User login
VA SHIELD: A Biorepository for Veterans and the Nation
The Veterans Health Administration (VHA) clinicians, clinician-investigators, and investigators perform basic and translational research for the benefit of our nation and are widely recognized for treating patients and discovering cures.1,2 In May 2020, the US Department of Veterans Affairs (VA) launched the VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). The goal of this novel enterprise was to assemble a comprehensive specimen and data repository for emerging life-threatening diseases and to address future challenges. VA SHIELD was specifically charged with creating a biorepository to advance research, improve diagnostic and therapeutic capabilities, and develop strategies for immediate deployment to VA clinical environments. One main objective of VA SHIELD is to harness the clinical and scientific strengths of the VA in order to create a more cohesive collaboration between preexisting clinical research efforts within the VA.
ANATOMY OF VA SHIELD
The charge and scope of VA SHIELD is unique.3 As an entity, this program leverages the strengths of the diverse VHA network, has a broad potential impact on national health care, is positioned to respond rapidly to national and international health-related events, and substantially contributes to clinical research and development. In addition, VA SHIELD upholds VA’s Fourth Mission, which is to contribute to national emergencies and support emergency management, public health, safety, and homeland security efforts.
VA SHIELD is part of the VA Office of Research and Development (ORD). The coordinating center (CC), headquartered in Cleveland, Ohio, is the central operational partner, leading VA SHIELD and interacting with other important VA programs, including laboratory, clinical science, rehabilitation, and health services. The VA SHIELD CC oversees all aspects of operations, including biospecimen collection, creating and enforcing of standard operating procedures, ensuring the quality of the samples, processing research applications, distribution of samples, financing, and progress reports. The CC also initiates and maintains interagency collaborations, convenes stakeholders, and develops strategic plans to address emerging diseases.
The VA SHIELD Executive Steering Committee (ESC) is composed of infectious disease, biorepository, and public health specialists. The ESC provides scientific and programmatic direction to the CC, including operational activities and guidance regarding biorepository priorities and scientific agenda, and measuring and reporting on VA SHIELD accomplishments.
The primary function of the Programmatic and Scientific Review Board (PSRB) is to evaluate incoming research proposals for specimen and data use for feasibility and make recommendations to the VA SHIELD CC. The PSRB evaluates and ensures that data and specimen use align with VA SHIELD ethical, clinical, and scientific objectives.
VA SHIELD IN PRACTICE
VA SHIELD consisted of 11 specimen collection sites (Atlanta, GA; Boise, ID; Bronx, NY; Cincinnati, OH; Cleveland, OH; Durham, NC; Houston, TX; Los Angeles, CA; Mountain Home, TN; Palo Alto, CA; and Tucson, AZ), a data processing center in Boston, MA, and 2 central biorepositories in Palo Alto, CA, and Tucson, AZ. Information flow is a coordinated process among specimen collection sites, data processing centers, and the biorepositories. Initially, each local collection site identifies residual specimens that would have been discarded after clinical laboratory testing. These samples currently account for the majority of biological material within VA SHIELD via a novel collection protocol known as “Sweep,” which allows residual clinical discarded samples as well as samples from patients with new emerging infectious and noninfectious diseases of concern to be collected at the time of first emergence and submitted to VA SHIELD during the course of routine veteran health care.3 These clinical discarded samples are de-identified and transferred from the clinical laboratory to VA SHIELD. The VA Central Institutional Review Board (cIRB) has approved the use of these samples as nonhuman subject research. Biological samples are collected, processed, aliquoted, shipped to, and stored at the central biorepository sites.
The Umbrella amendment to Sweep that has been approved also by the VA cIRB, will allow VA SHIELD sites to prospectively consent veterans and collect biospecimens and additional clinical and self-reported information. The implementation of Umbrella could significantly enhance collection and research. Although Sweep is a onetime collection of samples, the Umbrella protocol will allow the longitudinal collection of samples from the same patient. Additionally, the Umbrella amendment will allow VA SHIELD to accept samples from other preexisting biorepositories or specimen collections.
Central Biorepositories
VA SHIELD has a federated organization with 2 central specimen biorepositories (Palo Alto, CA and Tucson, AZ), and an enterprise data processing center (Boston, MA). The specimen biorepositories receive de-identified specimens that are stored until distribution to approved research projects. The samples and data are linked using an electronic honest broker system to protect privacy, which integrates de-identified specimens with requested clinical and demographic data as needed for approved projects. The honest broker system is operated by independent personnel and does not have vested interest in any studies being performed under VA SHIELD. The integration of sample and associated data is done only as needed when characterization of the donor/participant is necessary byresearch aims or project outcomes. The process is facilitated by a nationally supported laboratory information management system (LIMS), managed by the VA SHIELD data center, that assists with all data requests. The clinical and demographic data are collected from VA electronic health record (EHR), available through VA Corporate Data Warehouse (CDW) and VA Informatics and Computing Infrastructure (VINCI) as needed and integrated with the biorepository samples information for approved VA SHIELD studies. The CDW is the largest longitudinal EHR data collection in the US and has the ability to provide access to national clinical and demographic data.
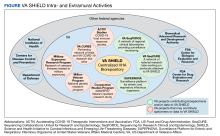
VA SHIELD interacts with multiple VA programs and other entities (Figure). For example, Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) is a network of 5 VA medical centers supported by the Centers for Disease Control and Prevention.4 Its initial goal was to perform surveillance for acute gastroenteritis. In 2020, SUPERNOVA shifted to conduct surveillance for COVID-19 variants among veterans.5 VA SHIELD also interacts with VHA genomic surveillance and sequencing programs: the VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) and VA Sequencing for Research Clinical and Epidemiology (SeqFORCE), described by Krishnan and colleagues.6
Working Groups
To encourage research projects that use biospecimens, VA SHIELD developed content-oriented research working groups. The goal is to inspire collaborations between VA scientists and prevent redundant or overlapping projects. Currently working groups are focused on long COVID, and COVID-19 neurology, pathogen host response, epidemiology and sequencing, cancer and cancer biomarkers, antimicrobial resistance, and vector-borne diseases. Working groups meet regularly to discuss projects and report progress. Working groups also may consider samples that might benefit VA health research and identify potential veteran populations for future research. Working groups connect VA SHIELD and investigators and guide the collection and use of resources.
Ethical Considerations
We recognize the significant ethical concerns for biobanking of specimens. However, there is no general consensus or guideline that addresses all of the complex ethical issues regarding biobanking.7 To address these ethical concerns, we applied the VA Ethical Framework Principles for Access to and Use of Veteran Data principles to VA SHIELD, including all parties who oversee the access to, sharing of, or the use of data, or who access or use its data.8
Conclusions
The VA has assembled a scientific enterprise dedicated to combating emerging infectious diseases and other threats to our patients. This enterprise has been modeled in its structure and oversight to support VA SHIELD. The establishment of a real-time biorepository and data procurement system linked to clinical samples is a bold step forward to address current and future challenges. Similarly, the integration and cooperation of multiple arms within the VA that transcend disciplines and boundaries promise to shepherd a new era of system-wide investigation. In the future, VA SHIELD will integrate with other existing government agencies to advance mutual scientific agendas. VA SHIELD has established the data and biorepository infrastructure to develop innovative and novel technologies to address future challenges. The alignment of basic science, clinical, and translational research goals under one governance is a significant advancement compared with previous models of research coordination.
VA SHIELD was developed to meet an immediate need; it was also framed to be a research enterprise that harnesses the robust clinical and research environment in VHA. The VA SHIELD infrastructure was conceptualized to harmonize specimen and data collection across the VA, allowing researchers to leverage broader collection efforts. Building a biorepository and data collection system within the largest integrated health care system has the potential to provide a lasting impact on VHA and on our nation’s health.
Acknowledgments
The authors wish to acknowledge Ms. Daphne Swancutt for her contribution as copywriter for this manuscript. The authors wish to acknowledge the VA SHIELD investigators: Mary Cloud Ammons, David Beenhouwer, Sheldon T. Brown, Victoria Davey, Abhinav Diwan, John B. Harley, Mark Holodniy, Vincent C. Marconi, Jonathan Moorman, Emerson B. Padiernos, Ian F. Robey, Maria Rodriguez-Barradas, Jason Wertheim, Christopher W. Woods.
1. Lipshy KA, Itani K, Chu D, et al. Sentinel contributions of US Department of Veterans Affairs surgeons in shaping the face of health care. JAMA Surg. 2021;156(4):380-386. doi:10.1001/jamasurg.2020.6372
2. Zucker S, Crabbe JC, Cooper G 4th, et al. Veterans Administration support for medical research: opinions of the endangered species of physician-scientists. FASEB J. 2004;18(13):1481-1486. doi:10.1096/fj.04-1573lfe
3. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
4. Meites E, Bajema KL, Kambhampati A, et al; SUPERNOVA COVID-19 Surveillance Group. Adapting the Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) for COVID-19 among hospitalized adults: surveillance protocol. Front Public Health. 2021;9:739076. doi:10.3389/fpubh.2021.739076
5. Bajema KL, Dahl RM, Evener SL, et al; SUPERNOVA COVID-19 Surveillance Group; Surveillance Platform for Enteric and Respiratory Infectious Organisms at the VA (SUPERNOVA) COVID-19 Surveillance Group. Comparative effectiveness and antibody responses to Moderna and Pfizer-BioNTech COVID-19 vaccines among hospitalized veterans–five Veterans Affairs Medical Centers, United States, February 1-September 30, 2021. MMWR Morb Mortal Wkly Rep. 2021;70(49):1700-1705. doi:10.15585/mmwr.mm7049a2external icon
6. Krishnan J, Woods C, Holodniy M, et al. Nationwide genomic surveillance and response to coronavirus disease 2019 (COVID-19): SeqCURE and SeqFORCE consortiums. Fed Pract. 2023;40(suppl 5):S44-S47. doi:10.12788/fp.0417
7. Ashcroft JW, Macpherson CC. The complex ethical landscape of biobanking. Lancet Public Health. 2019;(6):e274-e275. doi:10.1016/S2468-2667(19)30081-7
8. Principle-Based Ethics Framework for Access to and Use of Veteran Data. Fed Regist. 2022;87(129):40451-40452.
The Veterans Health Administration (VHA) clinicians, clinician-investigators, and investigators perform basic and translational research for the benefit of our nation and are widely recognized for treating patients and discovering cures.1,2 In May 2020, the US Department of Veterans Affairs (VA) launched the VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). The goal of this novel enterprise was to assemble a comprehensive specimen and data repository for emerging life-threatening diseases and to address future challenges. VA SHIELD was specifically charged with creating a biorepository to advance research, improve diagnostic and therapeutic capabilities, and develop strategies for immediate deployment to VA clinical environments. One main objective of VA SHIELD is to harness the clinical and scientific strengths of the VA in order to create a more cohesive collaboration between preexisting clinical research efforts within the VA.
ANATOMY OF VA SHIELD
The charge and scope of VA SHIELD is unique.3 As an entity, this program leverages the strengths of the diverse VHA network, has a broad potential impact on national health care, is positioned to respond rapidly to national and international health-related events, and substantially contributes to clinical research and development. In addition, VA SHIELD upholds VA’s Fourth Mission, which is to contribute to national emergencies and support emergency management, public health, safety, and homeland security efforts.
VA SHIELD is part of the VA Office of Research and Development (ORD). The coordinating center (CC), headquartered in Cleveland, Ohio, is the central operational partner, leading VA SHIELD and interacting with other important VA programs, including laboratory, clinical science, rehabilitation, and health services. The VA SHIELD CC oversees all aspects of operations, including biospecimen collection, creating and enforcing of standard operating procedures, ensuring the quality of the samples, processing research applications, distribution of samples, financing, and progress reports. The CC also initiates and maintains interagency collaborations, convenes stakeholders, and develops strategic plans to address emerging diseases.
The VA SHIELD Executive Steering Committee (ESC) is composed of infectious disease, biorepository, and public health specialists. The ESC provides scientific and programmatic direction to the CC, including operational activities and guidance regarding biorepository priorities and scientific agenda, and measuring and reporting on VA SHIELD accomplishments.
The primary function of the Programmatic and Scientific Review Board (PSRB) is to evaluate incoming research proposals for specimen and data use for feasibility and make recommendations to the VA SHIELD CC. The PSRB evaluates and ensures that data and specimen use align with VA SHIELD ethical, clinical, and scientific objectives.
VA SHIELD IN PRACTICE
VA SHIELD consisted of 11 specimen collection sites (Atlanta, GA; Boise, ID; Bronx, NY; Cincinnati, OH; Cleveland, OH; Durham, NC; Houston, TX; Los Angeles, CA; Mountain Home, TN; Palo Alto, CA; and Tucson, AZ), a data processing center in Boston, MA, and 2 central biorepositories in Palo Alto, CA, and Tucson, AZ. Information flow is a coordinated process among specimen collection sites, data processing centers, and the biorepositories. Initially, each local collection site identifies residual specimens that would have been discarded after clinical laboratory testing. These samples currently account for the majority of biological material within VA SHIELD via a novel collection protocol known as “Sweep,” which allows residual clinical discarded samples as well as samples from patients with new emerging infectious and noninfectious diseases of concern to be collected at the time of first emergence and submitted to VA SHIELD during the course of routine veteran health care.3 These clinical discarded samples are de-identified and transferred from the clinical laboratory to VA SHIELD. The VA Central Institutional Review Board (cIRB) has approved the use of these samples as nonhuman subject research. Biological samples are collected, processed, aliquoted, shipped to, and stored at the central biorepository sites.
The Umbrella amendment to Sweep that has been approved also by the VA cIRB, will allow VA SHIELD sites to prospectively consent veterans and collect biospecimens and additional clinical and self-reported information. The implementation of Umbrella could significantly enhance collection and research. Although Sweep is a onetime collection of samples, the Umbrella protocol will allow the longitudinal collection of samples from the same patient. Additionally, the Umbrella amendment will allow VA SHIELD to accept samples from other preexisting biorepositories or specimen collections.
Central Biorepositories
VA SHIELD has a federated organization with 2 central specimen biorepositories (Palo Alto, CA and Tucson, AZ), and an enterprise data processing center (Boston, MA). The specimen biorepositories receive de-identified specimens that are stored until distribution to approved research projects. The samples and data are linked using an electronic honest broker system to protect privacy, which integrates de-identified specimens with requested clinical and demographic data as needed for approved projects. The honest broker system is operated by independent personnel and does not have vested interest in any studies being performed under VA SHIELD. The integration of sample and associated data is done only as needed when characterization of the donor/participant is necessary byresearch aims or project outcomes. The process is facilitated by a nationally supported laboratory information management system (LIMS), managed by the VA SHIELD data center, that assists with all data requests. The clinical and demographic data are collected from VA electronic health record (EHR), available through VA Corporate Data Warehouse (CDW) and VA Informatics and Computing Infrastructure (VINCI) as needed and integrated with the biorepository samples information for approved VA SHIELD studies. The CDW is the largest longitudinal EHR data collection in the US and has the ability to provide access to national clinical and demographic data.
