User login
Method enables complex, large-scale genetic analysis

Image by Spencer Phillips
A new algorithm makes it possible to perform genetic analyses on as many as 500,000 individuals—and many traits—at the same time, according to an article in Nature Methods.
The authors noted that determining how genetic variations affect health and disease involves analyzing associations between many different variants and multiple traits and making the best use of data from large cohorts that include hundreds of thousands of individuals.
“It is very challenging to identify genetic variants that underlie phenotypes, or traits, and, usually, we do this by analyzing each phenotype and each variant one by one,” explained Oliver Stegle, PhD, of the European Molecular Biology Laboratory-European Bioinformatics Institute in Cambridge, UK.
“But the simple models we use to do this are too simplistic to uncover the complex dependencies between sets of genetic variants and disease phenotypes.”
On the other hand, complex models that reveal the combined action of many different variants have involved so much computation that it would take a year to run a single complex query. But Dr Stegle and his colleagues said their algorithm can change that.
“The breakthrough here is that we’ve made it possible to perform an integrative analysis involving many variants and phenotypes at the same speed as current approaches,” Dr Stegle said.
He and his colleagues tested their new method, called mtSet, on data from 2 studies and compared the results with existing tools used for genetic analysis. The researchers found that mtSet was substantially faster than existing methods and could explain a larger proportion of traits in terms of the genetics that drive them.
The team said mtSet will allow researchers to explore several variants of a gene at once, while comparing them with several related phenotypes. This makes it easier to pinpoint which genes—or locations on genes—are involved in a particular function.
“What’s important about this work is that it improves statistical power and provides the tools people need to analyze multiple traits in very large cohorts,” Dr Stegle said. “Our algorithm can be used to study up to half a million individuals. That hasn’t been possible until now.” ![]()

Image by Spencer Phillips
A new algorithm makes it possible to perform genetic analyses on as many as 500,000 individuals—and many traits—at the same time, according to an article in Nature Methods.
The authors noted that determining how genetic variations affect health and disease involves analyzing associations between many different variants and multiple traits and making the best use of data from large cohorts that include hundreds of thousands of individuals.
“It is very challenging to identify genetic variants that underlie phenotypes, or traits, and, usually, we do this by analyzing each phenotype and each variant one by one,” explained Oliver Stegle, PhD, of the European Molecular Biology Laboratory-European Bioinformatics Institute in Cambridge, UK.
“But the simple models we use to do this are too simplistic to uncover the complex dependencies between sets of genetic variants and disease phenotypes.”
On the other hand, complex models that reveal the combined action of many different variants have involved so much computation that it would take a year to run a single complex query. But Dr Stegle and his colleagues said their algorithm can change that.
“The breakthrough here is that we’ve made it possible to perform an integrative analysis involving many variants and phenotypes at the same speed as current approaches,” Dr Stegle said.
He and his colleagues tested their new method, called mtSet, on data from 2 studies and compared the results with existing tools used for genetic analysis. The researchers found that mtSet was substantially faster than existing methods and could explain a larger proportion of traits in terms of the genetics that drive them.
The team said mtSet will allow researchers to explore several variants of a gene at once, while comparing them with several related phenotypes. This makes it easier to pinpoint which genes—or locations on genes—are involved in a particular function.
“What’s important about this work is that it improves statistical power and provides the tools people need to analyze multiple traits in very large cohorts,” Dr Stegle said. “Our algorithm can be used to study up to half a million individuals. That hasn’t been possible until now.” ![]()

Image by Spencer Phillips
A new algorithm makes it possible to perform genetic analyses on as many as 500,000 individuals—and many traits—at the same time, according to an article in Nature Methods.
The authors noted that determining how genetic variations affect health and disease involves analyzing associations between many different variants and multiple traits and making the best use of data from large cohorts that include hundreds of thousands of individuals.
“It is very challenging to identify genetic variants that underlie phenotypes, or traits, and, usually, we do this by analyzing each phenotype and each variant one by one,” explained Oliver Stegle, PhD, of the European Molecular Biology Laboratory-European Bioinformatics Institute in Cambridge, UK.
