devices that meet patients’ needs, from their unique anatomy to specific medicine requirements. And as tools in interventional radiology, these devices are part of treatment options that are less invasive than traditional surgery.”
Using 3D printing technology and resorbable bioplastics, Dr D’Agostino and his colleagues developed bioactive filaments, chemotherapy beads, and catheters and stents containing antibiotics or chemotherapeutic agents.
The team then tested these devices in cell cultures. They found the antibiotic-containing catheters inhibited E coli growth, and filaments carrying chemotherapeutic agents inhibited the growth of osteosarcoma cells.
“We treat a wide variety of patients and, with some patients, the current one-size-fits-all devices are not an option,” Dr D’Agostino noted. “3D printing gives us the ability to craft devices that are better suited for certain patient populations that are traditionally tough to treat, such as children and the obese, who have different anatomy. There’s limitless potential to be explored with this technology.”
The researchers were also able to print biodegradable filaments, catheters, and stents that contain antibiotics and chemotherapeutic agents. These devices might help patients avoid the need to undergo a second procedure or treatment to remove or destroy the delivery vehicle.
Dr D’Agostino said this early success with 3D-printed instruments in the lab warrants further studies, with the goal of receiving approval to use these devices in humans. He also sees an opportunity to collaborate with other medical specialties to deliver higher-quality, personalized care to all types of patients. ![]()
Findings could lead to new methods of malaria control

Photo courtesy of CDC
Sexual biology may be the key to uncovering why Anopheles mosquitoes are unique in their ability to transmit malaria to humans, according to research published in Science.
By analyzing 16 Anopheles genomes, investigators found these mosquitoes’ reproductive traits evolved along with their capacity to transmit the Plasmodium parasite.
The team believes these findings may provide a new target for malaria control, particularly in regions hardest hit by the disease.
“Our study is the first to reveal the evolutionary dynamics between the sexes that are likely responsible for shaping the ability of Anopheles mosquitoes to transmit malaria to humans,” said study author Flaminia Catteruccia, PhD, of the University of Perugia in Italy.
She and her colleagues analyzed 9 globally dispersed Anopheles species, enabling reconstruction of the evolutionary history of the mosquitoes’ reproductive traits and capacity to transmit malaria.
Results showed that 2 key male reproductive traits in Anopheles are acquired and evolved together over time. The first is transferring ejaculate as a gelatinous, rod-shaped structure called the mating plug. And the second is the ability to synthesize a steroid hormone known as 20-hydroxyecdysone (20E), which is contained in the mating plug.
The investigators also demonstrated that the evolution of these male traits drove reciprocal adaptations in females that are strongly linked to the mosquitoes’ capacity to transmit malaria.
With prior research, the team had shown that sexual transfer of 20E induces a series of dramatic changes in the female mosquito, fundamentally altering her physiology and behavior. These changes affect a female’s reproductive output, longevity, and immune response to Plasmodium parasites, all key factors in malaria transmission.
All 4 species of Anopheles mosquitoes that transfer large levels of 20E are major malaria vectors originating from Africa and India, the regions of highest malaria burden.
The investigators believe that, by identifying factors important for malaria transmission, they have paved the way for the development of compounds to specifically target those factors. Such compounds could be incorporated into existing mosquito control technologies, boosting their overall effectiveness.
The team also thinks their findings might be applicable to Dengue and West Nile virus, which are transmitted by the Aedes and Culex mosquitoes, respectively. In these species, some aspects of reproductive biology are similar to Anopheles. ![]()

Photo courtesy of CDC
Sexual biology may be the key to uncovering why Anopheles mosquitoes are unique in their ability to transmit malaria to humans, according to research published in Science.
By analyzing 16 Anopheles genomes, investigators found these mosquitoes’ reproductive traits evolved along with their capacity to transmit the Plasmodium parasite.
The team believes these findings may provide a new target for malaria control, particularly in regions hardest hit by the disease.
“Our study is the first to reveal the evolutionary dynamics between the sexes that are likely responsible for shaping the ability of Anopheles mosquitoes to transmit malaria to humans,” said study author Flaminia Catteruccia, PhD, of the University of Perugia in Italy.
She and her colleagues analyzed 9 globally dispersed Anopheles species, enabling reconstruction of the evolutionary history of the mosquitoes’ reproductive traits and capacity to transmit malaria.
Results showed that 2 key male reproductive traits in Anopheles are acquired and evolved together over time. The first is transferring ejaculate as a gelatinous, rod-shaped structure called the mating plug. And the second is the ability to synthesize a steroid hormone known as 20-hydroxyecdysone (20E), which is contained in the mating plug.
The investigators also demonstrated that the evolution of these male traits drove reciprocal adaptations in females that are strongly linked to the mosquitoes’ capacity to transmit malaria.
With prior research, the team had shown that sexual transfer of 20E induces a series of dramatic changes in the female mosquito, fundamentally altering her physiology and behavior. These changes affect a female’s reproductive output, longevity, and immune response to Plasmodium parasites, all key factors in malaria transmission.
All 4 species of Anopheles mosquitoes that transfer large levels of 20E are major malaria vectors originating from Africa and India, the regions of highest malaria burden.
The investigators believe that, by identifying factors important for malaria transmission, they have paved the way for the development of compounds to specifically target those factors. Such compounds could be incorporated into existing mosquito control technologies, boosting their overall effectiveness.
The team also thinks their findings might be applicable to Dengue and West Nile virus, which are transmitted by the Aedes and Culex mosquitoes, respectively. In these species, some aspects of reproductive biology are similar to Anopheles. ![]()