VA SHIELD interacts with multiple VA programs and other entities (Figure). For example, Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) is a network of 5 VA medical centers supported by the Centers for Disease Control and Prevention.4 Its initial goal was to perform surveillance for acute gastroenteritis. In 2020, SUPERNOVA shifted to conduct surveillance for COVID-19 variants among veterans.5 VA SHIELD also interacts with VHA genomic surveillance and sequencing programs: the VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) and VA Sequencing for Research Clinical and Epidemiology (SeqFORCE), described by Krishnan and colleagues.6
Working Groups
To encourage research projects that use biospecimens, VA SHIELD developed content-oriented research working groups. The goal is to inspire collaborations between VA scientists and prevent redundant or overlapping projects. Currently working groups are focused on long COVID, and COVID-19 neurology, pathogen host response, epidemiology and sequencing, cancer and cancer biomarkers, antimicrobial resistance, and vector-borne diseases. Working groups meet regularly to discuss projects and report progress. Working groups also may consider samples that might benefit VA health research and identify potential veteran populations for future research. Working groups connect VA SHIELD and investigators and guide the collection and use of resources.
Ethical Considerations
We recognize the significant ethical concerns for biobanking of specimens. However, there is no general consensus or guideline that addresses all of the complex ethical issues regarding biobanking.7 To address these ethical concerns, we applied the VA Ethical Framework Principles for Access to and Use of Veteran Data principles to VA SHIELD, including all parties who oversee the access to, sharing of, or the use of data, or who access or use its data.8
Conclusions
The VA has assembled a scientific enterprise dedicated to combating emerging infectious diseases and other threats to our patients. This enterprise has been modeled in its structure and oversight to support VA SHIELD. The establishment of a real-time biorepository and data procurement system linked to clinical samples is a bold step forward to address current and future challenges. Similarly, the integration and cooperation of multiple arms within the VA that transcend disciplines and boundaries promise to shepherd a new era of system-wide investigation. In the future, VA SHIELD will integrate with other existing government agencies to advance mutual scientific agendas. VA SHIELD has established the data and biorepository infrastructure to develop innovative and novel technologies to address future challenges. The alignment of basic science, clinical, and translational research goals under one governance is a significant advancement compared with previous models of research coordination.
VA SHIELD was developed to meet an immediate need; it was also framed to be a research enterprise that harnesses the robust clinical and research environment in VHA. The VA SHIELD infrastructure was conceptualized to harmonize specimen and data collection across the VA, allowing researchers to leverage broader collection efforts. Building a biorepository and data collection system within the largest integrated health care system has the potential to provide a lasting impact on VHA and on our nation’s health.
Acknowledgments
The authors wish to acknowledge Ms. Daphne Swancutt for her contribution as copywriter for this manuscript. The authors wish to acknowledge the VA SHIELD investigators: Mary Cloud Ammons, David Beenhouwer, Sheldon T. Brown, Victoria Davey, Abhinav Diwan, John B. Harley, Mark Holodniy, Vincent C. Marconi, Jonathan Moorman, Emerson B. Padiernos, Ian F. Robey, Maria Rodriguez-Barradas, Jason Wertheim, Christopher W. Woods.
The Veterans Health Administration (VHA) clinicians, clinician-investigators, and investigators perform basic and translational research for the benefit of our nation and are widely recognized for treating patients and discovering cures.1,2 In May 2020, the US Department of Veterans Affairs (VA) launched the VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). The goal of this novel enterprise was to assemble a comprehensive specimen and data repository for emerging life-threatening diseases and to address future challenges. VA SHIELD was specifically charged with creating a biorepository to advance research, improve diagnostic and therapeutic capabilities, and develop strategies for immediate deployment to VA clinical environments. One main objective of VA SHIELD is to harness the clinical and scientific strengths of the VA in order to create a more cohesive collaboration between preexisting clinical research efforts within the VA.
ANATOMY OF VA SHIELD
The charge and scope of VA SHIELD is unique.3 As an entity, this program leverages the strengths of the diverse VHA network, has a broad potential impact on national health care, is positioned to respond rapidly to national and international health-related events, and substantially contributes to clinical research and development. In addition, VA SHIELD upholds VA’s Fourth Mission, which is to contribute to national emergencies and support emergency management, public health, safety, and homeland security efforts.
VA SHIELD is part of the VA Office of Research and Development (ORD). The coordinating center (CC), headquartered in Cleveland, Ohio, is the central operational partner, leading VA SHIELD and interacting with other important VA programs, including laboratory, clinical science, rehabilitation, and health services. The VA SHIELD CC oversees all aspects of operations, including biospecimen collection, creating and enforcing of standard operating procedures, ensuring the quality of the samples, processing research applications, distribution of samples, financing, and progress reports. The CC also initiates and maintains interagency collaborations, convenes stakeholders, and develops strategic plans to address emerging diseases.
The VA SHIELD Executive Steering Committee (ESC) is composed of infectious disease, biorepository, and public health specialists. The ESC provides scientific and programmatic direction to the CC, including operational activities and guidance regarding biorepository priorities and scientific agenda, and measuring and reporting on VA SHIELD accomplishments.
The primary function of the Programmatic and Scientific Review Board (PSRB) is to evaluate incoming research proposals for specimen and data use for feasibility and make recommendations to the VA SHIELD CC. The PSRB evaluates and ensures that data and specimen use align with VA SHIELD ethical, clinical, and scientific objectives.
VA SHIELD IN PRACTICE
VA SHIELD consisted of 11 specimen collection sites (Atlanta, GA; Boise, ID; Bronx, NY; Cincinnati, OH; Cleveland, OH; Durham, NC; Houston, TX; Los Angeles, CA; Mountain Home, TN; Palo Alto, CA; and Tucson, AZ), a data processing center in Boston, MA, and 2 central biorepositories in Palo Alto, CA, and Tucson, AZ. Information flow is a coordinated process among specimen collection sites, data processing centers, and the biorepositories. Initially, each local collection site identifies residual specimens that would have been discarded after clinical laboratory testing. These samples currently account for the majority of biological material within VA SHIELD via a novel collection protocol known as “Sweep,” which allows residual clinical discarded samples as well as samples from patients with new emerging infectious and noninfectious diseases of concern to be collected at the time of first emergence and submitted to VA SHIELD during the course of routine veteran health care.3 These clinical discarded samples are de-identified and transferred from the clinical laboratory to VA SHIELD. The VA Central Institutional Review Board (cIRB) has approved the use of these samples as nonhuman subject research. Biological samples are collected, processed, aliquoted, shipped to, and stored at the central biorepository sites.
The Umbrella amendment to Sweep that has been approved also by the VA cIRB, will allow VA SHIELD sites to prospectively consent veterans and collect biospecimens and additional clinical and self-reported information. The implementation of Umbrella could significantly enhance collection and research. Although Sweep is a onetime collection of samples, the Umbrella protocol will allow the longitudinal collection of samples from the same patient. Additionally, the Umbrella amendment will allow VA SHIELD to accept samples from other preexisting biorepositories or specimen collections.
Central Biorepositories
VA SHIELD has a federated organization with 2 central specimen biorepositories (Palo Alto, CA and Tucson, AZ), and an enterprise data processing center (Boston, MA). The specimen biorepositories receive de-identified specimens that are stored until distribution to approved research projects. The samples and data are linked using an electronic honest broker system to protect privacy, which integrates de-identified specimens with requested clinical and demographic data as needed for approved projects. The honest broker system is operated by independent personnel and does not have vested interest in any studies being performed under VA SHIELD. The integration of sample and associated data is done only as needed when characterization of the donor/participant is necessary byresearch aims or project outcomes. The process is facilitated by a nationally supported laboratory information management system (LIMS), managed by the VA SHIELD data center, that assists with all data requests. The clinical and demographic data are collected from VA electronic health record (EHR), available through VA Corporate Data Warehouse (CDW) and VA Informatics and Computing Infrastructure (VINCI) as needed and integrated with the biorepository samples information for approved VA SHIELD studies. The CDW is the largest longitudinal EHR data collection in the US and has the ability to provide access to national clinical and demographic data.
VA SHIELD interacts with multiple VA programs and other entities (Figure). For example, Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) is a network of 5 VA medical centers supported by the Centers for Disease Control and Prevention.4 Its initial goal was to perform surveillance for acute gastroenteritis. In 2020, SUPERNOVA shifted to conduct surveillance for COVID-19 variants among veterans.5 VA SHIELD also interacts with VHA genomic surveillance and sequencing programs: the VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) and VA Sequencing for Research Clinical and Epidemiology (SeqFORCE), described by Krishnan and colleagues.6
Working Groups
To encourage research projects that use biospecimens, VA SHIELD developed content-oriented research working groups. The goal is to inspire collaborations between VA scientists and prevent redundant or overlapping projects. Currently working groups are focused on long COVID, and COVID-19 neurology, pathogen host response, epidemiology and sequencing, cancer and cancer biomarkers, antimicrobial resistance, and vector-borne diseases. Working groups meet regularly to discuss projects and report progress. Working groups also may consider samples that might benefit VA health research and identify potential veteran populations for future research. Working groups connect VA SHIELD and investigators and guide the collection and use of resources.
Ethical Considerations
We recognize the significant ethical concerns for biobanking of specimens. However, there is no general consensus or guideline that addresses all of the complex ethical issues regarding biobanking.7 To address these ethical concerns, we applied the VA Ethical Framework Principles for Access to and Use of Veteran Data principles to VA SHIELD, including all parties who oversee the access to, sharing of, or the use of data, or who access or use its data.8
Conclusions
The VA has assembled a scientific enterprise dedicated to combating emerging infectious diseases and other threats to our patients. This enterprise has been modeled in its structure and oversight to support VA SHIELD. The establishment of a real-time biorepository and data procurement system linked to clinical samples is a bold step forward to address current and future challenges. Similarly, the integration and cooperation of multiple arms within the VA that transcend disciplines and boundaries promise to shepherd a new era of system-wide investigation. In the future, VA SHIELD will integrate with other existing government agencies to advance mutual scientific agendas. VA SHIELD has established the data and biorepository infrastructure to develop innovative and novel technologies to address future challenges. The alignment of basic science, clinical, and translational research goals under one governance is a significant advancement compared with previous models of research coordination.
VA SHIELD was developed to meet an immediate need; it was also framed to be a research enterprise that harnesses the robust clinical and research environment in VHA. The VA SHIELD infrastructure was conceptualized to harmonize specimen and data collection across the VA, allowing researchers to leverage broader collection efforts. Building a biorepository and data collection system within the largest integrated health care system has the potential to provide a lasting impact on VHA and on our nation’s health.
Acknowledgments
The authors wish to acknowledge Ms. Daphne Swancutt for her contribution as copywriter for this manuscript. The authors wish to acknowledge the VA SHIELD investigators: Mary Cloud Ammons, David Beenhouwer, Sheldon T. Brown, Victoria Davey, Abhinav Diwan, John B. Harley, Mark Holodniy, Vincent C. Marconi, Jonathan Moorman, Emerson B. Padiernos, Ian F. Robey, Maria Rodriguez-Barradas, Jason Wertheim, Christopher W. Woods.
1. Lipshy KA, Itani K, Chu D, et al. Sentinel contributions of US Department of Veterans Affairs surgeons in shaping the face of health care. JAMA Surg. 2021;156(4):380-386. doi:10.1001/jamasurg.2020.6372
2. Zucker S, Crabbe JC, Cooper G 4th, et al. Veterans Administration support for medical research: opinions of the endangered species of physician-scientists. FASEB J. 2004;18(13):1481-1486. doi:10.1096/fj.04-1573lfe
3. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
4. Meites E, Bajema KL, Kambhampati A, et al; SUPERNOVA COVID-19 Surveillance Group. Adapting the Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) for COVID-19 among hospitalized adults: surveillance protocol. Front Public Health. 2021;9:739076. doi:10.3389/fpubh.2021.739076
5. Bajema KL, Dahl RM, Evener SL, et al; SUPERNOVA COVID-19 Surveillance Group; Surveillance Platform for Enteric and Respiratory Infectious Organisms at the VA (SUPERNOVA) COVID-19 Surveillance Group. Comparative effectiveness and antibody responses to Moderna and Pfizer-BioNTech COVID-19 vaccines among hospitalized veterans–five Veterans Affairs Medical Centers, United States, February 1-September 30, 2021. MMWR Morb Mortal Wkly Rep. 2021;70(49):1700-1705. doi:10.15585/mmwr.mm7049a2external icon
6. Krishnan J, Woods C, Holodniy M, et al. Nationwide genomic surveillance and response to coronavirus disease 2019 (COVID-19): SeqCURE and SeqFORCE consortiums. Fed Pract. 2023;40(suppl 5):S44-S47. doi:10.12788/fp.0417
7. Ashcroft JW, Macpherson CC. The complex ethical landscape of biobanking. Lancet Public Health. 2019;(6):e274-e275. doi:10.1016/S2468-2667(19)30081-7
8. Principle-Based Ethics Framework for Access to and Use of Veteran Data. Fed Regist. 2022;87(129):40451-40452.
1. Lipshy KA, Itani K, Chu D, et al. Sentinel contributions of US Department of Veterans Affairs surgeons in shaping the face of health care. JAMA Surg. 2021;156(4):380-386. doi:10.1001/jamasurg.2020.6372
2. Zucker S, Crabbe JC, Cooper G 4th, et al. Veterans Administration support for medical research: opinions of the endangered species of physician-scientists. FASEB J. 2004;18(13):1481-1486. doi:10.1096/fj.04-1573lfe
3. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
4. Meites E, Bajema KL, Kambhampati A, et al; SUPERNOVA COVID-19 Surveillance Group. Adapting the Surveillance Platform for Enteric and Respiratory Infectious Organisms at United States Veterans Affairs Medical Centers (SUPERNOVA) for COVID-19 among hospitalized adults: surveillance protocol. Front Public Health. 2021;9:739076. doi:10.3389/fpubh.2021.739076
5. Bajema KL, Dahl RM, Evener SL, et al; SUPERNOVA COVID-19 Surveillance Group; Surveillance Platform for Enteric and Respiratory Infectious Organisms at the VA (SUPERNOVA) COVID-19 Surveillance Group. Comparative effectiveness and antibody responses to Moderna and Pfizer-BioNTech COVID-19 vaccines among hospitalized veterans–five Veterans Affairs Medical Centers, United States, February 1-September 30, 2021. MMWR Morb Mortal Wkly Rep. 2021;70(49):1700-1705. doi:10.15585/mmwr.mm7049a2external icon
6. Krishnan J, Woods C, Holodniy M, et al. Nationwide genomic surveillance and response to coronavirus disease 2019 (COVID-19): SeqCURE and SeqFORCE consortiums. Fed Pract. 2023;40(suppl 5):S44-S47. doi:10.12788/fp.0417
7. Ashcroft JW, Macpherson CC. The complex ethical landscape of biobanking. Lancet Public Health. 2019;(6):e274-e275. doi:10.1016/S2468-2667(19)30081-7
8. Principle-Based Ethics Framework for Access to and Use of Veteran Data. Fed Regist. 2022;87(129):40451-40452.
Nationwide Genomic Surveillance and Response to COVID-19: The VA SeqFORCE and SeqCURE Consortiums
The COVID-19 virus and its associated pandemic have highlighted the urgent need for a national infrastructure to rapidly identify and respond to emerging pathogens. The importance of understanding viral population dynamics through genetic sequencing has become apparent over time, particularly as the vaccine responses, clinical implications, and therapeutic effectiveness of treatments have varied substantially with COVID-19 variants.1,2
As the largest integrated health care system in the US, the US Department of Veterans Affairs (VA) is uniquely situated to help with pandemic detection and response. This article highlights 2 VA programs dedicated to COVID-19 sequencing at the forefront of pandemic response and research: VA Sequencing for Research Clinical and Epidemiology (SeqFORCE) and VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) (Table).