“But the simple models we use to do this are too simplistic to uncover the complex dependencies between sets of genetic variants and disease phenotypes.”
On the other hand, complex models that reveal the combined action of many different variants have involved so much computation that it would take a year to run a single complex query. But Dr Stegle and his colleagues said their algorithm can change that.
“The breakthrough here is that we’ve made it possible to perform an integrative analysis involving many variants and phenotypes at the same speed as current approaches,” Dr Stegle said.
He and his colleagues tested their new method, called mtSet, on data from 2 studies and compared the results with existing tools used for genetic analysis. The researchers found that mtSet was substantially faster than existing methods and could explain a larger proportion of traits in terms of the genetics that drive them.
The team said mtSet will allow researchers to explore several variants of a gene at once, while comparing them with several related phenotypes. This makes it easier to pinpoint which genes—or locations on genes—are involved in a particular function.
“What’s important about this work is that it improves statistical power and provides the tools people need to analyze multiple traits in very large cohorts,” Dr Stegle said. “Our algorithm can be used to study up to half a million individuals. That hasn’t been possible until now.” ![]()
Team identifies new target for malaria vaccine

Photo courtesy of the CDC
Researchers say they have located a new, more precise target for a mosquito-based vaccine to block the transmission of malaria.
The team created a 3-D crystal structure of AnAPN1, a protein found in the gut of the Anopheles mosquito that is considered essential to malaria transmission.
In studying the entire protein, the researchers found that previous incarnations of a proposed vaccine included irrelevant regions of AnAPN1.
Rhoel R. Dinglasan, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland, and his colleagues described this work in Nature Structural & Molecular Biology.
AnAPN1 is found on the apical surface of the Anopheles gambiae midgut and is potentially a receptor for the Plasmodium parasite. As a vaccine antigen, AnAPN1 prompts antibody production, but only some of these antibodies block parasite transmission.
“This dilution of the overall antibody response to AnAPN1 is problematic,” Dr Dinglasan said. “To further improve vaccine immunogenicity at the preclinical stage, we need to immuno-focus the antibody response to only the critical, transmission-blocking regions of the protein.”
Using the Australian Synchrotron, the researchers were able to visualize the crystal structure of AnAPN1 for the first time. This allowed the team to pinpoint the binding site of AnAPN1 antibodies that can and cannot block parasite development.
“We now know much more about which parts of the AnAPN1 protein are involved in generating transmission-blocking antibodies and have a new hypothesis as to how they might work,” said Natalie Borg, PhD, of Monash University in Clayton, Victoria, Australia.
The researchers identified a particularly potent antibody, 4H5B7, and tested it in blood samples from children carrying the Plasmodium falciparum parasite. 4H5B7 exhibited “complete transmission-blocking activity” against these naturally circulating strains of P falciparum.
The team said their data indicate the mechanism underlying this activity is the recognition of a conformation-dependent epitope that is predominantly found on peptide 4, which is part of peptide 7 on domain 1 of AnAPN1.
These findings suggest that, previously, researchers were asking the immune system to target too many regions on AnAPN1, which diluted the response to the relevant regions of the protein. ![]()

Photo courtesy of the CDC
Researchers say they have located a new, more precise target for a mosquito-based vaccine to block the transmission of malaria.
The team created a 3-D crystal structure of AnAPN1, a protein found in the gut of the Anopheles mosquito that is considered essential to malaria transmission.
In studying the entire protein, the researchers found that previous incarnations of a proposed vaccine included irrelevant regions of AnAPN1.
Rhoel R. Dinglasan, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland, and his colleagues described this work in Nature Structural & Molecular Biology.
AnAPN1 is found on the apical surface of the Anopheles gambiae midgut and is potentially a receptor for the Plasmodium parasite. As a vaccine antigen, AnAPN1 prompts antibody production, but only some of these antibodies block parasite transmission.