Photo courtesy of CDC
Sexual biology may be the key to uncovering why Anopheles mosquitoes are unique in their ability to transmit malaria to humans, according to research published in Science.
By analyzing 16 Anopheles genomes, investigators found these mosquitoes’ reproductive traits evolved along with their capacity to transmit the Plasmodium parasite.
The team believes these findings may provide a new target for malaria control, particularly in regions hardest hit by the disease.
“Our study is the first to reveal the evolutionary dynamics between the sexes that are likely responsible for shaping the ability of Anopheles mosquitoes to transmit malaria to humans,” said study author Flaminia Catteruccia, PhD, of the University of Perugia in Italy.
She and her colleagues analyzed 9 globally dispersed Anopheles species, enabling reconstruction of the evolutionary history of the mosquitoes’ reproductive traits and capacity to transmit malaria.
Results showed that 2 key male reproductive traits in Anopheles are acquired and evolved together over time. The first is transferring ejaculate as a gelatinous, rod-shaped structure called the mating plug. And the second is the ability to synthesize a steroid hormone known as 20-hydroxyecdysone (20E), which is contained in the mating plug.
The investigators also demonstrated that the evolution of these male traits drove reciprocal adaptations in females that are strongly linked to the mosquitoes’ capacity to transmit malaria.
With prior research, the team had shown that sexual transfer of 20E induces a series of dramatic changes in the female mosquito, fundamentally altering her physiology and behavior. These changes affect a female’s reproductive output, longevity, and immune response to Plasmodium parasites, all key factors in malaria transmission.
All 4 species of Anopheles mosquitoes that transfer large levels of 20E are major malaria vectors originating from Africa and India, the regions of highest malaria burden.
The investigators believe that, by identifying factors important for malaria transmission, they have paved the way for the development of compounds to specifically target those factors. Such compounds could be incorporated into existing mosquito control technologies, boosting their overall effectiveness.
The team also thinks their findings might be applicable to Dengue and West Nile virus, which are transmitted by the Aedes and Culex mosquitoes, respectively. In these species, some aspects of reproductive biology are similar to Anopheles. ![]()
Use of animals in research on the rise in the US

The use of animals in experimental research conducted at leading US laboratories has risen by more than 70% in recent years, according to a study published in the Journal of Medical Ethics.
The data contradict industry claims of reduced animal use and are at odds with government policies designed to curb and replace the use of animals in experiments, according to researchers.
The US is the world’s largest user of animals in experiments. And government data show declines in the use of cats, dogs, primates, rabbits, hamsters, and other larger mammals.
But the exclusion of the species most commonly used in laboratory research—mice, rats, birds, and all cold-blooded animals—from federal regulations has resulted in an absence of published data on how many of these animals are used in experiments.
To fill this gap, researchers at People for the Ethical Treatment of Animals (PETA) analyzed previously unpublished data collected by the National Institutes of Health (NIH) on the use of all vertebrate species at the top 25 institutions in receipt of NIH grants.
This includes such institutions as Harvard University, Yale University, Columbia University, the University of California-San Francisco, the University of Wisconsin-Madison, and Johns Hopkins University. Together, these institutions account for 27% of all NIH grants disbursed.
PETA’s analysis showed that the use of animals in laboratory research at these facilities rose by 72.7% between 1997 and 2012. This increase was largely driven by an increase in the use of mice. The use of other species remained mostly unchanged.
This study is the first ever to document figures on the use of mice, rats, birds, fish, reptiles, and amphibians in US laboratories. These species are not protected under the federal law governing the treatment of animals used in experiments—the Animal Welfare Act—and therefore are excluded from the law’s reporting requirements.
PETA’s study showed that these unregulated species comprise 98.8% of animals in laboratories.
The researchers said possible explanations for these results include personal and legal biases toward certain animal species. But the figures highlight a need for greater efforts to curb the use of animals in scientific research and more transparency in reporting on whether such efforts are succeeding.
A linked viewpoint article acknowledges the ongoing tensions between scientists and animal rights advocates but suggests that people on both sides of the divide do want to better understand one another.
It recommends that institutional policies be updated to better inform the public about the use of animals in scientific research, as well as opening up dialogue between a broad base of players to replace the often poorly informed and emotionally charged debate. ![]()