VA Seq FORCE
VA SeqFORCE was established March 2021 to facilitate clinical surveillance of COVID-19 variants in the US veteran population and in VA employees. VA SeqFORCE consists of 9 Clinical Laboratory Improvement Amendment (CLIA)–certified laboratories in VA medical centers, including the VA Public Health Reference Laboratory in Palo Alto, California, and 8 Veterans Health Administration (VHA) clinical laboratories (Los Angeles, California; Boise, Idaho; Iowa City, Iowa; Bronx, New York; West Haven, Connecticut; Indianapolis, Indiana; Denver, Colorado; and Orlando, Florida).3 Specimen standards (eg, real-time polymerase chain reaction [RT-PCR] cycle threshold [Ct] ≤ 30, minimum volume, etc) and clinical criteria (eg, COVID-19–related deaths, COVID-19 vaccine escape, etc) for submitting samples to VA SeqFORCE laboratories were established, and logistics for sample sequencing was centralized, including providing centralized instructions for sample preparation and to which VA SeqFORCE laboratory samples should be sent.
These laboratories sequenced samples from patients and employees with COVID-19 to understand patterns of variant evolution, vaccine, antiviral and monoclonal antibody response, health care–associated outbreaks, and COVID-19 transmission. As clinically relevant findings, such as monoclonal antibody treatment failure, emerged with novel viral variants, VA SeqFORCE was well positioned to rapidly detect the emergent variants and inform better clinical care of patients with COVID-19. Other clinical indications identified for sequencing within VA SeqFORCE included outbreak investigation, re-infection with COVID-19 > 90 days but < 6 months after a prior infection, extended hospitalization of > 21 days, death due to COVID-19, infection with a history of recent nondomestic travel, rebound of symptoms after improvement on oral antiviral therapy, and epidemiologic surveillance.
VA SeqFORCE laboratories use a variety of sequencing platforms, although a federated system was developed that electronically linked all laboratories using a software system (PraediGene, Bitscopic) for sample management, COVID-19 variant analytics, and automated result reporting of clade and lineage into the Veterans Health Information Systems and Technology Architecture (VistA) Computerized Patient Record System. In addition, generated nucleic acid sequence alignment through FASTA consensus sequence files have been archived for secondary research analyses. By archiving the consensus sequences, retrospective studies within the VA have the added benefit of being able to clinically annotate investigations into COVID-19 variant patterns. As of August 2023, 43,003 samples containing COVID-19 have been sequenced, and FASTA file and metadata upload are ongoing to the Global Initiative on Sharing Avian Influenza Data, which houses > 15 million COVID-19 files from global submissions.
VA SeqFORCE’s clinical sequencing efforts have created opportunities for multicenter collaboration in variant surveillance. In work from December 2021, investigators from the James J. Peters VA Medical Center in Bronx, New York, collaborated with the VHA Pathology and Laboratory Medicine Services and Public Health national program offices in Washington, DC, to develop an RT-PCR assay to rapidly differentiate Omicron from Delta variants.4 Samples from VA hospitals across the nation were used in this study.
Lessons from VA SeqFORCE have also been cited as inspiration to address COVID-19 clinical problems, including outbreak investigations in hospital settings and beyond. Researchers at the Iowa City VA Health Care System, for example, proposed a novel probabilistic quantitative method for determining genetic-relatedness among COVID-19 viral strains in an outbreak setting.5 They extended the scope of work to develop COVID-19 outbreak screening tools combining publicly available algorithms with targeted sequencing data to identify outbreaks as they arise.6 We expect VA SeqFORCE, in conjunction with its complement VA SeqCURE, will continue to further pandemic surveillance and response.
VA Seq CURE
As the research-focused complement to VA SeqFORCE, VA SeqCURE is dedicated to a broader study of the COVID-19 genome through sequencing. Established January 2021, the VA SeqCURE network consists of 6 research laboratories in Boise, Idaho; Bronx, New York; Cleveland, Ohio; Durham, North Carolina; Iowa City, Iowa; and Temple, Texas.
Samples are collected as a subset of the broader VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD) biorepository sweep protocol for discarded blood and nasal swab specimens of VHA patients hospitalized with COVID-19, as described by Epstein and colleagues.7-9 While VA SeqFORCE sequences samples positive for COVID-19 by RT-PCR with a Ct value of ≤ 30 for diagnostic purposes, VA SeqCURE laboratories sequence more broadly for nondiagnostic purposes, including samples with a Ct value > 30. The 6 VA SeqCURE laboratories generate sequencing data using various platforms, amplification kits, and formats. To ensure maximum quality and metadata on the sequences generated across the different laboratories, a sequence intake pipeline has been developed, adapting the ViralRecon bioinformatics platform.10 This harmonized analysis pipeline accommodates different file formats and performs quality control, alignment, variant calling, lineage assignment, clade assignment, and annotation. As of August 2023, VA SeqCURE has identified viral sequences from 24,107 unique specimens. Annotated COVID-19 sequences with the appropriate metadata will be available to VA researchers through VA SHIELD.
Research projects include descriptive epidemiology of COVID-19 variants in individuals who receive VHA care, COVID-19 vaccine and therapy effectiveness, and the unique distribution of variants and vaccine effectiveness in rural settings.3 True to its core mission, members of the VA SeqCURE consortium have contributed to the COVID-19 viral sequencing literature over the past 2 years. Researchers also are accessing VA SeqCURE to study COVID-19 persistence and rebound among individuals with mild disease taking nirmatrelvir/ritonavir compared with other COVID-19 therapeutics and untreated controls. Finally, COVID-19 samples and their sequences are stored in the VA SHIELD biorepository, which leverages these samples and data to advance scientific understanding of COVID-19 and future emerging infectious diseases.7-9
Important work from investigators at the Central Texas Veterans Health Care System confronted the issue of whole genome sequencing data from COVID-19 samples with low viral loads, a common issue with COVID-19 sequencing. They found that yields of 2 sequencing protocols, which generated high-sequence coverage, were enhanced further by combining the results of both methods.11 This project, which has potentially broad applications for sequencing in research and clinical settings, is an example of VA SeqCURE’s efforts to address the COVID-19 pandemic. The VA SeqCURE program has substantial potential as a large viral sequencing repository with broad geographic and demographic representation, such that future large-scale sequencing analyses may be generated from preexisting nested cohorts within the repository.
NEXT STEPS
Promising new directions of clinical and laboratory-based research are planned for VA SeqFORCE and VA SeqCURE. While the impact of COVID-19 and other viruses with epidemic potential is perhaps most feared in urban settings, evidence suggests that the distribution of COVID-19 in rural settings is unique and associated with worse outcomes.12,13 Given the wide catchment areas of VA hospitals that encompass both rural and urban settings, the VA’s ongoing COVID-19 sequencing programs and repositories are uniquely positioned to understand viral dynamics in areas of differing population density.
While rates of infection, hospitalization, and death resulting from COVID-19 have substantially dropped, the long-term impact of the pandemic is just beginning to be recognized in conditions such as long COVID or postacute COVID-19 syndrome. Long COVID has already proven to be biologically multifaceted, difficult to diagnose, and unpredictable in identifying the most at-risk patients.14-16 Much remains to be determined in our understanding of long COVID, including a unified definition that can effectively be used in clinical settings to diagnose and treat patients. However, research indicates that comorbidities common in veterans, such as diabetes and cardiovascular disease, are associated with worse long-term outcomes.17,18 Collaborations between VA scientists, clinicians, and national cooperative programs (such as a network of VHA long COVID clinics) create an unmatched opportunity for VA SeqFORCE and VA SeqCURE programs to provide insight into a disease likely to become a chronic disease outcome of the pandemic.
With VA SeqFORCE and VA SeqCURE programs, the VA now has infrastructure ready to respond to new infectious diseases. During the mpox outbreak of 2022, the VA Public Health Reference Laboratory received > 80% of all VA mpox samples for orthopox screening and mpox confirmatory testing. A subset of these samples underwent whole genome sequencing with the identification of 10 unique lineages across VA, and > 200 positive and 400 negative samples have been aliquoted and submitted to VA SHIELD for research. Furthermore, the VA SeqFORCE and VA SeqCURE sequencing processes might be adapted to identify outbreaks of multidrug-resistant organisms among VA patients trialed at other institutions.19 We are hopeful that VA SeqFORCE and VA SeqCURE will become invaluable components of health care delivery and infection prevention at the hospital level and beyond.
Finally, the robust data infrastructure and associated repositories of VA SeqFORCE and VA SeqCURE may be leveraged to study noninfectious diseases. Research groups are starting to apply these programs to cancer sequencing. We anticipate that these efforts may have a substantial impact on our understanding of cancer epidemiology and region-specific risk factors for malignancy, given the size and breadth of VA SeqFORCE and VA SeqCURE. Common oncogenic mutations identified through these programs could be targets for precision oncology therapeutics. Similarly, we envision applications of the VA SeqFORCE and VA SeqCURE data infrastructures and repositories toward other precision medicine fields, including pharmacogenomics and nutrition, to tailor interventions to meet the specific individual needs of veterans.
CONCLUSIONS
The productivity of VA SeqFORCE and VA SeqCURE programs over the past 2 years continues to increase in response to the COVID-19 pandemic. We anticipate that they will be vital components in our nation’s responses to infectious threats and beyond.
1. Iuliano AD, Brunkard JM, Boehmer TK, et al. Trends in disease severity and health care utilization during the early Omicron variant period compared with previous SARS-CoV-2 high transmission periods - United States, December 2020-January 2022. MMWR Morb Mortal Wkly Rep. 2022;71(4):146-152. Published 2022 Jan 28. doi:10.15585/mmwr.mm7104e4
2. Nyberg T, Ferguson NM, Nash SG, et al. Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. Lancet. 2022;399(10332):1303-1312. doi:10.1016/S0140-6736(22)00462-7
3. Veterans Health Administration. Coronavirus Disease 2019 (COVID-19) response report - annex C. December 5, 2022. Accessed August 28, 2023. https://www.va.gov/HEALTH/docs/VHA-COVID-19-Response-2022-Annex-C.pdf 4. Barasch NJ, Iqbal J, Coombs M, et al. Utilization of a SARS-CoV-2 variant assay for the rapid differentiation of Omicron and Delta. medRxiv. Preprint posted online December 27, 2021. doi:10.1101/2021.12.22.21268195
5. Bilal MY. Similarity Index-probabilistic confidence estimation of SARS-CoV-2 strain relatedness in localized outbreaks. Epidemiologia (Basel). 2022;3(2):238-249. doi:10.3390/epidemiologia3020019
6. Bilal MY, Klutts JS. Molecular Epidemiological investigations of localized SARS-CoV-2 outbreaks-utility of public algorithms. Epidemiologia (Basel). 2022;3(3):402-411. doi:10.3390/epidemiologia3030031
7. Veterans Health Administration, Office of Research & Development. VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). Updated November 23, 2022. Accessed August 28, 2023. https://www.research.va.gov/programs/shield/about.cfm
8. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
9. Epstein L, Shive C, Garcia AP, et al. VA SHIELD: a biorepository for our veterans and the nation. Fed Pract. 2023;40(suppl 5):S48-S51. doi:10.12788/fp.0424
10. Patel H, Varona S, Monzón S, et al. Version 2.5. nf-core/viralrecon: nf-core/viralrecon v2.5 - Manganese Monkey (2.5). Zenodo. July 13, 2022. doi:10.5281/zenodo.6827984
11. Choi H, Hwang M, Navarathna DH, Xu J, Lukey J, Jinadatha C. Performance of COVIDSeq and swift normalase amplicon SARS-CoV-2 panels for SARS-CoV-2 genome sequencing: practical guide and combining FASTQ strategy. J Clin Microbiol. 2022;60(4):e0002522. doi:10.1128/jcm.00025-22
12. Cuadros DF, Branscum AJ, Mukandavire Z, Miller FD, MacKinnon N. Dynamics of the COVID-19 epidemic in urban and rural areas in the United States. Ann Epidemiol. 2021;59:16-20. doi:10.1016/j.annepidem.2021.04.007
13. Anzalone AJ, Horswell R, Hendricks BM, et al. Higher hospitalization and mortality rates among SARS-CoV-2-infected persons in rural America. J Rural Health. 2023;39(1):39-54. doi:10.1111/jrh.12689
14. Su Y, Yuan D, Chen DG, et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell. 2022;185(5):881-895.e20. doi:10.1016/j.cell.2022.01.014
15. Pfaff ER, Girvin AT, Bennett TD, et al. Identifying who has long COVID in the USA: a machine learning approach using N3C data. Lancet Digit Health. 2022;4(7):e532-e541. doi:10.1016/S2589-7500(22)00048-6
16. Subramanian A, Nirantharakumar K, Hughes S, et al. Symptoms and risk factors for long COVID in non-hospitalized adults. Nat Med. 2022;28(8):1706-1714. doi:10.1038/s41591-022-01909-w
17. Munblit D, O’Hara ME, Akrami A, Perego E, Olliaro P, Needham DM. Long COVID: aiming for a consensus. Lancet Respir Med. 2022;10(7):632-634. doi:10.1016/S2213-2600(22)00135-7
18. Thaweethai T, Jolley SE, Karlson EW, et al. Development of a definition of postacute sequelae of SARS-CoV-2 infection. JAMA. 2023;329(22):1934-1946. doi:10.1001/jama.2023.8823
19. Sundermann AJ, Chen J, Kumar P, et al. Whole-genome sequencing surveillance and machine learning of the electronic health record for enhanced healthcare outbreak detection. Clin Infect Dis. 2022;75(3):476-482. doi:10.1093/cid/ciab946
The COVID-19 virus and its associated pandemic have highlighted the urgent need for a national infrastructure to rapidly identify and respond to emerging pathogens. The importance of understanding viral population dynamics through genetic sequencing has become apparent over time, particularly as the vaccine responses, clinical implications, and therapeutic effectiveness of treatments have varied substantially with COVID-19 variants.1,2
As the largest integrated health care system in the US, the US Department of Veterans Affairs (VA) is uniquely situated to help with pandemic detection and response. This article highlights 2 VA programs dedicated to COVID-19 sequencing at the forefront of pandemic response and research: VA Sequencing for Research Clinical and Epidemiology (SeqFORCE) and VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) (Table).