“This dilution of the overall antibody response to AnAPN1 is problematic,” Dr Dinglasan said. “To further improve vaccine immunogenicity at the preclinical stage, we need to immuno-focus the antibody response to only the critical, transmission-blocking regions of the protein.”
Using the Australian Synchrotron, the researchers were able to visualize the crystal structure of AnAPN1 for the first time. This allowed the team to pinpoint the binding site of AnAPN1 antibodies that can and cannot block parasite development.
“We now know much more about which parts of the AnAPN1 protein are involved in generating transmission-blocking antibodies and have a new hypothesis as to how they might work,” said Natalie Borg, PhD, of Monash University in Clayton, Victoria, Australia.
The researchers identified a particularly potent antibody, 4H5B7, and tested it in blood samples from children carrying the Plasmodium falciparum parasite. 4H5B7 exhibited “complete transmission-blocking activity” against these naturally circulating strains of P falciparum.
The team said their data indicate the mechanism underlying this activity is the recognition of a conformation-dependent epitope that is predominantly found on peptide 4, which is part of peptide 7 on domain 1 of AnAPN1.
These findings suggest that, previously, researchers were asking the immune system to target too many regions on AnAPN1, which diluted the response to the relevant regions of the protein. ![]()

Photo courtesy of the CDC
Researchers say they have located a new, more precise target for a mosquito-based vaccine to block the transmission of malaria.
The team created a 3-D crystal structure of AnAPN1, a protein found in the gut of the Anopheles mosquito that is considered essential to malaria transmission.
In studying the entire protein, the researchers found that previous incarnations of a proposed vaccine included irrelevant regions of AnAPN1.
Rhoel R. Dinglasan, PhD, of the Johns Hopkins Bloomberg School of Public Health in Baltimore, Maryland, and his colleagues described this work in Nature Structural & Molecular Biology.
AnAPN1 is found on the apical surface of the Anopheles gambiae midgut and is potentially a receptor for the Plasmodium parasite. As a vaccine antigen, AnAPN1 prompts antibody production, but only some of these antibodies block parasite transmission.
“This dilution of the overall antibody response to AnAPN1 is problematic,” Dr Dinglasan said. “To further improve vaccine immunogenicity at the preclinical stage, we need to immuno-focus the antibody response to only the critical, transmission-blocking regions of the protein.”
Using the Australian Synchrotron, the researchers were able to visualize the crystal structure of AnAPN1 for the first time. This allowed the team to pinpoint the binding site of AnAPN1 antibodies that can and cannot block parasite development.
“We now know much more about which parts of the AnAPN1 protein are involved in generating transmission-blocking antibodies and have a new hypothesis as to how they might work,” said Natalie Borg, PhD, of Monash University in Clayton, Victoria, Australia.
The researchers identified a particularly potent antibody, 4H5B7, and tested it in blood samples from children carrying the Plasmodium falciparum parasite. 4H5B7 exhibited “complete transmission-blocking activity” against these naturally circulating strains of P falciparum.
The team said their data indicate the mechanism underlying this activity is the recognition of a conformation-dependent epitope that is predominantly found on peptide 4, which is part of peptide 7 on domain 1 of AnAPN1.
These findings suggest that, previously, researchers were asking the immune system to target too many regions on AnAPN1, which diluted the response to the relevant regions of the protein. ![]()
New trial design accounts for patient behavior

Photo by Esther Dyson
New research suggests patient behavior can have a serious impact on the effectiveness of a treatment, and procedures used in double-blind randomized controlled trials (DBRCTs) may not allow researchers to assess the effects of behavior on treatment.
To solve this problem, a group of investigators has proposed a new trial design, called a two-by-two trial, that can account for behavior-treatment interactions.
They described this design in PLOS ONE.
Study rationale
The study authors pointed out that patient behaviors can directly relate to a trial. For example, a patient who believes in the drug might religiously stick to his or her treatment regimen, while someone more skeptical might skip a few doses.