The use of animals in experimental research conducted at leading US laboratories has risen by more than 70% in recent years, according to a study published in the Journal of Medical Ethics.
The data contradict industry claims of reduced animal use and are at odds with government policies designed to curb and replace the use of animals in experiments, according to researchers.
The US is the world’s largest user of animals in experiments. And government data show declines in the use of cats, dogs, primates, rabbits, hamsters, and other larger mammals.
But the exclusion of the species most commonly used in laboratory research—mice, rats, birds, and all cold-blooded animals—from federal regulations has resulted in an absence of published data on how many of these animals are used in experiments.
To fill this gap, researchers at People for the Ethical Treatment of Animals (PETA) analyzed previously unpublished data collected by the National Institutes of Health (NIH) on the use of all vertebrate species at the top 25 institutions in receipt of NIH grants.
This includes such institutions as Harvard University, Yale University, Columbia University, the University of California-San Francisco, the University of Wisconsin-Madison, and Johns Hopkins University. Together, these institutions account for 27% of all NIH grants disbursed.
PETA’s analysis showed that the use of animals in laboratory research at these facilities rose by 72.7% between 1997 and 2012. This increase was largely driven by an increase in the use of mice. The use of other species remained mostly unchanged.
This study is the first ever to document figures on the use of mice, rats, birds, fish, reptiles, and amphibians in US laboratories. These species are not protected under the federal law governing the treatment of animals used in experiments—the Animal Welfare Act—and therefore are excluded from the law’s reporting requirements.
PETA’s study showed that these unregulated species comprise 98.8% of animals in laboratories.
The researchers said possible explanations for these results include personal and legal biases toward certain animal species. But the figures highlight a need for greater efforts to curb the use of animals in scientific research and more transparency in reporting on whether such efforts are succeeding.
A linked viewpoint article acknowledges the ongoing tensions between scientists and animal rights advocates but suggests that people on both sides of the divide do want to better understand one another.
It recommends that institutional policies be updated to better inform the public about the use of animals in scientific research, as well as opening up dialogue between a broad base of players to replace the often poorly informed and emotionally charged debate. ![]()

The use of animals in experimental research conducted at leading US laboratories has risen by more than 70% in recent years, according to a study published in the Journal of Medical Ethics.
The data contradict industry claims of reduced animal use and are at odds with government policies designed to curb and replace the use of animals in experiments, according to researchers.
The US is the world’s largest user of animals in experiments. And government data show declines in the use of cats, dogs, primates, rabbits, hamsters, and other larger mammals.
But the exclusion of the species most commonly used in laboratory research—mice, rats, birds, and all cold-blooded animals—from federal regulations has resulted in an absence of published data on how many of these animals are used in experiments.
To fill this gap, researchers at People for the Ethical Treatment of Animals (PETA) analyzed previously unpublished data collected by the National Institutes of Health (NIH) on the use of all vertebrate species at the top 25 institutions in receipt of NIH grants.
This includes such institutions as Harvard University, Yale University, Columbia University, the University of California-San Francisco, the University of Wisconsin-Madison, and Johns Hopkins University. Together, these institutions account for 27% of all NIH grants disbursed.
PETA’s analysis showed that the use of animals in laboratory research at these facilities rose by 72.7% between 1997 and 2012. This increase was largely driven by an increase in the use of mice. The use of other species remained mostly unchanged.
This study is the first ever to document figures on the use of mice, rats, birds, fish, reptiles, and amphibians in US laboratories. These species are not protected under the federal law governing the treatment of animals used in experiments—the Animal Welfare Act—and therefore are excluded from the law’s reporting requirements.
PETA’s study showed that these unregulated species comprise 98.8% of animals in laboratories.
The researchers said possible explanations for these results include personal and legal biases toward certain animal species. But the figures highlight a need for greater efforts to curb the use of animals in scientific research and more transparency in reporting on whether such efforts are succeeding.
A linked viewpoint article acknowledges the ongoing tensions between scientists and animal rights advocates but suggests that people on both sides of the divide do want to better understand one another.
It recommends that institutional policies be updated to better inform the public about the use of animals in scientific research, as well as opening up dialogue between a broad base of players to replace the often poorly informed and emotionally charged debate. ![]()
Team uses 3D printing to create drug carrier

Photo by Aaron Logan
Researchers have used a 3D printer to create a carrier that allows for local and sustained delivery of the immunosuppressive drug cyclosporine A (CsA) after cell transplantation.
The carrier is a combination of microspheres and hydrogel. In murine experiments, it delivered a local, sustained load of CsA in an amount that eliminated the need for additional drugs to treat immune rejection.
The researchers described these results in Cell Transplantation.
“Our objective was to show the feasibility of using a subcutaneous, 3D-printed drug delivery system to achieve local and sustained CsA release and to investigate the local immunosuppressive effects of the CsA after cell transplantation,” said study author Dong-Woo Cho, PhD, of the Pohang University of Science and Technology in Korea.
“The improved load-bearing capacity of the combined microsphere and hydrogel system, and its ability to maintain its integrity and shape during the implantation period, helped to deliver a sustained CsA release, preventing the acceleration of the secretion of cytokines related to immune rejection.”
The researchers noted that CsA improves the success rate of transplants, but systemic administration requires high doses that can have severe side effects. The benefit of a carrier is that it provides local drug delivery.
Other research groups have attempted CsA delivery via either microspheres or hydrogels, but most encountered serious problems, such as embolisms or organ damage due to migration of the microspheres from the injection site.
In addition, weak mechanical properties in some delivery systems caused premature dissolution and placed limitations on drug load quantity.
However, Dr Cho’s group said their carrier’s improved structure and load-bearing capacity allowed for sustained release of CsA at the desired site.
Their carrier is a hybrid of a CsA-poly (lactic-co-glycolic) acid microsphere-loaded hydrogel and a polymeric framework, which ensures the carrier can endure external force under physiological conditions.
In in vitro experiments with the carrier, the researchers observed decreased expression of cytokines, which are secreted by spleen cells activated by Concanavalin A and are related to immune rejection.
The team also implanted in mice drug carriers seeded with xenogeneic cells, and they observed significant suppression of T-cell-mediated rejection for 4 weeks.
The researchers believe this study could help overcome existing cell transplantation limitations caused by systemic immunosuppression. They said their carrier could be a promising solution for treating a range of diseases that require cell-based therapy. ![]()