VA Seq FORCE
VA SeqFORCE was established March 2021 to facilitate clinical surveillance of COVID-19 variants in the US veteran population and in VA employees. VA SeqFORCE consists of 9 Clinical Laboratory Improvement Amendment (CLIA)–certified laboratories in VA medical centers, including the VA Public Health Reference Laboratory in Palo Alto, California, and 8 Veterans Health Administration (VHA) clinical laboratories (Los Angeles, California; Boise, Idaho; Iowa City, Iowa; Bronx, New York; West Haven, Connecticut; Indianapolis, Indiana; Denver, Colorado; and Orlando, Florida).3 Specimen standards (eg, real-time polymerase chain reaction [RT-PCR] cycle threshold [Ct] ≤ 30, minimum volume, etc) and clinical criteria (eg, COVID-19–related deaths, COVID-19 vaccine escape, etc) for submitting samples to VA SeqFORCE laboratories were established, and logistics for sample sequencing was centralized, including providing centralized instructions for sample preparation and to which VA SeqFORCE laboratory samples should be sent.
These laboratories sequenced samples from patients and employees with COVID-19 to understand patterns of variant evolution, vaccine, antiviral and monoclonal antibody response, health care–associated outbreaks, and COVID-19 transmission. As clinically relevant findings, such as monoclonal antibody treatment failure, emerged with novel viral variants, VA SeqFORCE was well positioned to rapidly detect the emergent variants and inform better clinical care of patients with COVID-19. Other clinical indications identified for sequencing within VA SeqFORCE included outbreak investigation, re-infection with COVID-19 > 90 days but < 6 months after a prior infection, extended hospitalization of > 21 days, death due to COVID-19, infection with a history of recent nondomestic travel, rebound of symptoms after improvement on oral antiviral therapy, and epidemiologic surveillance.
VA SeqFORCE laboratories use a variety of sequencing platforms, although a federated system was developed that electronically linked all laboratories using a software system (PraediGene, Bitscopic) for sample management, COVID-19 variant analytics, and automated result reporting of clade and lineage into the Veterans Health Information Systems and Technology Architecture (VistA) Computerized Patient Record System. In addition, generated nucleic acid sequence alignment through FASTA consensus sequence files have been archived for secondary research analyses. By archiving the consensus sequences, retrospective studies within the VA have the added benefit of being able to clinically annotate investigations into COVID-19 variant patterns. As of August 2023, 43,003 samples containing COVID-19 have been sequenced, and FASTA file and metadata upload are ongoing to the Global Initiative on Sharing Avian Influenza Data, which houses > 15 million COVID-19 files from global submissions.
VA SeqFORCE’s clinical sequencing efforts have created opportunities for multicenter collaboration in variant surveillance. In work from December 2021, investigators from the James J. Peters VA Medical Center in Bronx, New York, collaborated with the VHA Pathology and Laboratory Medicine Services and Public Health national program offices in Washington, DC, to develop an RT-PCR assay to rapidly differentiate Omicron from Delta variants.4 Samples from VA hospitals across the nation were used in this study.
Lessons from VA SeqFORCE have also been cited as inspiration to address COVID-19 clinical problems, including outbreak investigations in hospital settings and beyond. Researchers at the Iowa City VA Health Care System, for example, proposed a novel probabilistic quantitative method for determining genetic-relatedness among COVID-19 viral strains in an outbreak setting.5 They extended the scope of work to develop COVID-19 outbreak screening tools combining publicly available algorithms with targeted sequencing data to identify outbreaks as they arise.6 We expect VA SeqFORCE, in conjunction with its complement VA SeqCURE, will continue to further pandemic surveillance and response.
VA Seq CURE
As the research-focused complement to VA SeqFORCE, VA SeqCURE is dedicated to a broader study of the COVID-19 genome through sequencing. Established January 2021, the VA SeqCURE network consists of 6 research laboratories in Boise, Idaho; Bronx, New York; Cleveland, Ohio; Durham, North Carolina; Iowa City, Iowa; and Temple, Texas.
Samples are collected as a subset of the broader VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD) biorepository sweep protocol for discarded blood and nasal swab specimens of VHA patients hospitalized with COVID-19, as described by Epstein and colleagues.7-9 While VA SeqFORCE sequences samples positive for COVID-19 by RT-PCR with a Ct value of ≤ 30 for diagnostic purposes, VA SeqCURE laboratories sequence more broadly for nondiagnostic purposes, including samples with a Ct value > 30. The 6 VA SeqCURE laboratories generate sequencing data using various platforms, amplification kits, and formats. To ensure maximum quality and metadata on the sequences generated across the different laboratories, a sequence intake pipeline has been developed, adapting the ViralRecon bioinformatics platform.10 This harmonized analysis pipeline accommodates different file formats and performs quality control, alignment, variant calling, lineage assignment, clade assignment, and annotation. As of August 2023, VA SeqCURE has identified viral sequences from 24,107 unique specimens. Annotated COVID-19 sequences with the appropriate metadata will be available to VA researchers through VA SHIELD.
Research projects include descriptive epidemiology of COVID-19 variants in individuals who receive VHA care, COVID-19 vaccine and therapy effectiveness, and the unique distribution of variants and vaccine effectiveness in rural settings.3 True to its core mission, members of the VA SeqCURE consortium have contributed to the COVID-19 viral sequencing literature over the past 2 years. Researchers also are accessing VA SeqCURE to study COVID-19 persistence and rebound among individuals with mild disease taking nirmatrelvir/ritonavir compared with other COVID-19 therapeutics and untreated controls. Finally, COVID-19 samples and their sequences are stored in the VA SHIELD biorepository, which leverages these samples and data to advance scientific understanding of COVID-19 and future emerging infectious diseases.7-9
Important work from investigators at the Central Texas Veterans Health Care System confronted the issue of whole genome sequencing data from COVID-19 samples with low viral loads, a common issue with COVID-19 sequencing. They found that yields of 2 sequencing protocols, which generated high-sequence coverage, were enhanced further by combining the results of both methods.11 This project, which has potentially broad applications for sequencing in research and clinical settings, is an example of VA SeqCURE’s efforts to address the COVID-19 pandemic. The VA SeqCURE program has substantial potential as a large viral sequencing repository with broad geographic and demographic representation, such that future large-scale sequencing analyses may be generated from preexisting nested cohorts within the repository.
NEXT STEPS
Promising new directions of clinical and laboratory-based research are planned for VA SeqFORCE and VA SeqCURE. While the impact of COVID-19 and other viruses with epidemic potential is perhaps most feared in urban settings, evidence suggests that the distribution of COVID-19 in rural settings is unique and associated with worse outcomes.12,13 Given the wide catchment areas of VA hospitals that encompass both rural and urban settings, the VA’s ongoing COVID-19 sequencing programs and repositories are uniquely positioned to understand viral dynamics in areas of differing population density.
While rates of infection, hospitalization, and death resulting from COVID-19 have substantially dropped, the long-term impact of the pandemic is just beginning to be recognized in conditions such as long COVID or postacute COVID-19 syndrome. Long COVID has already proven to be biologically multifaceted, difficult to diagnose, and unpredictable in identifying the most at-risk patients.14-16 Much remains to be determined in our understanding of long COVID, including a unified definition that can effectively be used in clinical settings to diagnose and treat patients. However, research indicates that comorbidities common in veterans, such as diabetes and cardiovascular disease, are associated with worse long-term outcomes.17,18 Collaborations between VA scientists, clinicians, and national cooperative programs (such as a network of VHA long COVID clinics) create an unmatched opportunity for VA SeqFORCE and VA SeqCURE programs to provide insight into a disease likely to become a chronic disease outcome of the pandemic.
With VA SeqFORCE and VA SeqCURE programs, the VA now has infrastructure ready to respond to new infectious diseases. During the mpox outbreak of 2022, the VA Public Health Reference Laboratory received > 80% of all VA mpox samples for orthopox screening and mpox confirmatory testing. A subset of these samples underwent whole genome sequencing with the identification of 10 unique lineages across VA, and > 200 positive and 400 negative samples have been aliquoted and submitted to VA SHIELD for research. Furthermore, the VA SeqFORCE and VA SeqCURE sequencing processes might be adapted to identify outbreaks of multidrug-resistant organisms among VA patients trialed at other institutions.19 We are hopeful that VA SeqFORCE and VA SeqCURE will become invaluable components of health care delivery and infection prevention at the hospital level and beyond.
Finally, the robust data infrastructure and associated repositories of VA SeqFORCE and VA SeqCURE may be leveraged to study noninfectious diseases. Research groups are starting to apply these programs to cancer sequencing. We anticipate that these efforts may have a substantial impact on our understanding of cancer epidemiology and region-specific risk factors for malignancy, given the size and breadth of VA SeqFORCE and VA SeqCURE. Common oncogenic mutations identified through these programs could be targets for precision oncology therapeutics. Similarly, we envision applications of the VA SeqFORCE and VA SeqCURE data infrastructures and repositories toward other precision medicine fields, including pharmacogenomics and nutrition, to tailor interventions to meet the specific individual needs of veterans.
CONCLUSIONS
The productivity of VA SeqFORCE and VA SeqCURE programs over the past 2 years continues to increase in response to the COVID-19 pandemic. We anticipate that they will be vital components in our nation’s responses to infectious threats and beyond.
The COVID-19 virus and its associated pandemic have highlighted the urgent need for a national infrastructure to rapidly identify and respond to emerging pathogens. The importance of understanding viral population dynamics through genetic sequencing has become apparent over time, particularly as the vaccine responses, clinical implications, and therapeutic effectiveness of treatments have varied substantially with COVID-19 variants.1,2
As the largest integrated health care system in the US, the US Department of Veterans Affairs (VA) is uniquely situated to help with pandemic detection and response. This article highlights 2 VA programs dedicated to COVID-19 sequencing at the forefront of pandemic response and research: VA Sequencing for Research Clinical and Epidemiology (SeqFORCE) and VA Sequencing Collaborations United for Research and Epidemiology (SeqCURE) (Table).
VA Seq FORCE
VA SeqFORCE was established March 2021 to facilitate clinical surveillance of COVID-19 variants in the US veteran population and in VA employees. VA SeqFORCE consists of 9 Clinical Laboratory Improvement Amendment (CLIA)–certified laboratories in VA medical centers, including the VA Public Health Reference Laboratory in Palo Alto, California, and 8 Veterans Health Administration (VHA) clinical laboratories (Los Angeles, California; Boise, Idaho; Iowa City, Iowa; Bronx, New York; West Haven, Connecticut; Indianapolis, Indiana; Denver, Colorado; and Orlando, Florida).3 Specimen standards (eg, real-time polymerase chain reaction [RT-PCR] cycle threshold [Ct] ≤ 30, minimum volume, etc) and clinical criteria (eg, COVID-19–related deaths, COVID-19 vaccine escape, etc) for submitting samples to VA SeqFORCE laboratories were established, and logistics for sample sequencing was centralized, including providing centralized instructions for sample preparation and to which VA SeqFORCE laboratory samples should be sent.
These laboratories sequenced samples from patients and employees with COVID-19 to understand patterns of variant evolution, vaccine, antiviral and monoclonal antibody response, health care–associated outbreaks, and COVID-19 transmission. As clinically relevant findings, such as monoclonal antibody treatment failure, emerged with novel viral variants, VA SeqFORCE was well positioned to rapidly detect the emergent variants and inform better clinical care of patients with COVID-19. Other clinical indications identified for sequencing within VA SeqFORCE included outbreak investigation, re-infection with COVID-19 > 90 days but < 6 months after a prior infection, extended hospitalization of > 21 days, death due to COVID-19, infection with a history of recent nondomestic travel, rebound of symptoms after improvement on oral antiviral therapy, and epidemiologic surveillance.
VA SeqFORCE laboratories use a variety of sequencing platforms, although a federated system was developed that electronically linked all laboratories using a software system (PraediGene, Bitscopic) for sample management, COVID-19 variant analytics, and automated result reporting of clade and lineage into the Veterans Health Information Systems and Technology Architecture (VistA) Computerized Patient Record System. In addition, generated nucleic acid sequence alignment through FASTA consensus sequence files have been archived for secondary research analyses. By archiving the consensus sequences, retrospective studies within the VA have the added benefit of being able to clinically annotate investigations into COVID-19 variant patterns. As of August 2023, 43,003 samples containing COVID-19 have been sequenced, and FASTA file and metadata upload are ongoing to the Global Initiative on Sharing Avian Influenza Data, which houses > 15 million COVID-19 files from global submissions.
VA SeqFORCE’s clinical sequencing efforts have created opportunities for multicenter collaboration in variant surveillance. In work from December 2021, investigators from the James J. Peters VA Medical Center in Bronx, New York, collaborated with the VHA Pathology and Laboratory Medicine Services and Public Health national program offices in Washington, DC, to develop an RT-PCR assay to rapidly differentiate Omicron from Delta variants.4 Samples from VA hospitals across the nation were used in this study.
Lessons from VA SeqFORCE have also been cited as inspiration to address COVID-19 clinical problems, including outbreak investigations in hospital settings and beyond. Researchers at the Iowa City VA Health Care System, for example, proposed a novel probabilistic quantitative method for determining genetic-relatedness among COVID-19 viral strains in an outbreak setting.5 They extended the scope of work to develop COVID-19 outbreak screening tools combining publicly available algorithms with targeted sequencing data to identify outbreaks as they arise.6 We expect VA SeqFORCE, in conjunction with its complement VA SeqCURE, will continue to further pandemic surveillance and response.
VA Seq CURE
As the research-focused complement to VA SeqFORCE, VA SeqCURE is dedicated to a broader study of the COVID-19 genome through sequencing. Established January 2021, the VA SeqCURE network consists of 6 research laboratories in Boise, Idaho; Bronx, New York; Cleveland, Ohio; Durham, North Carolina; Iowa City, Iowa; and Temple, Texas.
Samples are collected as a subset of the broader VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD) biorepository sweep protocol for discarded blood and nasal swab specimens of VHA patients hospitalized with COVID-19, as described by Epstein and colleagues.7-9 While VA SeqFORCE sequences samples positive for COVID-19 by RT-PCR with a Ct value of ≤ 30 for diagnostic purposes, VA SeqCURE laboratories sequence more broadly for nondiagnostic purposes, including samples with a Ct value > 30. The 6 VA SeqCURE laboratories generate sequencing data using various platforms, amplification kits, and formats. To ensure maximum quality and metadata on the sequences generated across the different laboratories, a sequence intake pipeline has been developed, adapting the ViralRecon bioinformatics platform.10 This harmonized analysis pipeline accommodates different file formats and performs quality control, alignment, variant calling, lineage assignment, clade assignment, and annotation. As of August 2023, VA SeqCURE has identified viral sequences from 24,107 unique specimens. Annotated COVID-19 sequences with the appropriate metadata will be available to VA researchers through VA SHIELD.
Research projects include descriptive epidemiology of COVID-19 variants in individuals who receive VHA care, COVID-19 vaccine and therapy effectiveness, and the unique distribution of variants and vaccine effectiveness in rural settings.3 True to its core mission, members of the VA SeqCURE consortium have contributed to the COVID-19 viral sequencing literature over the past 2 years. Researchers also are accessing VA SeqCURE to study COVID-19 persistence and rebound among individuals with mild disease taking nirmatrelvir/ritonavir compared with other COVID-19 therapeutics and untreated controls. Finally, COVID-19 samples and their sequences are stored in the VA SHIELD biorepository, which leverages these samples and data to advance scientific understanding of COVID-19 and future emerging infectious diseases.7-9
Important work from investigators at the Central Texas Veterans Health Care System confronted the issue of whole genome sequencing data from COVID-19 samples with low viral loads, a common issue with COVID-19 sequencing. They found that yields of 2 sequencing protocols, which generated high-sequence coverage, were enhanced further by combining the results of both methods.11 This project, which has potentially broad applications for sequencing in research and clinical settings, is an example of VA SeqCURE’s efforts to address the COVID-19 pandemic. The VA SeqCURE program has substantial potential as a large viral sequencing repository with broad geographic and demographic representation, such that future large-scale sequencing analyses may be generated from preexisting nested cohorts within the repository.