Alternatively, patient behaviors may relate to how the person acts in general, such as preferences in diet, exercise, and social engagement. And in the design of today’s standard trials, these behaviors are not accounted for, said author Erik Snowberg, PhD, of the California Institute of Technology in Pasadena.
He noted that DBRCTs typically assign patients to an experimental group that receives the new treatment and a control group that does not. As the trial is double-blinded, neither the subjects nor their doctors know who falls into which group.
However, patients do know their probability of getting the treatment, and that 50% likelihood of getting the new treatment might not be enough to encourage a patient to change behaviors that could influence the efficacy of the drug, Dr Snowberg said.
For example, if a subject wants to lose weight and knows he has a high probability—a 70% chance—of being in the experimental group for a new weight loss drug, he may be more likely to take the drug as directed and to make other healthy lifestyle choices that can contribute to weight loss. As a result, he might lose more weight, boosting the apparent effectiveness of the drug.
However, if the subject knows he only has a 30% chance of being in the experimental group, he might be less motivated to both take the drug as directed and to make those other changes. As a result, he might lose less weight—even if he is in the treatment group—and the same drug would seem less effective.
“Most medical research just wants to know if a drug will work or not,” Dr Snowberg said. “We wanted to go a step further, designing new trials that would take into account the way people behave. As social scientists, we naturally turned to the mathematical tools of formal social science to do this.”
New trial design
Dr Snowberg and his colleagues found that, with the two-by-two trial, they can tease out the effects of behavior and the interaction of behavior and treatment, as well as the effects of treatment alone. The new trial design, which still randomizes treatment, also randomizes the probability of treatment, which can change a patient’s behavior.
In a two-by-two trial, instead of patients first being assigned to either the experimental or control groups, they are randomly assigned to either a “high probability of treatment” group or a “low probability of treatment” group.
The patients in the high probability group are then randomized to either the treatment or the control group, giving them a 70% chance of receiving the treatment. Patients in the low probability group are also randomized to treatment or control, and their likelihood of receiving the treatment is 30%. The patients are then informed of their probability of treatment.
By randomizing both the treatment and the probability of treatment, researchers can quantify the effects of treatment, the effects of behavior, and the effects of the interaction between treatment and behavior, Dr Snowberg said. And determining each is essential for understanding the overall efficacy of treatment.
“It’s a very small change to the design of the trial, but it’s important,” he said. “The effect of a treatment has these two constituent parts: pure treatment effect and the treatment-behavior interaction. Standard blind trials just randomize the likelihood of treatment, so you can’t see this interaction. Although you can’t just tell someone to randomize their behavior, we found a way that you can randomize the probability that a patient will get something that will change their behavior.”
Testing the design
Because it is difficult to implement new trial design changes in active trials, Dr Snowberg and his colleagues wanted to first test their idea with a meta-analysis of data from previous clinical trials. They devised a new mathematical formula that can be used to analyze DBRCT data.
The formula teases out the health outcomes resulting from treatment alone as well as outcomes resulting from an interaction between treatment and behavior.
The investigators used the formula to analyze 6 DBRCTs evaluating the antidepressants imipramine (a tricyclic antidepressant also known as Tofranil) and paroxetine (a selective serotonin reuptake inhibitor sold as Paxil).
First, the researchers wanted to see if there was evidence that patients behave differently when they have a high probability of treatment and when they have a low probability of treatment.
The trials recorded how many patients dropped out of the study, so this was the behavior Dr Snowberg and his colleagues analyzed. They found that, in trials where patients happened to have a relatively high probability of treatment—near 70%—the dropout rate was significantly lower than in trials where patients had a lower probability of treatment—around 50%.
Although the team did not have any specific behaviors to analyze, other than dropping out of the study, they also wanted to determine if behavior in general could have added to the effect of the treatments.
Using their statistical techniques, the investigators found that imipramine seemed to have a pure treatment effect but no effect from the interaction between treatment and behavior. That is, the drug seemed to work fine, regardless of any behavioral differences that may have been present.