Photo by Aaron Logan
Researchers have used a 3D printer to create a carrier that allows for local and sustained delivery of the immunosuppressive drug cyclosporine A (CsA) after cell transplantation.
The carrier is a combination of microspheres and hydrogel. In murine experiments, it delivered a local, sustained load of CsA in an amount that eliminated the need for additional drugs to treat immune rejection.
The researchers described these results in Cell Transplantation.
“Our objective was to show the feasibility of using a subcutaneous, 3D-printed drug delivery system to achieve local and sustained CsA release and to investigate the local immunosuppressive effects of the CsA after cell transplantation,” said study author Dong-Woo Cho, PhD, of the Pohang University of Science and Technology in Korea.
“The improved load-bearing capacity of the combined microsphere and hydrogel system, and its ability to maintain its integrity and shape during the implantation period, helped to deliver a sustained CsA release, preventing the acceleration of the secretion of cytokines related to immune rejection.”
The researchers noted that CsA improves the success rate of transplants, but systemic administration requires high doses that can have severe side effects. The benefit of a carrier is that it provides local drug delivery.
Other research groups have attempted CsA delivery via either microspheres or hydrogels, but most encountered serious problems, such as embolisms or organ damage due to migration of the microspheres from the injection site.
In addition, weak mechanical properties in some delivery systems caused premature dissolution and placed limitations on drug load quantity.
However, Dr Cho’s group said their carrier’s improved structure and load-bearing capacity allowed for sustained release of CsA at the desired site.
Their carrier is a hybrid of a CsA-poly (lactic-co-glycolic) acid microsphere-loaded hydrogel and a polymeric framework, which ensures the carrier can endure external force under physiological conditions.
In in vitro experiments with the carrier, the researchers observed decreased expression of cytokines, which are secreted by spleen cells activated by Concanavalin A and are related to immune rejection.
The team also implanted in mice drug carriers seeded with xenogeneic cells, and they observed significant suppression of T-cell-mediated rejection for 4 weeks.
The researchers believe this study could help overcome existing cell transplantation limitations caused by systemic immunosuppression. They said their carrier could be a promising solution for treating a range of diseases that require cell-based therapy. ![]()

Photo by Aaron Logan
Researchers have used a 3D printer to create a carrier that allows for local and sustained delivery of the immunosuppressive drug cyclosporine A (CsA) after cell transplantation.
The carrier is a combination of microspheres and hydrogel. In murine experiments, it delivered a local, sustained load of CsA in an amount that eliminated the need for additional drugs to treat immune rejection.
The researchers described these results in Cell Transplantation.
“Our objective was to show the feasibility of using a subcutaneous, 3D-printed drug delivery system to achieve local and sustained CsA release and to investigate the local immunosuppressive effects of the CsA after cell transplantation,” said study author Dong-Woo Cho, PhD, of the Pohang University of Science and Technology in Korea.
“The improved load-bearing capacity of the combined microsphere and hydrogel system, and its ability to maintain its integrity and shape during the implantation period, helped to deliver a sustained CsA release, preventing the acceleration of the secretion of cytokines related to immune rejection.”
The researchers noted that CsA improves the success rate of transplants, but systemic administration requires high doses that can have severe side effects. The benefit of a carrier is that it provides local drug delivery.
Other research groups have attempted CsA delivery via either microspheres or hydrogels, but most encountered serious problems, such as embolisms or organ damage due to migration of the microspheres from the injection site.
In addition, weak mechanical properties in some delivery systems caused premature dissolution and placed limitations on drug load quantity.
However, Dr Cho’s group said their carrier’s improved structure and load-bearing capacity allowed for sustained release of CsA at the desired site.
Their carrier is a hybrid of a CsA-poly (lactic-co-glycolic) acid microsphere-loaded hydrogel and a polymeric framework, which ensures the carrier can endure external force under physiological conditions.
In in vitro experiments with the carrier, the researchers observed decreased expression of cytokines, which are secreted by spleen cells activated by Concanavalin A and are related to immune rejection.
The team also implanted in mice drug carriers seeded with xenogeneic cells, and they observed significant suppression of T-cell-mediated rejection for 4 weeks.
The researchers believe this study could help overcome existing cell transplantation limitations caused by systemic immunosuppression. They said their carrier could be a promising solution for treating a range of diseases that require cell-based therapy.
Study provides new insights regarding HSCs, FA
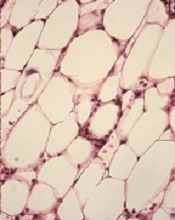
with Fanconi anemia
Image by Michael Milsom
Environmental stress is a major factor driving DNA damage in hematopoietic stem cells (HSCs), according to research published in Nature.
Investigators found that repeated exposure to such stress induces accelerated tissue aging and may cause cancer.
In examining HSCs’ response to stress, the team also gained new insight regarding Fanconi anemia.
The investigators noted that, under normal conditions, HSCs exist in a state of dormancy. They rarely divide and have very low energy demands.
“Our theory was that this state of dormancy protected hematopoietic stem cells from DNA damage and therefore protects them from premature aging,” said study author Michael Milsom, PhD, of The Heidelberg Institute for Stem Cell Technology and Experimental Medicine in Germany.
However, under conditions of stress, such as during chronic blood loss or infection, HSCs are driven into a state of rapid cell division in order to produce new blood cells and repair the damaged tissue.
“The stem cells go from a state of rest to very high activity within a short space of time, requiring them to rapidly increase their metabolic rate, synthesize new DNA, and coordinate cell division,” Dr Milsom explained. “Suddenly having to simultaneously execute these complicated functions dramatically increases the likelihood that something will go wrong.”
Indeed, the investigators’ experiments showed that the increased energy demands during stress results in elevated production of reactive metabolites that can directly damage DNA. If this happens at the same time a cell is trying to replicate its DNA, this can cause cell death or the acquisition of mutations that may cause cancer.
Normal stem cells can repair the majority of this stress-induced DNA damage, the investigators noted. However, the more HSCs are exposed to stress, the more likely they are to inefficiently repair the damage and then die or mutate.
“We believe that this model perfectly explains the gradual accumulation of DNA damage in stem cells with age and the associated reduction in the ability of a tissue to maintain and repair itself as you get older,” Dr Milsom said.
He and his colleagues also examined how this stress response impacts a mouse model of Fanconi anemia. These mice have the same DNA repair defect found in humans with the disease, but the mice never spontaneously develop the bone marrow failure observed in nearly all patients.
“We felt that stress-induced DNA damage was the missing ingredient that was required to cause hematopoietic stem cell depletion in these mice,” Dr Milsom said.
When mice with Fanconi anemia were exposed to stimulation mimicking a prolonged viral infection, they were unable to efficiently repair the resulting DNA damage, and their stem cells failed.
In the same space of time that normal mice showed a gradual decline in HSC numbers, the stem cells in Fanconi anemia mice were almost completely depleted, resulting in bone marrow failure and an inadequate production of blood cells to sustain life.
“This perfectly recapitulates what happens to Fanconi anemia patients and now gives us an opportunity to understand how this disease works and how we might better treat it,” Dr Milsom said.