NEXT STEPS
Promising new directions of clinical and laboratory-based research are planned for VA SeqFORCE and VA SeqCURE. While the impact of COVID-19 and other viruses with epidemic potential is perhaps most feared in urban settings, evidence suggests that the distribution of COVID-19 in rural settings is unique and associated with worse outcomes.12,13 Given the wide catchment areas of VA hospitals that encompass both rural and urban settings, the VA’s ongoing COVID-19 sequencing programs and repositories are uniquely positioned to understand viral dynamics in areas of differing population density.
While rates of infection, hospitalization, and death resulting from COVID-19 have substantially dropped, the long-term impact of the pandemic is just beginning to be recognized in conditions such as long COVID or postacute COVID-19 syndrome. Long COVID has already proven to be biologically multifaceted, difficult to diagnose, and unpredictable in identifying the most at-risk patients.14-16 Much remains to be determined in our understanding of long COVID, including a unified definition that can effectively be used in clinical settings to diagnose and treat patients. However, research indicates that comorbidities common in veterans, such as diabetes and cardiovascular disease, are associated with worse long-term outcomes.17,18 Collaborations between VA scientists, clinicians, and national cooperative programs (such as a network of VHA long COVID clinics) create an unmatched opportunity for VA SeqFORCE and VA SeqCURE programs to provide insight into a disease likely to become a chronic disease outcome of the pandemic.
With VA SeqFORCE and VA SeqCURE programs, the VA now has infrastructure ready to respond to new infectious diseases. During the mpox outbreak of 2022, the VA Public Health Reference Laboratory received > 80% of all VA mpox samples for orthopox screening and mpox confirmatory testing. A subset of these samples underwent whole genome sequencing with the identification of 10 unique lineages across VA, and > 200 positive and 400 negative samples have been aliquoted and submitted to VA SHIELD for research. Furthermore, the VA SeqFORCE and VA SeqCURE sequencing processes might be adapted to identify outbreaks of multidrug-resistant organisms among VA patients trialed at other institutions.19 We are hopeful that VA SeqFORCE and VA SeqCURE will become invaluable components of health care delivery and infection prevention at the hospital level and beyond.
Finally, the robust data infrastructure and associated repositories of VA SeqFORCE and VA SeqCURE may be leveraged to study noninfectious diseases. Research groups are starting to apply these programs to cancer sequencing. We anticipate that these efforts may have a substantial impact on our understanding of cancer epidemiology and region-specific risk factors for malignancy, given the size and breadth of VA SeqFORCE and VA SeqCURE. Common oncogenic mutations identified through these programs could be targets for precision oncology therapeutics. Similarly, we envision applications of the VA SeqFORCE and VA SeqCURE data infrastructures and repositories toward other precision medicine fields, including pharmacogenomics and nutrition, to tailor interventions to meet the specific individual needs of veterans.
CONCLUSIONS
The productivity of VA SeqFORCE and VA SeqCURE programs over the past 2 years continues to increase in response to the COVID-19 pandemic. We anticipate that they will be vital components in our nation’s responses to infectious threats and beyond.
1. Iuliano AD, Brunkard JM, Boehmer TK, et al. Trends in disease severity and health care utilization during the early Omicron variant period compared with previous SARS-CoV-2 high transmission periods - United States, December 2020-January 2022. MMWR Morb Mortal Wkly Rep. 2022;71(4):146-152. Published 2022 Jan 28. doi:10.15585/mmwr.mm7104e4
2. Nyberg T, Ferguson NM, Nash SG, et al. Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. Lancet. 2022;399(10332):1303-1312. doi:10.1016/S0140-6736(22)00462-7
3. Veterans Health Administration. Coronavirus Disease 2019 (COVID-19) response report - annex C. December 5, 2022. Accessed August 28, 2023. https://www.va.gov/HEALTH/docs/VHA-COVID-19-Response-2022-Annex-C.pdf 4. Barasch NJ, Iqbal J, Coombs M, et al. Utilization of a SARS-CoV-2 variant assay for the rapid differentiation of Omicron and Delta. medRxiv. Preprint posted online December 27, 2021. doi:10.1101/2021.12.22.21268195
5. Bilal MY. Similarity Index-probabilistic confidence estimation of SARS-CoV-2 strain relatedness in localized outbreaks. Epidemiologia (Basel). 2022;3(2):238-249. doi:10.3390/epidemiologia3020019
6. Bilal MY, Klutts JS. Molecular Epidemiological investigations of localized SARS-CoV-2 outbreaks-utility of public algorithms. Epidemiologia (Basel). 2022;3(3):402-411. doi:10.3390/epidemiologia3030031
7. Veterans Health Administration, Office of Research & Development. VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). Updated November 23, 2022. Accessed August 28, 2023. https://www.research.va.gov/programs/shield/about.cfm
8. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
9. Epstein L, Shive C, Garcia AP, et al. VA SHIELD: a biorepository for our veterans and the nation. Fed Pract. 2023;40(suppl 5):S48-S51. doi:10.12788/fp.0424
10. Patel H, Varona S, Monzón S, et al. Version 2.5. nf-core/viralrecon: nf-core/viralrecon v2.5 - Manganese Monkey (2.5). Zenodo. July 13, 2022. doi:10.5281/zenodo.6827984
11. Choi H, Hwang M, Navarathna DH, Xu J, Lukey J, Jinadatha C. Performance of COVIDSeq and swift normalase amplicon SARS-CoV-2 panels for SARS-CoV-2 genome sequencing: practical guide and combining FASTQ strategy. J Clin Microbiol. 2022;60(4):e0002522. doi:10.1128/jcm.00025-22
12. Cuadros DF, Branscum AJ, Mukandavire Z, Miller FD, MacKinnon N. Dynamics of the COVID-19 epidemic in urban and rural areas in the United States. Ann Epidemiol. 2021;59:16-20. doi:10.1016/j.annepidem.2021.04.007
13. Anzalone AJ, Horswell R, Hendricks BM, et al. Higher hospitalization and mortality rates among SARS-CoV-2-infected persons in rural America. J Rural Health. 2023;39(1):39-54. doi:10.1111/jrh.12689
14. Su Y, Yuan D, Chen DG, et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell. 2022;185(5):881-895.e20. doi:10.1016/j.cell.2022.01.014
15. Pfaff ER, Girvin AT, Bennett TD, et al. Identifying who has long COVID in the USA: a machine learning approach using N3C data. Lancet Digit Health. 2022;4(7):e532-e541. doi:10.1016/S2589-7500(22)00048-6
16. Subramanian A, Nirantharakumar K, Hughes S, et al. Symptoms and risk factors for long COVID in non-hospitalized adults. Nat Med. 2022;28(8):1706-1714. doi:10.1038/s41591-022-01909-w
17. Munblit D, O’Hara ME, Akrami A, Perego E, Olliaro P, Needham DM. Long COVID: aiming for a consensus. Lancet Respir Med. 2022;10(7):632-634. doi:10.1016/S2213-2600(22)00135-7
18. Thaweethai T, Jolley SE, Karlson EW, et al. Development of a definition of postacute sequelae of SARS-CoV-2 infection. JAMA. 2023;329(22):1934-1946. doi:10.1001/jama.2023.8823
19. Sundermann AJ, Chen J, Kumar P, et al. Whole-genome sequencing surveillance and machine learning of the electronic health record for enhanced healthcare outbreak detection. Clin Infect Dis. 2022;75(3):476-482. doi:10.1093/cid/ciab946
1. Iuliano AD, Brunkard JM, Boehmer TK, et al. Trends in disease severity and health care utilization during the early Omicron variant period compared with previous SARS-CoV-2 high transmission periods - United States, December 2020-January 2022. MMWR Morb Mortal Wkly Rep. 2022;71(4):146-152. Published 2022 Jan 28. doi:10.15585/mmwr.mm7104e4
2. Nyberg T, Ferguson NM, Nash SG, et al. Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. Lancet. 2022;399(10332):1303-1312. doi:10.1016/S0140-6736(22)00462-7
3. Veterans Health Administration. Coronavirus Disease 2019 (COVID-19) response report - annex C. December 5, 2022. Accessed August 28, 2023. https://www.va.gov/HEALTH/docs/VHA-COVID-19-Response-2022-Annex-C.pdf 4. Barasch NJ, Iqbal J, Coombs M, et al. Utilization of a SARS-CoV-2 variant assay for the rapid differentiation of Omicron and Delta. medRxiv. Preprint posted online December 27, 2021. doi:10.1101/2021.12.22.21268195
5. Bilal MY. Similarity Index-probabilistic confidence estimation of SARS-CoV-2 strain relatedness in localized outbreaks. Epidemiologia (Basel). 2022;3(2):238-249. doi:10.3390/epidemiologia3020019
6. Bilal MY, Klutts JS. Molecular Epidemiological investigations of localized SARS-CoV-2 outbreaks-utility of public algorithms. Epidemiologia (Basel). 2022;3(3):402-411. doi:10.3390/epidemiologia3030031
7. Veterans Health Administration, Office of Research & Development. VA Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD). Updated November 23, 2022. Accessed August 28, 2023. https://www.research.va.gov/programs/shield/about.cfm
8. Harley JB, Pyarajan S, Partan ES, et al. The US Department of Veterans Affairs Science and Health Initiative to Combat Infectious and Emerging Life-Threatening Diseases (VA SHIELD): a biorepository addressing national health threats. Open Forum Infect Dis. 2022;9(12):ofac641. doi:10.1093/ofid/ofac641
9. Epstein L, Shive C, Garcia AP, et al. VA SHIELD: a biorepository for our veterans and the nation. Fed Pract. 2023;40(suppl 5):S48-S51. doi:10.12788/fp.0424
10. Patel H, Varona S, Monzón S, et al. Version 2.5. nf-core/viralrecon: nf-core/viralrecon v2.5 - Manganese Monkey (2.5). Zenodo. July 13, 2022. doi:10.5281/zenodo.6827984
11. Choi H, Hwang M, Navarathna DH, Xu J, Lukey J, Jinadatha C. Performance of COVIDSeq and swift normalase amplicon SARS-CoV-2 panels for SARS-CoV-2 genome sequencing: practical guide and combining FASTQ strategy. J Clin Microbiol. 2022;60(4):e0002522. doi:10.1128/jcm.00025-22
12. Cuadros DF, Branscum AJ, Mukandavire Z, Miller FD, MacKinnon N. Dynamics of the COVID-19 epidemic in urban and rural areas in the United States. Ann Epidemiol. 2021;59:16-20. doi:10.1016/j.annepidem.2021.04.007
13. Anzalone AJ, Horswell R, Hendricks BM, et al. Higher hospitalization and mortality rates among SARS-CoV-2-infected persons in rural America. J Rural Health. 2023;39(1):39-54. doi:10.1111/jrh.12689
14. Su Y, Yuan D, Chen DG, et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell. 2022;185(5):881-895.e20. doi:10.1016/j.cell.2022.01.014
15. Pfaff ER, Girvin AT, Bennett TD, et al. Identifying who has long COVID in the USA: a machine learning approach using N3C data. Lancet Digit Health. 2022;4(7):e532-e541. doi:10.1016/S2589-7500(22)00048-6
16. Subramanian A, Nirantharakumar K, Hughes S, et al. Symptoms and risk factors for long COVID in non-hospitalized adults. Nat Med. 2022;28(8):1706-1714. doi:10.1038/s41591-022-01909-w
17. Munblit D, O’Hara ME, Akrami A, Perego E, Olliaro P, Needham DM. Long COVID: aiming for a consensus. Lancet Respir Med. 2022;10(7):632-634. doi:10.1016/S2213-2600(22)00135-7
18. Thaweethai T, Jolley SE, Karlson EW, et al. Development of a definition of postacute sequelae of SARS-CoV-2 infection. JAMA. 2023;329(22):1934-1946. doi:10.1001/jama.2023.8823
19. Sundermann AJ, Chen J, Kumar P, et al. Whole-genome sequencing surveillance and machine learning of the electronic health record for enhanced healthcare outbreak detection. Clin Infect Dis. 2022;75(3):476-482. doi:10.1093/cid/ciab946
Leveraging the Million Veteran Program Infrastructure and Data for a Rapid Research Response to COVID-19
The Million Veteran Program (MVP) was launched in 2011 by the US Department of Veterans Affairs (VA) to enroll at least 1 million veterans in a longitudinal cohort to better understand how genes, lifestyle, military experience, and environmental exposures interact to influence health and illness and ultimately enable precision health care. The MVP has established a national, centralized infrastructure for recruitment and enrollment, biospecimen and data collection and storage, data generation and curation, and secure data access. When the COVID-19 pandemic hit in 2020, the MVP was leveraged to support research utilizing the following key infrastructure components: (1) MVP recruitment and enrollment platform to provide support for COVID-19 vaccine and treatment trials and to collect COVID-19 data from MVP participants; (2) using MVP Phenomics for COVID-19 research data cleaning and curation, assisting with the development of a VA Severity Index for COVID-19, and forming 6 scientific working groups to coordinate COVID-19 research questions; and (3) the VA/MVP and US Department of Energy (DOE) partnership to assist in responding to COVID-19 research questions identified by the US Food and Drug Administration (FDA). This article describes these infrastructure components in more detail and highlights key findings from the MVP COVID-19 research efforts.
MVP Infrastructure
The Veterans Health Administration (VHA) Office of Research and Development (ORD) oversaw efforts to develop the VA Coronavirus Research Volunteer List (the COVID-19 registry). To support the registry, the MVP leveraged its infrastructure to facilitate a rapid response. The MVP is designed as a full-service and centralized recruitment and enrollment platform. This includes MVP office oversight; MVP coordinating centers that manage the centralized platform; an information center that handles inbound and outbound calls; an informatics system built for recruitment and enrollment monitoring and tracking; and a network of more than 70 participating MVP sites with dedicated staff to conduct recruitment and enrollment activities. The MVP used its informatics infrastructure to support secure data storage for the registry volunteer information. MVP coordinating center staff worked with the COVID-19 registry to invite > 125,000 MVP participants from approximately 20 MVP sites. Additionally, MVP information center staff made > 4000 calls to prospective registry volunteers. This work resulted in 1300 volunteers agreeing to be
New Data Collection
The MVP protocol was approved by the VA Central Institutional Review Board (IRB) in 2011. As part of initial enrollment in MVP, participants consented to recontact for additional self-report information along with access to their electronic health record (EHR). This allows for the linkage of EHR and survey response data, thus providing a comprehensive understanding of health history before and after a self-reported COVID-19 diagnosis. Between May 2020 and September 2021, the MVP COVID-19 survey was distributed to existing MVP participants via mail, telephone, and email with the ability to complete the survey by paper and pencil or through the MVP online system. Dissemination of the survey was approved by the VA Central IRB in 2020, with nearly 730,000 eligible MVP participants contacted. As of June 2022, 255,737 MVP participants (35% of the eligible cohort) had completed the survey; 86% completed a paper survey while 14% completed it online. Respondents were primarily older (≥ 65 years); 90% were male; close to 7% reported Hispanic ethnicity, and 11% reported Black race.