The researchers also found that paroxetine seemed to have no effect from the treatment alone or behavior alone. However, an interaction between the treatment and behavior did effectively decrease depression.
Because this study was conducted in the past, the investigators could not determine which specific behavior was responsible for the interaction, but with the mathematical formula, they were able to tell that this behavior was necessary for the drug to be effective.
In their paper, Dr Snowberg and his colleagues speculate how a situation like this might come about.
“Maybe there is a drug, for instance, that makes people feel better in social situations, and if you’re in the high probability group, then maybe you’d be more willing to go out to parties to see if the drug helps you talk to people,” Dr Snowberg explained.
“Your behavior drives you to go to the party, and once you’re at the party, the drug helps you feel comfortable talking to people. That would be an example of an interaction effect. You couldn’t get that if people just took this drug alone at home.”
Although this specific example is just speculation, Dr Snowberg said the researchers’ actual results reveal that some behavior or set of behaviors interact with paroxetine to effectively treat depression. And, without this behavior, the drug appears to be ineffective.
“Normally, what you get when you run a standard blind trial is some sort of mishmash of the treatment effect and the treatment-behavior interaction effect,” Dr Snowberg said. “But knowing the full interaction effect is important.”
“Our work indicates that clinical trials underestimate the efficacy of a drug where behavior matters. It may be the case that 50% probability isn’t high enough for people to change any of their behaviors, especially if it’s a really uncertain new treatment. Then, it’s just going to look like the drug doesn’t work, and that isn’t the case.”
Because the meta-analysis supported the team’s hypothesis—that the interaction between treatment and behavior can have an effect on health outcomes—the next step is incorporating these new ideas into an active clinical trial.
Dr Snowberg said the best fit would be a drug trial for a condition, such as a mental health disorder or an addiction, that is known to be associated with behavior. At the very least, he hopes these results will lead the medical research community to a conversation about ways to improve the DBRCT. ![]()

Photo by Esther Dyson
New research suggests patient behavior can have a serious impact on the effectiveness of a treatment, and procedures used in double-blind randomized controlled trials (DBRCTs) may not allow researchers to assess the effects of behavior on treatment.
To solve this problem, a group of investigators has proposed a new trial design, called a two-by-two trial, that can account for behavior-treatment interactions.
They described this design in PLOS ONE.
Study rationale
The study authors pointed out that patient behaviors can directly relate to a trial. For example, a patient who believes in the drug might religiously stick to his or her treatment regimen, while someone more skeptical might skip a few doses.
Alternatively, patient behaviors may relate to how the person acts in general, such as preferences in diet, exercise, and social engagement. And in the design of today’s standard trials, these behaviors are not accounted for, said author Erik Snowberg, PhD, of the California Institute of Technology in Pasadena.
He noted that DBRCTs typically assign patients to an experimental group that receives the new treatment and a control group that does not. As the trial is double-blinded, neither the subjects nor their doctors know who falls into which group.
However, patients do know their probability of getting the treatment, and that 50% likelihood of getting the new treatment might not be enough to encourage a patient to change behaviors that could influence the efficacy of the drug, Dr Snowberg said.
For example, if a subject wants to lose weight and knows he has a high probability—a 70% chance—of being in the experimental group for a new weight loss drug, he may be more likely to take the drug as directed and to make other healthy lifestyle choices that can contribute to weight loss. As a result, he might lose more weight, boosting the apparent effectiveness of the drug.
However, if the subject knows he only has a 30% chance of being in the experimental group, he might be less motivated to both take the drug as directed and to make those other changes. As a result, he might lose less weight—even if he is in the treatment group—and the same drug would seem less effective.
“Most medical research just wants to know if a drug will work or not,” Dr Snowberg said. “We wanted to go a step further, designing new trials that would take into account the way people behave. As social scientists, we naturally turned to the mathematical tools of formal social science to do this.”
New trial design
Dr Snowberg and his colleagues found that, with the two-by-two trial, they can tease out the effects of behavior and the interaction of behavior and treatment, as well as the effects of treatment alone. The new trial design, which still randomizes treatment, also randomizes the probability of treatment, which can change a patient’s behavior.