with Fanconi anemia
Image by Michael Milsom
Environmental stress is a major factor driving DNA damage in hematopoietic stem cells (HSCs), according to research published in Nature.
Investigators found that repeated exposure to such stress induces accelerated tissue aging and may cause cancer.
In examining HSCs’ response to stress, the team also gained new insight regarding Fanconi anemia.
The investigators noted that, under normal conditions, HSCs exist in a state of dormancy. They rarely divide and have very low energy demands.
“Our theory was that this state of dormancy protected hematopoietic stem cells from DNA damage and therefore protects them from premature aging,” said study author Michael Milsom, PhD, of The Heidelberg Institute for Stem Cell Technology and Experimental Medicine in Germany.
However, under conditions of stress, such as during chronic blood loss or infection, HSCs are driven into a state of rapid cell division in order to produce new blood cells and repair the damaged tissue.
“The stem cells go from a state of rest to very high activity within a short space of time, requiring them to rapidly increase their metabolic rate, synthesize new DNA, and coordinate cell division,” Dr Milsom explained. “Suddenly having to simultaneously execute these complicated functions dramatically increases the likelihood that something will go wrong.”
Indeed, the investigators’ experiments showed that the increased energy demands during stress results in elevated production of reactive metabolites that can directly damage DNA. If this happens at the same time a cell is trying to replicate its DNA, this can cause cell death or the acquisition of mutations that may cause cancer.
Normal stem cells can repair the majority of this stress-induced DNA damage, the investigators noted. However, the more HSCs are exposed to stress, the more likely they are to inefficiently repair the damage and then die or mutate.
“We believe that this model perfectly explains the gradual accumulation of DNA damage in stem cells with age and the associated reduction in the ability of a tissue to maintain and repair itself as you get older,” Dr Milsom said.
He and his colleagues also examined how this stress response impacts a mouse model of Fanconi anemia. These mice have the same DNA repair defect found in humans with the disease, but the mice never spontaneously develop the bone marrow failure observed in nearly all patients.
“We felt that stress-induced DNA damage was the missing ingredient that was required to cause hematopoietic stem cell depletion in these mice,” Dr Milsom said.
When mice with Fanconi anemia were exposed to stimulation mimicking a prolonged viral infection, they were unable to efficiently repair the resulting DNA damage, and their stem cells failed.
In the same space of time that normal mice showed a gradual decline in HSC numbers, the stem cells in Fanconi anemia mice were almost completely depleted, resulting in bone marrow failure and an inadequate production of blood cells to sustain life.
“This perfectly recapitulates what happens to Fanconi anemia patients and now gives us an opportunity to understand how this disease works and how we might better treat it,” Dr Milsom said.