Findings from this survey provide insight into pandemic behaviors not consistently captured in EHRs, such as psychosocial aspects, including social and emotional support, loss of tangible and intangible resources, as well as COVID-19–related behaviors, such as social distancing and self-protective practices.1 MVP COVID-19 survey data combined with veteran EHRs, responses to other MVP surveys, and genetic data enable MVP researchers to better understand epidemiological, clinical, and psychosocial aspects of the disease. Future COVID-19 studies may use self-reported survey responses to enrich understanding about the effects of the disease on a veteran’s daily life, and possibly validate existing EHR COVID-19 diagnoses and hospitalization findings. This comprehensive data resource provides a unique opportunity to identify new targets for disease prevention, treatment, and management with an emphasis on individual variability in genes, environment, and lifestyle.
COVID-19 Research
In early 2020, the burden of COVID-19 on the US was unprecedented, and little was known about risk factors for severe COVID-19 and deaths. The MVP Phenomics team quickly responded with a large-scale phenome-wide association study (PheWAS) of >
To broaden disease progression data curation and fit the specific needs of the VA, we operationalized and validated the World Health Organization clinical severity scale and used VA EHR data to create the VA Severity Index for COVID-19 (VASIC).3 The VASIC category is now part of the MVP core data repository, where volumes of data from multiple activities are integrated through an automated process to create monthly research-ready data cubes. These activities include extensive data curation, mapping, phenotyping, and adjudication that are performed to curate oxygen supplementation status and other procedures related to treatment that are processed and understood in real time. The data cubes were provisioned to MVP COVID-19 researchers. In addition, the VASIC scale variable is now integrated within the larger VA system for all researchers to use as part of its wider COVID-19 initiative. The VA Centralized Interactive Phenomics Resource (CIPHER) phenomics library now hosts the details of VASIC, codes, metadata, and related COVID-19 data products for all VA communities. In partnership with CIPHER and other internal and external COVID-19 initiatives, the MVP continues to play an integral part for the VA and beyond in the development of a phenomics algorithm for long COVID, or post-acute COVID-19 syndrome (PACS).
Host Genetics in COVID-19
As the SARS-CoV-2 virus continued to spread globally, it became clear that the symptoms and severity of infection experienced by patients varied across a broad spectrum, from being asymptomatic carriers to experiencing severe symptoms in 1 or more organ systems in the body, resulting in death. This variability suggested that host genetics and other host factors may play a role in determining the severity of COVID-19 infection. The MVP dataset, with genetic and health information on > 600,000 MVP participants, provided an ideal dataset to explore host contributions to COVID-19.
In late spring 2020, the MVP executive committee issued a call to the MVP research community to propose study aims around the COVID-19 pandemic that could leverage the phenotypic and genetic data and resources. The MVP quickly formed 6 rapid-response scientific working groups. Their mission was to cultivate collaboration and inclusivity and to coordinate COVID-19 research questions. A steering committee composed of the MVP executive committee, staff from computational environments, working group cochairs, and an administrator, who was responsible for daily oversight of the working groups. In addition, the ORD COVID-19 steering committee reviewed and approved research activities to ensure scientific rigor, as well as alignment with overall ongoing research activities.
The MVP COVID-19 working groups included dozens of researchers who used MVP data to identify disease mechanisms; understand the impact of host genetics on susceptibility, morbidity, and mortality; and identify potential targets for treatments and therapies. The working groups were further supported by MVP analysts to work cross-functionally on genomics, phenomics, statistical genetics, and PheWAS. Each working group chair was responsible for prioritizing concepts and moving them forward in coordination with the MVP and ORD COVID-19 steering committees. An overview of the MVP COVID-19 working groups follows (Table).4-9
Druggable genome. This working group researched drug-repurposing opportunities to prevent severe COVID-19, defined as hospitalization with oxygen therapy (high flow), intubation, mechanical ventilation, vasopressors, dialysis, or death from COVID-19; and prevent complications in patients hospitalized by COVID-19.
Pharmacogenomics. This working group focused on 2 main aims: the impact of apolipoprotein L1 risk variants on acute kidney injury (AKI) and death in Black veterans with COVID-19; and pharmacogenetic analysis of remdesivir-induced liver chemistry abnormalities.
Disease mechanisms. Understanding the underlying pathways and mechanisms behind COVID-19 has been a difficult but important challenge overall in the scientific community. This working group investigated specific genetic markers and effects on COVID-19, including polygenic predisposition to venous thromboembolism associated with increased COVID-19 susceptibility; renal comorbidities and new AKI and unfavorable outcomes among COVID-19–positive sickle cell trait carriers; and mucin 5B, oligomeric mucus/gel-forming gene polymorphism, and protective effects in COVID-19 infection.
Genomics for risk prediction, polygenic risk scores, and mendelian randomization. Risk prediction for COVID-19 has been widely studied mostly aiming at comorbidities and preexisting conditions. The MVP cohort provided a unique opportunity to understand how genetic information can enhance our understanding of COVID-19 risk. This working group focused on: (1) ABO blood group typing and the protective effects of the O blood group on COVID-19 infection; (2) polygenic risk scores and COVID-19 outcomes; (3) human leukocyte antigen typing and COVID-19 outcomes; and (4) a transcriptome-wide association study of COVID-19–positive MVP participants.
Genome-Wide Association Study (GWAS) and Downstream Analysis. This working group performed GWAS of the main COVID-19 outcomes. Results from GWAS unveiled new genetic loci to suggest further investigation on these candidate genes. The results were used by other MVP COVID-19 working groups for their activities. The results also contributed to external collaborations, such as the COVID-19 Host Genetics Initiative.
COVID-19–Related PheWAS. This working group focused on understanding the potential clinical significance of genetic variants associated with susceptibility to, or outcomes of, COVID-19 infection. They worked to identify traits that share genetic variants associated with severe COVID-19 from the Host Genetics Initiative. The group also studied the phenotypic consequences of acquired mosaic chromosomal alterations with early data linking to COVID-19 susceptibility.
COVID-19 Research Partnerships
In 2016, the VA and DOE formed an interagency partnership known as Computational Health Analytics for Medical Precision to Improve Outcomes Now (CHAMPION) to demonstrate the power of combining the VA EHR system, MVP genetic data, and clinical research expertise with DOE high-performance computing infrastructure and artificial intelligence expertise. The VA EHR captures longitudinal care information on veterans with records that go back decades. Furthermore, the VA covers the costs of medications and
The DOE Oak Ridge National Laboratory (ORNL) in Tennessee securely maintains this rich database for the VA. The ORNL Summit supercomputer can complete trillions of calculations per second to provide critical and timely analyses, applying the most advanced and powerful artificial intelligence methods, which would not be possible in more conventional research settings. CHAMPION taught the VA and DOE how to bring their disparate research cultures together for innovative collaborative investigation. Moreover, this collaboration produced a cadre of VA and DOE scientists familiar with VA patient data and experienced in conducting joint research successfully and integrating omics data with clinical data for a better mechanistic understanding. Because of this preexisting collaboration between the VA and DOE, interagency teams were prepared at the start of the COVID-19 pandemic.10-15
Other recently completed studies have developed and validated short-term mortality indices in individuals with COVID-19 based on their preexisting conditions, assessed the generalizability of VA COVID-19 experiences to the US population, and evaluated the effectiveness of hydroxychloroquine with and without azithromycin in VA patients with COVID-19.12,15 A recent study demonstrated the benefit of prophylactic anticoagulation at initial hospitalization.14
The VA also provided the FDA with daily reports on aggregate VA COVID-19 cases and their distribution across the VA system, demographics of VA patients with COVID-19, and analyses of predictive models for positive test results and death. The VA regularly sent the FDA aggregated data showing patterns of medication use and retrospective analyses of the effectiveness of certain medications (including remdesivir and some antithrombotic agents). The FDA used these data along with other data to understand the scope of the pandemic and to predict drug shortages or needs for additional medical equipment, including ventilators.
Limitations
For the most part, MVP infrastructure and partnerships were efficiently leveraged to significantly advance our understanding of the biological basis of COVID-19 and to develop treatments and vaccines. However, there were a few limitations that may have slowed timely and optimal outcomes. An issue not limited to the MVP or VA was the continual evolution of the pandemic and its response. This included evolving definitions of disease, symptomatology, testing, vaccines, and public health recommendations. Keeping pace with the emerging knowledge from these domains was a struggle for the entire scientific community. A more discrete limitation was the number of participants in the MVP with positive COVID-19 test results and positive symptoms; however, this was mitigated by partnering with other groups like the COVID-19 Host Genetics Initiative to increase study participant numbers. Finally, there were logistical and regulatory challenges associated with coordination of national clinical trial recruitment across a VA system with > 100 discrete hospitals.
Conclusions
Having a centralized infrastructure for recruitment and enrollment, including a national research volunteer registry, information center, research staff, and coordinating centers, can allow for expedited enrollment in vaccine and treatment trials in the face of future public health emergencies. VA assets, including its rich EHR and MVP, the world’s largest genomic cohort, have contributed to improving our understanding and management of COVID-19.
1. Whitbourne SB, Nguyen XT, Song RJ, et al. Million Veteran Program’s response to COVID-19: survey development and preliminary findings. PLoS One. 2022;17(4):e0266381. doi:10.1371/journal.pone.0266381
2. Song RJ, Ho YL, Schubert P, et al. Phenome-wide association of 1809 phenotypes and COVID-19 disease progression in the Veterans Health Administration Million Veteran Program. PLoS One. 2021;16(5):e0251651. doi:10.1371/journal.pone.0251651
3. Galloway A, Park Y, Tanukonda V, et al. Impact of COVID-19 severity on long-term events in US veterans using the Veterans Affairs Severity Index for COVID-19 (VASIC). J Infect Dis. 2022;226(12):2113-2117. doi:10.1093/infdis/jiac182
4. Gaziano L, Giambartolomei C, Pereira AC, et al. Actionable druggable genome-wide Mendelian randomization identifies repurposing opportunities for COVID-19. Nat Med. 2021;27(4):668-676. doi:10.0138/s41591-021-01310-z
5. Hung AM, Sha SC, Bick AG, et al. APOL1 risk variants, acute kidney injury, and death in participants with African ancestry hospitalized with COVID-19 from the Million Veteran Program. JAMA Intern Med. 2022;182(4):386-395. doi:10.1001/jamainternmed.2021.8538
6. Verma A, Huffman JE, Gao L, et al. Association of kidney comorbidities and acute kidney failure with unfavorable outcomes after COVID-19 in individuals with the sickle cell trait. JAMA Intern Med. 2022;182(8):796-804. doi:10.1001/jamainternmed.2022.2141
7. Verma A, Tsao NL, Thomann LO, et al. A phenome-wide association study of genes associated with COVID-19 severity reveals shared genetics with complex diseases in the Million Veteran Program. PLoS Genet. 2022;18(4):e1010113. doi:10.1371/journal.pgen.1010113
8. Peloso GM, Tcheandjieu C, McGeary JE, et al. Genetic loci associated with COVID-19 positivity and hospitalization in White, Black, and Hispanic Veterans of the VA Million Veteran Program. Front Genetic. 2022;12:777076. doi:10.3389/fgene.2021.777076
9. Verma A, Minnier J, Wan ES, et al. A MUC5B gene polymorphism, rs35705950-T confers protective effects against COVID-19 hospitalization but not severe disease or mortality. Am J Respir Crit Care Med. 2022;182(8):796-804. doi:10.1164/rccm.202109-2166OC
10. Garvin MR, Alvarez C, Miller JI, et al. A mechanistic model and therapeutic interventions for COVID-19 involving a RAS-mediated bradykinin storm. Elife. 2020;e59177. doi:10.7554/eLife.59177
11. Rentsch CT, Kidwai-Khan F, Tate JP, et al. Patterns of COVID-19 testing and mortality by race and ethnicity among United States veterans: A nationwide cohort study. PLoS Med. 2020;17(9):e1003379. doi:10.1371/journal.pmed.1003379
12. King JT, Yoon JS, Rentsch CT, et al. Development and validation of a 30-day mortality index based on pre-existing medical administrative data from 13,323 COVID-19 patients: the Veterans Health Administration COVID-19 (VACO) Index. PLoS One. 2020;15(11):e0241825. doi:10.1371/journal.pone.0241825
13. Joubert W, Weighill D, Kainer D, et al. Attacking the opioid epidemic: determining the epistatic and pleiotropic genetic architectures for chronic pain and opioid addiction. SC18: International Conference for High Performance Computing, Networking, Storage and Analysis. Dallas, TX, USA, 2018:717-730. doi:10.1109/SC.2018.00060
14. Rentsch CT, Beckman JA, Tomlinson L, et al. Early initiation of prophylactic anticoagulation for prevention of COVID-19 mortality: a nationwide cohort study of hospitalized patients in the United States. BMJ. 2021;372:n311. doi:10.1136/bmj.n311
15. Gerlovin H, Posner DC, Ho YL, et al. Pharmacoepidemiology, machine learning, and COVID-19: an intent-to-treat analysis of hydroxychloroquine, with or without Azithromycin, and COVID-19 outcomes among hospitalized US Veterans. Am J Epidemiol. 2021;190(11): 2405-2419. doi:10.1093/aje/kwab183
The Million Veteran Program (MVP) was launched in 2011 by the US Department of Veterans Affairs (VA) to enroll at least 1 million veterans in a longitudinal cohort to better understand how genes, lifestyle, military experience, and environmental exposures interact to influence health and illness and ultimately enable precision health care. The MVP has established a national, centralized infrastructure for recruitment and enrollment, biospecimen and data collection and storage, data generation and curation, and secure data access. When the COVID-19 pandemic hit in 2020, the MVP was leveraged to support research utilizing the following key infrastructure components: (1) MVP recruitment and enrollment platform to provide support for COVID-19 vaccine and treatment trials and to collect COVID-19 data from MVP participants; (2) using MVP Phenomics for COVID-19 research data cleaning and curation, assisting with the development of a VA Severity Index for COVID-19, and forming 6 scientific working groups to coordinate COVID-19 research questions; and (3) the VA/MVP and US Department of Energy (DOE) partnership to assist in responding to COVID-19 research questions identified by the US Food and Drug Administration (FDA). This article describes these infrastructure components in more detail and highlights key findings from the MVP COVID-19 research efforts.