In a two-by-two trial, instead of patients first being assigned to either the experimental or control groups, they are randomly assigned to either a “high probability of treatment” group or a “low probability of treatment” group.
The patients in the high probability group are then randomized to either the treatment or the control group, giving them a 70% chance of receiving the treatment. Patients in the low probability group are also randomized to treatment or control, and their likelihood of receiving the treatment is 30%. The patients are then informed of their probability of treatment.
By randomizing both the treatment and the probability of treatment, researchers can quantify the effects of treatment, the effects of behavior, and the effects of the interaction between treatment and behavior, Dr Snowberg said. And determining each is essential for understanding the overall efficacy of treatment.
“It’s a very small change to the design of the trial, but it’s important,” he said. “The effect of a treatment has these two constituent parts: pure treatment effect and the treatment-behavior interaction. Standard blind trials just randomize the likelihood of treatment, so you can’t see this interaction. Although you can’t just tell someone to randomize their behavior, we found a way that you can randomize the probability that a patient will get something that will change their behavior.”
Testing the design
Because it is difficult to implement new trial design changes in active trials, Dr Snowberg and his colleagues wanted to first test their idea with a meta-analysis of data from previous clinical trials. They devised a new mathematical formula that can be used to analyze DBRCT data.
The formula teases out the health outcomes resulting from treatment alone as well as outcomes resulting from an interaction between treatment and behavior.
The investigators used the formula to analyze 6 DBRCTs evaluating the antidepressants imipramine (a tricyclic antidepressant also known as Tofranil) and paroxetine (a selective serotonin reuptake inhibitor sold as Paxil).
First, the researchers wanted to see if there was evidence that patients behave differently when they have a high probability of treatment and when they have a low probability of treatment.
The trials recorded how many patients dropped out of the study, so this was the behavior Dr Snowberg and his colleagues analyzed. They found that, in trials where patients happened to have a relatively high probability of treatment—near 70%—the dropout rate was significantly lower than in trials where patients had a lower probability of treatment—around 50%.
Although the team did not have any specific behaviors to analyze, other than dropping out of the study, they also wanted to determine if behavior in general could have added to the effect of the treatments.
Using their statistical techniques, the investigators found that imipramine seemed to have a pure treatment effect but no effect from the interaction between treatment and behavior. That is, the drug seemed to work fine, regardless of any behavioral differences that may have been present.
The researchers also found that paroxetine seemed to have no effect from the treatment alone or behavior alone. However, an interaction between the treatment and behavior did effectively decrease depression.
Because this study was conducted in the past, the investigators could not determine which specific behavior was responsible for the interaction, but with the mathematical formula, they were able to tell that this behavior was necessary for the drug to be effective.
In their paper, Dr Snowberg and his colleagues speculate how a situation like this might come about.
“Maybe there is a drug, for instance, that makes people feel better in social situations, and if you’re in the high probability group, then maybe you’d be more willing to go out to parties to see if the drug helps you talk to people,” Dr Snowberg explained.
“Your behavior drives you to go to the party, and once you’re at the party, the drug helps you feel comfortable talking to people. That would be an example of an interaction effect. You couldn’t get that if people just took this drug alone at home.”
Although this specific example is just speculation, Dr Snowberg said the researchers’ actual results reveal that some behavior or set of behaviors interact with paroxetine to effectively treat depression. And, without this behavior, the drug appears to be ineffective.
“Normally, what you get when you run a standard blind trial is some sort of mishmash of the treatment effect and the treatment-behavior interaction effect,” Dr Snowberg said. “But knowing the full interaction effect is important.”
“Our work indicates that clinical trials underestimate the efficacy of a drug where behavior matters. It may be the case that 50% probability isn’t high enough for people to change any of their behaviors, especially if it’s a really uncertain new treatment. Then, it’s just going to look like the drug doesn’t work, and that isn’t the case.”