with Fanconi anemia
Image by Michael Milsom
Environmental stress is a major factor driving DNA damage in hematopoietic stem cells (HSCs), according to research published in Nature.
Investigators found that repeated exposure to such stress induces accelerated tissue aging and may cause cancer.
In examining HSCs’ response to stress, the team also gained new insight regarding Fanconi anemia.
The investigators noted that, under normal conditions, HSCs exist in a state of dormancy. They rarely divide and have very low energy demands.
“Our theory was that this state of dormancy protected hematopoietic stem cells from DNA damage and therefore protects them from premature aging,” said study author Michael Milsom, PhD, of The Heidelberg Institute for Stem Cell Technology and Experimental Medicine in Germany.
However, under conditions of stress, such as during chronic blood loss or infection, HSCs are driven into a state of rapid cell division in order to produce new blood cells and repair the damaged tissue.
“The stem cells go from a state of rest to very high activity within a short space of time, requiring them to rapidly increase their metabolic rate, synthesize new DNA, and coordinate cell division,” Dr Milsom explained. “Suddenly having to simultaneously execute these complicated functions dramatically increases the likelihood that something will go wrong.”
Indeed, the investigators’ experiments showed that the increased energy demands during stress results in elevated production of reactive metabolites that can directly damage DNA. If this happens at the same time a cell is trying to replicate its DNA, this can cause cell death or the acquisition of mutations that may cause cancer.
Normal stem cells can repair the majority of this stress-induced DNA damage, the investigators noted. However, the more HSCs are exposed to stress, the more likely they are to inefficiently repair the damage and then die or mutate.
“We believe that this model perfectly explains the gradual accumulation of DNA damage in stem cells with age and the associated reduction in the ability of a tissue to maintain and repair itself as you get older,” Dr Milsom said.
He and his colleagues also examined how this stress response impacts a mouse model of Fanconi anemia. These mice have the same DNA repair defect found in humans with the disease, but the mice never spontaneously develop the bone marrow failure observed in nearly all patients.
“We felt that stress-induced DNA damage was the missing ingredient that was required to cause hematopoietic stem cell depletion in these mice,” Dr Milsom said.
When mice with Fanconi anemia were exposed to stimulation mimicking a prolonged viral infection, they were unable to efficiently repair the resulting DNA damage, and their stem cells failed.
In the same space of time that normal mice showed a gradual decline in HSC numbers, the stem cells in Fanconi anemia mice were almost completely depleted, resulting in bone marrow failure and an inadequate production of blood cells to sustain life.
“This perfectly recapitulates what happens to Fanconi anemia patients and now gives us an opportunity to understand how this disease works and how we might better treat it,” Dr Milsom said.
Artemisinin-resistant malaria found across Myanmar
Photo by James Gathany
Resistance to the antimalarial drug artemisinin is present in Myanmar and has reached within 25 km of the Indian border, according to research published in The Lancet Infectious Diseases.
Researchers believe the spread of artemisinin-resistant malaria parasites into neighboring India would pose a serious threat to the global control and eradication of malaria.
And if drug resistance continues to spread, millions of lives could be at risk.
Kyaw Myo Tun, MD, of the Myanmar Oxford Clinical Research Unit in Yangon, Myanmar, and colleagues uncovered artemisinin resistance by analyzing parasite samples collected at 55 malaria treatment centers across Myanmar.
The group set out to determine if the samples carried mutations in specific regions of the parasite’s kelch gene (K13)—a known genetic marker of artemisinin resistance. And they confirmed the existance of resistant parasites in Homalin, in the Sagaing Region, which is located only 25 km from the Indian border.
“Myanmar is considered the frontline in the battle against artemisinin resistance, as it forms a gateway for resistance to spread to the rest of the world,” said Charles Woodrow, MD, of the University of Oxford in the UK.
“With artemisinins, we are in the unusual position of having molecular markers for resistance before resistance has spread globally. The more we understand about the current situation in the border regions, the better prepared we are to adapt and implement strategies to overcome the spread of further drug resistance.”
The researchers obtained the DNA sequences of 940 samples of Plasmodium falciparum malaria parasites from across Myanmar and neighboring border regions in Thailand and Bangladesh between 2013 and 2014. Of those 940 samples, 371 (39%) carried a resistance-conferring K13 mutation.
“We were able to gather patient samples rapidly across Myanmar, sometimes using discarded malaria blood diagnostic tests, and then test these immediately for the K13 marker, and so generate real-time information on the spread of resistance” said Mallika Imwong, PhD, of Mahidol University in Bangkok, Thailand.
Using this information, the researchers developed maps to display the predicted extent of artemisinin resistance determined by the prevalence of K13 mutations. The maps suggested the overall prevalence of K13 mutations was greater than 10% in large areas of the east and north of Myanmar, including areas close to the border with India.
“The identification of the K13 markers of resistance has transformed our ability to monitor the spread and emergence of artemisinin resistance,” said Philippe Guerin, MD, of the Worldwide Antimalarial Resistance Network in Oxford, UK.
“However, this study highlights that the pace at which artemisinin resistance is spreading or emerging is alarming. We need a more vigorous international effort to address this issue in border regions.”