MVP Infrastructure
The Veterans Health Administration (VHA) Office of Research and Development (ORD) oversaw efforts to develop the VA Coronavirus Research Volunteer List (the COVID-19 registry). To support the registry, the MVP leveraged its infrastructure to facilitate a rapid response. The MVP is designed as a full-service and centralized recruitment and enrollment platform. This includes MVP office oversight; MVP coordinating centers that manage the centralized platform; an information center that handles inbound and outbound calls; an informatics system built for recruitment and enrollment monitoring and tracking; and a network of more than 70 participating MVP sites with dedicated staff to conduct recruitment and enrollment activities. The MVP used its informatics infrastructure to support secure data storage for the registry volunteer information. MVP coordinating center staff worked with the COVID-19 registry to invite > 125,000 MVP participants from approximately 20 MVP sites. Additionally, MVP information center staff made > 4000 calls to prospective registry volunteers. This work resulted in 1300 volunteers agreeing to be
New Data Collection
The MVP protocol was approved by the VA Central Institutional Review Board (IRB) in 2011. As part of initial enrollment in MVP, participants consented to recontact for additional self-report information along with access to their electronic health record (EHR). This allows for the linkage of EHR and survey response data, thus providing a comprehensive understanding of health history before and after a self-reported COVID-19 diagnosis. Between May 2020 and September 2021, the MVP COVID-19 survey was distributed to existing MVP participants via mail, telephone, and email with the ability to complete the survey by paper and pencil or through the MVP online system. Dissemination of the survey was approved by the VA Central IRB in 2020, with nearly 730,000 eligible MVP participants contacted. As of June 2022, 255,737 MVP participants (35% of the eligible cohort) had completed the survey; 86% completed a paper survey while 14% completed it online. Respondents were primarily older (≥ 65 years); 90% were male; close to 7% reported Hispanic ethnicity, and 11% reported Black race.
Findings from this survey provide insight into pandemic behaviors not consistently captured in EHRs, such as psychosocial aspects, including social and emotional support, loss of tangible and intangible resources, as well as COVID-19–related behaviors, such as social distancing and self-protective practices.1 MVP COVID-19 survey data combined with veteran EHRs, responses to other MVP surveys, and genetic data enable MVP researchers to better understand epidemiological, clinical, and psychosocial aspects of the disease. Future COVID-19 studies may use self-reported survey responses to enrich understanding about the effects of the disease on a veteran’s daily life, and possibly validate existing EHR COVID-19 diagnoses and hospitalization findings. This comprehensive data resource provides a unique opportunity to identify new targets for disease prevention, treatment, and management with an emphasis on individual variability in genes, environment, and lifestyle.
COVID-19 Research
In early 2020, the burden of COVID-19 on the US was unprecedented, and little was known about risk factors for severe COVID-19 and deaths. The MVP Phenomics team quickly responded with a large-scale phenome-wide association study (PheWAS) of >
To broaden disease progression data curation and fit the specific needs of the VA, we operationalized and validated the World Health Organization clinical severity scale and used VA EHR data to create the VA Severity Index for COVID-19 (VASIC).3 The VASIC category is now part of the MVP core data repository, where volumes of data from multiple activities are integrated through an automated process to create monthly research-ready data cubes. These activities include extensive data curation, mapping, phenotyping, and adjudication that are performed to curate oxygen supplementation status and other procedures related to treatment that are processed and understood in real time. The data cubes were provisioned to MVP COVID-19 researchers. In addition, the VASIC scale variable is now integrated within the larger VA system for all researchers to use as part of its wider COVID-19 initiative. The VA Centralized Interactive Phenomics Resource (CIPHER) phenomics library now hosts the details of VASIC, codes, metadata, and related COVID-19 data products for all VA communities. In partnership with CIPHER and other internal and external COVID-19 initiatives, the MVP continues to play an integral part for the VA and beyond in the development of a phenomics algorithm for long COVID, or post-acute COVID-19 syndrome (PACS).
Host Genetics in COVID-19
As the SARS-CoV-2 virus continued to spread globally, it became clear that the symptoms and severity of infection experienced by patients varied across a broad spectrum, from being asymptomatic carriers to experiencing severe symptoms in 1 or more organ systems in the body, resulting in death. This variability suggested that host genetics and other host factors may play a role in determining the severity of COVID-19 infection. The MVP dataset, with genetic and health information on > 600,000 MVP participants, provided an ideal dataset to explore host contributions to COVID-19.
In late spring 2020, the MVP executive committee issued a call to the MVP research community to propose study aims around the COVID-19 pandemic that could leverage the phenotypic and genetic data and resources. The MVP quickly formed 6 rapid-response scientific working groups. Their mission was to cultivate collaboration and inclusivity and to coordinate COVID-19 research questions. A steering committee composed of the MVP executive committee, staff from computational environments, working group cochairs, and an administrator, who was responsible for daily oversight of the working groups. In addition, the ORD COVID-19 steering committee reviewed and approved research activities to ensure scientific rigor, as well as alignment with overall ongoing research activities.
The MVP COVID-19 working groups included dozens of researchers who used MVP data to identify disease mechanisms; understand the impact of host genetics on susceptibility, morbidity, and mortality; and identify potential targets for treatments and therapies. The working groups were further supported by MVP analysts to work cross-functionally on genomics, phenomics, statistical genetics, and PheWAS. Each working group chair was responsible for prioritizing concepts and moving them forward in coordination with the MVP and ORD COVID-19 steering committees. An overview of the MVP COVID-19 working groups follows (Table).4-9
Druggable genome. This working group researched drug-repurposing opportunities to prevent severe COVID-19, defined as hospitalization with oxygen therapy (high flow), intubation, mechanical ventilation, vasopressors, dialysis, or death from COVID-19; and prevent complications in patients hospitalized by COVID-19.
Pharmacogenomics. This working group focused on 2 main aims: the impact of apolipoprotein L1 risk variants on acute kidney injury (AKI) and death in Black veterans with COVID-19; and pharmacogenetic analysis of remdesivir-induced liver chemistry abnormalities.
Disease mechanisms. Understanding the underlying pathways and mechanisms behind COVID-19 has been a difficult but important challenge overall in the scientific community. This working group investigated specific genetic markers and effects on COVID-19, including polygenic predisposition to venous thromboembolism associated with increased COVID-19 susceptibility; renal comorbidities and new AKI and unfavorable outcomes among COVID-19–positive sickle cell trait carriers; and mucin 5B, oligomeric mucus/gel-forming gene polymorphism, and protective effects in COVID-19 infection.
Genomics for risk prediction, polygenic risk scores, and mendelian randomization. Risk prediction for COVID-19 has been widely studied mostly aiming at comorbidities and preexisting conditions. The MVP cohort provided a unique opportunity to understand how genetic information can enhance our understanding of COVID-19 risk. This working group focused on: (1) ABO blood group typing and the protective effects of the O blood group on COVID-19 infection; (2) polygenic risk scores and COVID-19 outcomes; (3) human leukocyte antigen typing and COVID-19 outcomes; and (4) a transcriptome-wide association study of COVID-19–positive MVP participants.
Genome-Wide Association Study (GWAS) and Downstream Analysis. This working group performed GWAS of the main COVID-19 outcomes. Results from GWAS unveiled new genetic loci to suggest further investigation on these candidate genes. The results were used by other MVP COVID-19 working groups for their activities. The results also contributed to external collaborations, such as the COVID-19 Host Genetics Initiative.
COVID-19–Related PheWAS. This working group focused on understanding the potential clinical significance of genetic variants associated with susceptibility to, or outcomes of, COVID-19 infection. They worked to identify traits that share genetic variants associated with severe COVID-19 from the Host Genetics Initiative. The group also studied the phenotypic consequences of acquired mosaic chromosomal alterations with early data linking to COVID-19 susceptibility.
COVID-19 Research Partnerships
In 2016, the VA and DOE formed an interagency partnership known as Computational Health Analytics for Medical Precision to Improve Outcomes Now (CHAMPION) to demonstrate the power of combining the VA EHR system, MVP genetic data, and clinical research expertise with DOE high-performance computing infrastructure and artificial intelligence expertise. The VA EHR captures longitudinal care information on veterans with records that go back decades. Furthermore, the VA covers the costs of medications and
The DOE Oak Ridge National Laboratory (ORNL) in Tennessee securely maintains this rich database for the VA. The ORNL Summit supercomputer can complete trillions of calculations per second to provide critical and timely analyses, applying the most advanced and powerful artificial intelligence methods, which would not be possible in more conventional research settings. CHAMPION taught the VA and DOE how to bring their disparate research cultures together for innovative collaborative investigation. Moreover, this collaboration produced a cadre of VA and DOE scientists familiar with VA patient data and experienced in conducting joint research successfully and integrating omics data with clinical data for a better mechanistic understanding. Because of this preexisting collaboration between the VA and DOE, interagency teams were prepared at the start of the COVID-19 pandemic.10-15
Other recently completed studies have developed and validated short-term mortality indices in individuals with COVID-19 based on their preexisting conditions, assessed the generalizability of VA COVID-19 experiences to the US population, and evaluated the effectiveness of hydroxychloroquine with and without azithromycin in VA patients with COVID-19.12,15 A recent study demonstrated the benefit of prophylactic anticoagulation at initial hospitalization.14
The VA also provided the FDA with daily reports on aggregate VA COVID-19 cases and their distribution across the VA system, demographics of VA patients with COVID-19, and analyses of predictive models for positive test results and death. The VA regularly sent the FDA aggregated data showing patterns of medication use and retrospective analyses of the effectiveness of certain medications (including remdesivir and some antithrombotic agents). The FDA used these data along with other data to understand the scope of the pandemic and to predict drug shortages or needs for additional medical equipment, including ventilators.
Limitations
For the most part, MVP infrastructure and partnerships were efficiently leveraged to significantly advance our understanding of the biological basis of COVID-19 and to develop treatments and vaccines. However, there were a few limitations that may have slowed timely and optimal outcomes. An issue not limited to the MVP or VA was the continual evolution of the pandemic and its response. This included evolving definitions of disease, symptomatology, testing, vaccines, and public health recommendations. Keeping pace with the emerging knowledge from these domains was a struggle for the entire scientific community. A more discrete limitation was the number of participants in the MVP with positive COVID-19 test results and positive symptoms; however, this was mitigated by partnering with other groups like the COVID-19 Host Genetics Initiative to increase study participant numbers. Finally, there were logistical and regulatory challenges associated with coordination of national clinical trial recruitment across a VA system with > 100 discrete hospitals.
Conclusions
Having a centralized infrastructure for recruitment and enrollment, including a national research volunteer registry, information center, research staff, and coordinating centers, can allow for expedited enrollment in vaccine and treatment trials in the face of future public health emergencies. VA assets, including its rich EHR and MVP, the world’s largest genomic cohort, have contributed to improving our understanding and management of COVID-19.
The Million Veteran Program (MVP) was launched in 2011 by the US Department of Veterans Affairs (VA) to enroll at least 1 million veterans in a longitudinal cohort to better understand how genes, lifestyle, military experience, and environmental exposures interact to influence health and illness and ultimately enable precision health care. The MVP has established a national, centralized infrastructure for recruitment and enrollment, biospecimen and data collection and storage, data generation and curation, and secure data access. When the COVID-19 pandemic hit in 2020, the MVP was leveraged to support research utilizing the following key infrastructure components: (1) MVP recruitment and enrollment platform to provide support for COVID-19 vaccine and treatment trials and to collect COVID-19 data from MVP participants; (2) using MVP Phenomics for COVID-19 research data cleaning and curation, assisting with the development of a VA Severity Index for COVID-19, and forming 6 scientific working groups to coordinate COVID-19 research questions; and (3) the VA/MVP and US Department of Energy (DOE) partnership to assist in responding to COVID-19 research questions identified by the US Food and Drug Administration (FDA). This article describes these infrastructure components in more detail and highlights key findings from the MVP COVID-19 research efforts.
MVP Infrastructure
The Veterans Health Administration (VHA) Office of Research and Development (ORD) oversaw efforts to develop the VA Coronavirus Research Volunteer List (the COVID-19 registry). To support the registry, the MVP leveraged its infrastructure to facilitate a rapid response. The MVP is designed as a full-service and centralized recruitment and enrollment platform. This includes MVP office oversight; MVP coordinating centers that manage the centralized platform; an information center that handles inbound and outbound calls; an informatics system built for recruitment and enrollment monitoring and tracking; and a network of more than 70 participating MVP sites with dedicated staff to conduct recruitment and enrollment activities. The MVP used its informatics infrastructure to support secure data storage for the registry volunteer information. MVP coordinating center staff worked with the COVID-19 registry to invite > 125,000 MVP participants from approximately 20 MVP sites. Additionally, MVP information center staff made > 4000 calls to prospective registry volunteers. This work resulted in 1300 volunteers agreeing to be
New Data Collection
The MVP protocol was approved by the VA Central Institutional Review Board (IRB) in 2011. As part of initial enrollment in MVP, participants consented to recontact for additional self-report information along with access to their electronic health record (EHR). This allows for the linkage of EHR and survey response data, thus providing a comprehensive understanding of health history before and after a self-reported COVID-19 diagnosis. Between May 2020 and September 2021, the MVP COVID-19 survey was distributed to existing MVP participants via mail, telephone, and email with the ability to complete the survey by paper and pencil or through the MVP online system. Dissemination of the survey was approved by the VA Central IRB in 2020, with nearly 730,000 eligible MVP participants contacted. As of June 2022, 255,737 MVP participants (35% of the eligible cohort) had completed the survey; 86% completed a paper survey while 14% completed it online. Respondents were primarily older (≥ 65 years); 90% were male; close to 7% reported Hispanic ethnicity, and 11% reported Black race.
Findings from this survey provide insight into pandemic behaviors not consistently captured in EHRs, such as psychosocial aspects, including social and emotional support, loss of tangible and intangible resources, as well as COVID-19–related behaviors, such as social distancing and self-protective practices.1 MVP COVID-19 survey data combined with veteran EHRs, responses to other MVP surveys, and genetic data enable MVP researchers to better understand epidemiological, clinical, and psychosocial aspects of the disease. Future COVID-19 studies may use self-reported survey responses to enrich understanding about the effects of the disease on a veteran’s daily life, and possibly validate existing EHR COVID-19 diagnoses and hospitalization findings. This comprehensive data resource provides a unique opportunity to identify new targets for disease prevention, treatment, and management with an emphasis on individual variability in genes, environment, and lifestyle.
COVID-19 Research
In early 2020, the burden of COVID-19 on the US was unprecedented, and little was known about risk factors for severe COVID-19 and deaths. The MVP Phenomics team quickly responded with a large-scale phenome-wide association study (PheWAS) of >
To broaden disease progression data curation and fit the specific needs of the VA, we operationalized and validated the World Health Organization clinical severity scale and used VA EHR data to create the VA Severity Index for COVID-19 (VASIC).3 The VASIC category is now part of the MVP core data repository, where volumes of data from multiple activities are integrated through an automated process to create monthly research-ready data cubes. These activities include extensive data curation, mapping, phenotyping, and adjudication that are performed to curate oxygen supplementation status and other procedures related to treatment that are processed and understood in real time. The data cubes were provisioned to MVP COVID-19 researchers. In addition, the VASIC scale variable is now integrated within the larger VA system for all researchers to use as part of its wider COVID-19 initiative. The VA Centralized Interactive Phenomics Resource (CIPHER) phenomics library now hosts the details of VASIC, codes, metadata, and related COVID-19 data products for all VA communities. In partnership with CIPHER and other internal and external COVID-19 initiatives, the MVP continues to play an integral part for the VA and beyond in the development of a phenomics algorithm for long COVID, or post-acute COVID-19 syndrome (PACS).