Photo by James Gathany
Resistance to the antimalarial drug artemisinin is present in Myanmar and has reached within 25 km of the Indian border, according to research published in The Lancet Infectious Diseases.
Researchers believe the spread of artemisinin-resistant malaria parasites into neighboring India would pose a serious threat to the global control and eradication of malaria.
And if drug resistance continues to spread, millions of lives could be at risk.
Kyaw Myo Tun, MD, of the Myanmar Oxford Clinical Research Unit in Yangon, Myanmar, and colleagues uncovered artemisinin resistance by analyzing parasite samples collected at 55 malaria treatment centers across Myanmar.
The group set out to determine if the samples carried mutations in specific regions of the parasite’s kelch gene (K13)—a known genetic marker of artemisinin resistance. And they confirmed the existance of resistant parasites in Homalin, in the Sagaing Region, which is located only 25 km from the Indian border.
“Myanmar is considered the frontline in the battle against artemisinin resistance, as it forms a gateway for resistance to spread to the rest of the world,” said Charles Woodrow, MD, of the University of Oxford in the UK.
“With artemisinins, we are in the unusual position of having molecular markers for resistance before resistance has spread globally. The more we understand about the current situation in the border regions, the better prepared we are to adapt and implement strategies to overcome the spread of further drug resistance.”
The researchers obtained the DNA sequences of 940 samples of Plasmodium falciparum malaria parasites from across Myanmar and neighboring border regions in Thailand and Bangladesh between 2013 and 2014. Of those 940 samples, 371 (39%) carried a resistance-conferring K13 mutation.
“We were able to gather patient samples rapidly across Myanmar, sometimes using discarded malaria blood diagnostic tests, and then test these immediately for the K13 marker, and so generate real-time information on the spread of resistance” said Mallika Imwong, PhD, of Mahidol University in Bangkok, Thailand.
Using this information, the researchers developed maps to display the predicted extent of artemisinin resistance determined by the prevalence of K13 mutations. The maps suggested the overall prevalence of K13 mutations was greater than 10% in large areas of the east and north of Myanmar, including areas close to the border with India.
“The identification of the K13 markers of resistance has transformed our ability to monitor the spread and emergence of artemisinin resistance,” said Philippe Guerin, MD, of the Worldwide Antimalarial Resistance Network in Oxford, UK.
“However, this study highlights that the pace at which artemisinin resistance is spreading or emerging is alarming. We need a more vigorous international effort to address this issue in border regions.”

Photo by James Gathany
Resistance to the antimalarial drug artemisinin is present in Myanmar and has reached within 25 km of the Indian border, according to research published in The Lancet Infectious Diseases.
Researchers believe the spread of artemisinin-resistant malaria parasites into neighboring India would pose a serious threat to the global control and eradication of malaria.
And if drug resistance continues to spread, millions of lives could be at risk.
Kyaw Myo Tun, MD, of the Myanmar Oxford Clinical Research Unit in Yangon, Myanmar, and colleagues uncovered artemisinin resistance by analyzing parasite samples collected at 55 malaria treatment centers across Myanmar.
The group set out to determine if the samples carried mutations in specific regions of the parasite’s kelch gene (K13)—a known genetic marker of artemisinin resistance. And they confirmed the existance of resistant parasites in Homalin, in the Sagaing Region, which is located only 25 km from the Indian border.
“Myanmar is considered the frontline in the battle against artemisinin resistance, as it forms a gateway for resistance to spread to the rest of the world,” said Charles Woodrow, MD, of the University of Oxford in the UK.
“With artemisinins, we are in the unusual position of having molecular markers for resistance before resistance has spread globally. The more we understand about the current situation in the border regions, the better prepared we are to adapt and implement strategies to overcome the spread of further drug resistance.”
The researchers obtained the DNA sequences of 940 samples of Plasmodium falciparum malaria parasites from across Myanmar and neighboring border regions in Thailand and Bangladesh between 2013 and 2014. Of those 940 samples, 371 (39%) carried a resistance-conferring K13 mutation.
“We were able to gather patient samples rapidly across Myanmar, sometimes using discarded malaria blood diagnostic tests, and then test these immediately for the K13 marker, and so generate real-time information on the spread of resistance” said Mallika Imwong, PhD, of Mahidol University in Bangkok, Thailand.
Using this information, the researchers developed maps to display the predicted extent of artemisinin resistance determined by the prevalence of K13 mutations. The maps suggested the overall prevalence of K13 mutations was greater than 10% in large areas of the east and north of Myanmar, including areas close to the border with India.
“The identification of the K13 markers of resistance has transformed our ability to monitor the spread and emergence of artemisinin resistance,” said Philippe Guerin, MD, of the Worldwide Antimalarial Resistance Network in Oxford, UK.
“However, this study highlights that the pace at which artemisinin resistance is spreading or emerging is alarming. We need a more vigorous international effort to address this issue in border regions.”
Evolutionary findings may aid cancer drug development

Photo by Darren Baker
By tracking the evolution of Abl and Src, investigators have made discoveries that may aid the design of highly specific cancer drugs.
Abl and Src are 2 nearly identical protein kinases with a predilection to cause cancer in humans, mainly chronic myeloid leukemia and colon cancer.
The proteins are separated by 146 amino acids and one big difference: Abl is susceptible to treatment with the tyrosine kinase inhibitor imatinib (Gleevec), but Src is not.
Dorothee Kern, PhD, of Brandeis University in Waltham, Massachusetts, and her colleagues traced the journey of these 2 proteins over 1 billion years of evolution, pinpointing the exact evolutionary shifts that caused imatinib to bind well with one protein and poorly with the other.
This new approach to researching enzymes and their binding sites may have a major impact on the development of cancer drugs, the investigators said.
They published their findings in Science.
To determine why imatinib binds with Abl but not Src, Dr Kern and her colleagues turned back the evolutionary clock 1 billion years.
This revealed Abl and Src’s common ancestor, a primitive protein in yeast the team dubbed “ANC-AS.” They mapped out the family tree, searching for changes in amino acids and molecular mechanisms.
“Src and Abl differ by 146 amino acids, and we were looking for the handful that dictate Gleevec specificity,” Dr Kern said. “It was like finding a needle in a haystack and could only be done by our evolutionary approach.”
As ANC-AS evolved in more complex organisms, it began to specialize and branch into proteins with different regulation, roles, and catalysis processes—creating Abl and Src.
By following this progression, while testing the proteins’ affinity to imatinib along the way, the investigators were able to whittle down the 146 different amino acids to 15 that are responsible for imatinib specificity.
These 15 amino acids play a role in Abl’s conformational equilibrium—a process in which the protein transitions between 2 structures. The main difference between Abl and Src, when it comes to binding with imatinib, is the relative times the proteins spend in each configuration, resulting in a major difference in their binding energies.
By understanding how and why imatinib works on Abl—and doesn’t work on Src—scientists have a jumping off point to design other drugs with a high affinity and specificity, and a strong binding on cancerous proteins.
“Understanding the molecular basis for Gleevec specificity has opened the door wider to designing good drugs,” Dr Kern said. “Our results pave the way for a different approach to rational drug design.”