Host Genetics in COVID-19
As the SARS-CoV-2 virus continued to spread globally, it became clear that the symptoms and severity of infection experienced by patients varied across a broad spectrum, from being asymptomatic carriers to experiencing severe symptoms in 1 or more organ systems in the body, resulting in death. This variability suggested that host genetics and other host factors may play a role in determining the severity of COVID-19 infection. The MVP dataset, with genetic and health information on > 600,000 MVP participants, provided an ideal dataset to explore host contributions to COVID-19.
In late spring 2020, the MVP executive committee issued a call to the MVP research community to propose study aims around the COVID-19 pandemic that could leverage the phenotypic and genetic data and resources. The MVP quickly formed 6 rapid-response scientific working groups. Their mission was to cultivate collaboration and inclusivity and to coordinate COVID-19 research questions. A steering committee composed of the MVP executive committee, staff from computational environments, working group cochairs, and an administrator, who was responsible for daily oversight of the working groups. In addition, the ORD COVID-19 steering committee reviewed and approved research activities to ensure scientific rigor, as well as alignment with overall ongoing research activities.
The MVP COVID-19 working groups included dozens of researchers who used MVP data to identify disease mechanisms; understand the impact of host genetics on susceptibility, morbidity, and mortality; and identify potential targets for treatments and therapies. The working groups were further supported by MVP analysts to work cross-functionally on genomics, phenomics, statistical genetics, and PheWAS. Each working group chair was responsible for prioritizing concepts and moving them forward in coordination with the MVP and ORD COVID-19 steering committees. An overview of the MVP COVID-19 working groups follows (Table).4-9
Druggable genome. This working group researched drug-repurposing opportunities to prevent severe COVID-19, defined as hospitalization with oxygen therapy (high flow), intubation, mechanical ventilation, vasopressors, dialysis, or death from COVID-19; and prevent complications in patients hospitalized by COVID-19.
Pharmacogenomics. This working group focused on 2 main aims: the impact of apolipoprotein L1 risk variants on acute kidney injury (AKI) and death in Black veterans with COVID-19; and pharmacogenetic analysis of remdesivir-induced liver chemistry abnormalities.
Disease mechanisms. Understanding the underlying pathways and mechanisms behind COVID-19 has been a difficult but important challenge overall in the scientific community. This working group investigated specific genetic markers and effects on COVID-19, including polygenic predisposition to venous thromboembolism associated with increased COVID-19 susceptibility; renal comorbidities and new AKI and unfavorable outcomes among COVID-19–positive sickle cell trait carriers; and mucin 5B, oligomeric mucus/gel-forming gene polymorphism, and protective effects in COVID-19 infection.
Genomics for risk prediction, polygenic risk scores, and mendelian randomization. Risk prediction for COVID-19 has been widely studied mostly aiming at comorbidities and preexisting conditions. The MVP cohort provided a unique opportunity to understand how genetic information can enhance our understanding of COVID-19 risk. This working group focused on: (1) ABO blood group typing and the protective effects of the O blood group on COVID-19 infection; (2) polygenic risk scores and COVID-19 outcomes; (3) human leukocyte antigen typing and COVID-19 outcomes; and (4) a transcriptome-wide association study of COVID-19–positive MVP participants.
Genome-Wide Association Study (GWAS) and Downstream Analysis. This working group performed GWAS of the main COVID-19 outcomes. Results from GWAS unveiled new genetic loci to suggest further investigation on these candidate genes. The results were used by other MVP COVID-19 working groups for their activities. The results also contributed to external collaborations, such as the COVID-19 Host Genetics Initiative.
COVID-19–Related PheWAS. This working group focused on understanding the potential clinical significance of genetic variants associated with susceptibility to, or outcomes of, COVID-19 infection. They worked to identify traits that share genetic variants associated with severe COVID-19 from the Host Genetics Initiative. The group also studied the phenotypic consequences of acquired mosaic chromosomal alterations with early data linking to COVID-19 susceptibility.
COVID-19 Research Partnerships
In 2016, the VA and DOE formed an interagency partnership known as Computational Health Analytics for Medical Precision to Improve Outcomes Now (CHAMPION) to demonstrate the power of combining the VA EHR system, MVP genetic data, and clinical research expertise with DOE high-performance computing infrastructure and artificial intelligence expertise. The VA EHR captures longitudinal care information on veterans with records that go back decades. Furthermore, the VA covers the costs of medications and
The DOE Oak Ridge National Laboratory (ORNL) in Tennessee securely maintains this rich database for the VA. The ORNL Summit supercomputer can complete trillions of calculations per second to provide critical and timely analyses, applying the most advanced and powerful artificial intelligence methods, which would not be possible in more conventional research settings. CHAMPION taught the VA and DOE how to bring their disparate research cultures together for innovative collaborative investigation. Moreover, this collaboration produced a cadre of VA and DOE scientists familiar with VA patient data and experienced in conducting joint research successfully and integrating omics data with clinical data for a better mechanistic understanding. Because of this preexisting collaboration between the VA and DOE, interagency teams were prepared at the start of the COVID-19 pandemic.10-15
Other recently completed studies have developed and validated short-term mortality indices in individuals with COVID-19 based on their preexisting conditions, assessed the generalizability of VA COVID-19 experiences to the US population, and evaluated the effectiveness of hydroxychloroquine with and without azithromycin in VA patients with COVID-19.12,15 A recent study demonstrated the benefit of prophylactic anticoagulation at initial hospitalization.14
The VA also provided the FDA with daily reports on aggregate VA COVID-19 cases and their distribution across the VA system, demographics of VA patients with COVID-19, and analyses of predictive models for positive test results and death. The VA regularly sent the FDA aggregated data showing patterns of medication use and retrospective analyses of the effectiveness of certain medications (including remdesivir and some antithrombotic agents). The FDA used these data along with other data to understand the scope of the pandemic and to predict drug shortages or needs for additional medical equipment, including ventilators.
Limitations
For the most part, MVP infrastructure and partnerships were efficiently leveraged to significantly advance our understanding of the biological basis of COVID-19 and to develop treatments and vaccines. However, there were a few limitations that may have slowed timely and optimal outcomes. An issue not limited to the MVP or VA was the continual evolution of the pandemic and its response. This included evolving definitions of disease, symptomatology, testing, vaccines, and public health recommendations. Keeping pace with the emerging knowledge from these domains was a struggle for the entire scientific community. A more discrete limitation was the number of participants in the MVP with positive COVID-19 test results and positive symptoms; however, this was mitigated by partnering with other groups like the COVID-19 Host Genetics Initiative to increase study participant numbers. Finally, there were logistical and regulatory challenges associated with coordination of national clinical trial recruitment across a VA system with > 100 discrete hospitals.
Conclusions
Having a centralized infrastructure for recruitment and enrollment, including a national research volunteer registry, information center, research staff, and coordinating centers, can allow for expedited enrollment in vaccine and treatment trials in the face of future public health emergencies. VA assets, including its rich EHR and MVP, the world’s largest genomic cohort, have contributed to improving our understanding and management of COVID-19.
1. Whitbourne SB, Nguyen XT, Song RJ, et al. Million Veteran Program’s response to COVID-19: survey development and preliminary findings. PLoS One. 2022;17(4):e0266381. doi:10.1371/journal.pone.0266381
2. Song RJ, Ho YL, Schubert P, et al. Phenome-wide association of 1809 phenotypes and COVID-19 disease progression in the Veterans Health Administration Million Veteran Program. PLoS One. 2021;16(5):e0251651. doi:10.1371/journal.pone.0251651
3. Galloway A, Park Y, Tanukonda V, et al. Impact of COVID-19 severity on long-term events in US veterans using the Veterans Affairs Severity Index for COVID-19 (VASIC). J Infect Dis. 2022;226(12):2113-2117. doi:10.1093/infdis/jiac182
4. Gaziano L, Giambartolomei C, Pereira AC, et al. Actionable druggable genome-wide Mendelian randomization identifies repurposing opportunities for COVID-19. Nat Med. 2021;27(4):668-676. doi:10.0138/s41591-021-01310-z
5. Hung AM, Sha SC, Bick AG, et al. APOL1 risk variants, acute kidney injury, and death in participants with African ancestry hospitalized with COVID-19 from the Million Veteran Program. JAMA Intern Med. 2022;182(4):386-395. doi:10.1001/jamainternmed.2021.8538
6. Verma A, Huffman JE, Gao L, et al. Association of kidney comorbidities and acute kidney failure with unfavorable outcomes after COVID-19 in individuals with the sickle cell trait. JAMA Intern Med. 2022;182(8):796-804. doi:10.1001/jamainternmed.2022.2141
7. Verma A, Tsao NL, Thomann LO, et al. A phenome-wide association study of genes associated with COVID-19 severity reveals shared genetics with complex diseases in the Million Veteran Program. PLoS Genet. 2022;18(4):e1010113. doi:10.1371/journal.pgen.1010113
8. Peloso GM, Tcheandjieu C, McGeary JE, et al. Genetic loci associated with COVID-19 positivity and hospitalization in White, Black, and Hispanic Veterans of the VA Million Veteran Program. Front Genetic. 2022;12:777076. doi:10.3389/fgene.2021.777076
9. Verma A, Minnier J, Wan ES, et al. A MUC5B gene polymorphism, rs35705950-T confers protective effects against COVID-19 hospitalization but not severe disease or mortality. Am J Respir Crit Care Med. 2022;182(8):796-804. doi:10.1164/rccm.202109-2166OC
10. Garvin MR, Alvarez C, Miller JI, et al. A mechanistic model and therapeutic interventions for COVID-19 involving a RAS-mediated bradykinin storm. Elife. 2020;e59177. doi:10.7554/eLife.59177
11. Rentsch CT, Kidwai-Khan F, Tate JP, et al. Patterns of COVID-19 testing and mortality by race and ethnicity among United States veterans: A nationwide cohort study. PLoS Med. 2020;17(9):e1003379. doi:10.1371/journal.pmed.1003379
12. King JT, Yoon JS, Rentsch CT, et al. Development and validation of a 30-day mortality index based on pre-existing medical administrative data from 13,323 COVID-19 patients: the Veterans Health Administration COVID-19 (VACO) Index. PLoS One. 2020;15(11):e0241825. doi:10.1371/journal.pone.0241825
13. Joubert W, Weighill D, Kainer D, et al. Attacking the opioid epidemic: determining the epistatic and pleiotropic genetic architectures for chronic pain and opioid addiction. SC18: International Conference for High Performance Computing, Networking, Storage and Analysis. Dallas, TX, USA, 2018:717-730. doi:10.1109/SC.2018.00060
14. Rentsch CT, Beckman JA, Tomlinson L, et al. Early initiation of prophylactic anticoagulation for prevention of COVID-19 mortality: a nationwide cohort study of hospitalized patients in the United States. BMJ. 2021;372:n311. doi:10.1136/bmj.n311
15. Gerlovin H, Posner DC, Ho YL, et al. Pharmacoepidemiology, machine learning, and COVID-19: an intent-to-treat analysis of hydroxychloroquine, with or without Azithromycin, and COVID-19 outcomes among hospitalized US Veterans. Am J Epidemiol. 2021;190(11): 2405-2419. doi:10.1093/aje/kwab183
1. Whitbourne SB, Nguyen XT, Song RJ, et al. Million Veteran Program’s response to COVID-19: survey development and preliminary findings. PLoS One. 2022;17(4):e0266381. doi:10.1371/journal.pone.0266381
2. Song RJ, Ho YL, Schubert P, et al. Phenome-wide association of 1809 phenotypes and COVID-19 disease progression in the Veterans Health Administration Million Veteran Program. PLoS One. 2021;16(5):e0251651. doi:10.1371/journal.pone.0251651
3. Galloway A, Park Y, Tanukonda V, et al. Impact of COVID-19 severity on long-term events in US veterans using the Veterans Affairs Severity Index for COVID-19 (VASIC). J Infect Dis. 2022;226(12):2113-2117. doi:10.1093/infdis/jiac182
4. Gaziano L, Giambartolomei C, Pereira AC, et al. Actionable druggable genome-wide Mendelian randomization identifies repurposing opportunities for COVID-19. Nat Med. 2021;27(4):668-676. doi:10.0138/s41591-021-01310-z
5. Hung AM, Sha SC, Bick AG, et al. APOL1 risk variants, acute kidney injury, and death in participants with African ancestry hospitalized with COVID-19 from the Million Veteran Program. JAMA Intern Med. 2022;182(4):386-395. doi:10.1001/jamainternmed.2021.8538
6. Verma A, Huffman JE, Gao L, et al. Association of kidney comorbidities and acute kidney failure with unfavorable outcomes after COVID-19 in individuals with the sickle cell trait. JAMA Intern Med. 2022;182(8):796-804. doi:10.1001/jamainternmed.2022.2141
7. Verma A, Tsao NL, Thomann LO, et al. A phenome-wide association study of genes associated with COVID-19 severity reveals shared genetics with complex diseases in the Million Veteran Program. PLoS Genet. 2022;18(4):e1010113. doi:10.1371/journal.pgen.1010113
8. Peloso GM, Tcheandjieu C, McGeary JE, et al. Genetic loci associated with COVID-19 positivity and hospitalization in White, Black, and Hispanic Veterans of the VA Million Veteran Program. Front Genetic. 2022;12:777076. doi:10.3389/fgene.2021.777076
9. Verma A, Minnier J, Wan ES, et al. A MUC5B gene polymorphism, rs35705950-T confers protective effects against COVID-19 hospitalization but not severe disease or mortality. Am J Respir Crit Care Med. 2022;182(8):796-804. doi:10.1164/rccm.202109-2166OC
10. Garvin MR, Alvarez C, Miller JI, et al. A mechanistic model and therapeutic interventions for COVID-19 involving a RAS-mediated bradykinin storm. Elife. 2020;e59177. doi:10.7554/eLife.59177
11. Rentsch CT, Kidwai-Khan F, Tate JP, et al. Patterns of COVID-19 testing and mortality by race and ethnicity among United States veterans: A nationwide cohort study. PLoS Med. 2020;17(9):e1003379. doi:10.1371/journal.pmed.1003379
12. King JT, Yoon JS, Rentsch CT, et al. Development and validation of a 30-day mortality index based on pre-existing medical administrative data from 13,323 COVID-19 patients: the Veterans Health Administration COVID-19 (VACO) Index. PLoS One. 2020;15(11):e0241825. doi:10.1371/journal.pone.0241825
13. Joubert W, Weighill D, Kainer D, et al. Attacking the opioid epidemic: determining the epistatic and pleiotropic genetic architectures for chronic pain and opioid addiction. SC18: International Conference for High Performance Computing, Networking, Storage and Analysis. Dallas, TX, USA, 2018:717-730. doi:10.1109/SC.2018.00060
14. Rentsch CT, Beckman JA, Tomlinson L, et al. Early initiation of prophylactic anticoagulation for prevention of COVID-19 mortality: a nationwide cohort study of hospitalized patients in the United States. BMJ. 2021;372:n311. doi:10.1136/bmj.n311
15. Gerlovin H, Posner DC, Ho YL, et al. Pharmacoepidemiology, machine learning, and COVID-19: an intent-to-treat analysis of hydroxychloroquine, with or without Azithromycin, and COVID-19 outcomes among hospitalized US Veterans. Am J Epidemiol. 2021;190(11): 2405-2419. doi:10.1093/aje/kwab183