Photo by Darren Baker
By tracking the evolution of Abl and Src, investigators have made discoveries that may aid the design of highly specific cancer drugs.
Abl and Src are 2 nearly identical protein kinases with a predilection to cause cancer in humans, mainly chronic myeloid leukemia and colon cancer.
The proteins are separated by 146 amino acids and one big difference: Abl is susceptible to treatment with the tyrosine kinase inhibitor imatinib (Gleevec), but Src is not.
Dorothee Kern, PhD, of Brandeis University in Waltham, Massachusetts, and her colleagues traced the journey of these 2 proteins over 1 billion years of evolution, pinpointing the exact evolutionary shifts that caused imatinib to bind well with one protein and poorly with the other.
This new approach to researching enzymes and their binding sites may have a major impact on the development of cancer drugs, the investigators said.
They published their findings in Science.
To determine why imatinib binds with Abl but not Src, Dr Kern and her colleagues turned back the evolutionary clock 1 billion years.
This revealed Abl and Src’s common ancestor, a primitive protein in yeast the team dubbed “ANC-AS.” They mapped out the family tree, searching for changes in amino acids and molecular mechanisms.
“Src and Abl differ by 146 amino acids, and we were looking for the handful that dictate Gleevec specificity,” Dr Kern said. “It was like finding a needle in a haystack and could only be done by our evolutionary approach.”
As ANC-AS evolved in more complex organisms, it began to specialize and branch into proteins with different regulation, roles, and catalysis processes—creating Abl and Src.
By following this progression, while testing the proteins’ affinity to imatinib along the way, the investigators were able to whittle down the 146 different amino acids to 15 that are responsible for imatinib specificity.
These 15 amino acids play a role in Abl’s conformational equilibrium—a process in which the protein transitions between 2 structures. The main difference between Abl and Src, when it comes to binding with imatinib, is the relative times the proteins spend in each configuration, resulting in a major difference in their binding energies.
By understanding how and why imatinib works on Abl—and doesn’t work on Src—scientists have a jumping off point to design other drugs with a high affinity and specificity, and a strong binding on cancerous proteins.
“Understanding the molecular basis for Gleevec specificity has opened the door wider to designing good drugs,” Dr Kern said. “Our results pave the way for a different approach to rational drug design.”

Photo by Darren Baker
By tracking the evolution of Abl and Src, investigators have made discoveries that may aid the design of highly specific cancer drugs.
Abl and Src are 2 nearly identical protein kinases with a predilection to cause cancer in humans, mainly chronic myeloid leukemia and colon cancer.
The proteins are separated by 146 amino acids and one big difference: Abl is susceptible to treatment with the tyrosine kinase inhibitor imatinib (Gleevec), but Src is not.
Dorothee Kern, PhD, of Brandeis University in Waltham, Massachusetts, and her colleagues traced the journey of these 2 proteins over 1 billion years of evolution, pinpointing the exact evolutionary shifts that caused imatinib to bind well with one protein and poorly with the other.
This new approach to researching enzymes and their binding sites may have a major impact on the development of cancer drugs, the investigators said.
They published their findings in Science.
To determine why imatinib binds with Abl but not Src, Dr Kern and her colleagues turned back the evolutionary clock 1 billion years.
This revealed Abl and Src’s common ancestor, a primitive protein in yeast the team dubbed “ANC-AS.” They mapped out the family tree, searching for changes in amino acids and molecular mechanisms.
“Src and Abl differ by 146 amino acids, and we were looking for the handful that dictate Gleevec specificity,” Dr Kern said. “It was like finding a needle in a haystack and could only be done by our evolutionary approach.”
As ANC-AS evolved in more complex organisms, it began to specialize and branch into proteins with different regulation, roles, and catalysis processes—creating Abl and Src.
By following this progression, while testing the proteins’ affinity to imatinib along the way, the investigators were able to whittle down the 146 different amino acids to 15 that are responsible for imatinib specificity.
These 15 amino acids play a role in Abl’s conformational equilibrium—a process in which the protein transitions between 2 structures. The main difference between Abl and Src, when it comes to binding with imatinib, is the relative times the proteins spend in each configuration, resulting in a major difference in their binding energies.
By understanding how and why imatinib works on Abl—and doesn’t work on Src—scientists have a jumping off point to design other drugs with a high affinity and specificity, and a strong binding on cancerous proteins.
“Understanding the molecular basis for Gleevec specificity has opened the door wider to designing good drugs,” Dr Kern said. “Our results pave the way for a different approach to rational drug design.”