Because the meta-analysis supported the team’s hypothesis—that the interaction between treatment and behavior can have an effect on health outcomes—the next step is incorporating these new ideas into an active clinical trial.
Dr Snowberg said the best fit would be a drug trial for a condition, such as a mental health disorder or an addiction, that is known to be associated with behavior. At the very least, he hopes these results will lead the medical research community to a conversation about ways to improve the DBRCT. ![]()

Photo by Esther Dyson
New research suggests patient behavior can have a serious impact on the effectiveness of a treatment, and procedures used in double-blind randomized controlled trials (DBRCTs) may not allow researchers to assess the effects of behavior on treatment.
To solve this problem, a group of investigators has proposed a new trial design, called a two-by-two trial, that can account for behavior-treatment interactions.
They described this design in PLOS ONE.
Study rationale
The study authors pointed out that patient behaviors can directly relate to a trial. For example, a patient who believes in the drug might religiously stick to his or her treatment regimen, while someone more skeptical might skip a few doses.
Alternatively, patient behaviors may relate to how the person acts in general, such as preferences in diet, exercise, and social engagement. And in the design of today’s standard trials, these behaviors are not accounted for, said author Erik Snowberg, PhD, of the California Institute of Technology in Pasadena.
He noted that DBRCTs typically assign patients to an experimental group that receives the new treatment and a control group that does not. As the trial is double-blinded, neither the subjects nor their doctors know who falls into which group.
However, patients do know their probability of getting the treatment, and that 50% likelihood of getting the new treatment might not be enough to encourage a patient to change behaviors that could influence the efficacy of the drug, Dr Snowberg said.
For example, if a subject wants to lose weight and knows he has a high probability—a 70% chance—of being in the experimental group for a new weight loss drug, he may be more likely to take the drug as directed and to make other healthy lifestyle choices that can contribute to weight loss. As a result, he might lose more weight, boosting the apparent effectiveness of the drug.
However, if the subject knows he only has a 30% chance of being in the experimental group, he might be less motivated to both take the drug as directed and to make those other changes. As a result, he might lose less weight—even if he is in the treatment group—and the same drug would seem less effective.
“Most medical research just wants to know if a drug will work or not,” Dr Snowberg said. “We wanted to go a step further, designing new trials that would take into account the way people behave. As social scientists, we naturally turned to the mathematical tools of formal social science to do this.”
New trial design
Dr Snowberg and his colleagues found that, with the two-by-two trial, they can tease out the effects of behavior and the interaction of behavior and treatment, as well as the effects of treatment alone. The new trial design, which still randomizes treatment, also randomizes the probability of treatment, which can change a patient’s behavior.
In a two-by-two trial, instead of patients first being assigned to either the experimental or control groups, they are randomly assigned to either a “high probability of treatment” group or a “low probability of treatment” group.
The patients in the high probability group are then randomized to either the treatment or the control group, giving them a 70% chance of receiving the treatment. Patients in the low probability group are also randomized to treatment or control, and their likelihood of receiving the treatment is 30%. The patients are then informed of their probability of treatment.
By randomizing both the treatment and the probability of treatment, researchers can quantify the effects of treatment, the effects of behavior, and the effects of the interaction between treatment and behavior, Dr Snowberg said. And determining each is essential for understanding the overall efficacy of treatment.
“It’s a very small change to the design of the trial, but it’s important,” he said. “The effect of a treatment has these two constituent parts: pure treatment effect and the treatment-behavior interaction. Standard blind trials just randomize the likelihood of treatment, so you can’t see this interaction. Although you can’t just tell someone to randomize their behavior, we found a way that you can randomize the probability that a patient will get something that will change their behavior.”
Testing the design
Because it is difficult to implement new trial design changes in active trials, Dr Snowberg and his colleagues wanted to first test their idea with a meta-analysis of data from previous clinical trials. They devised a new mathematical formula that can be used to analyze DBRCT data.
The formula teases out the health outcomes resulting from treatment alone as well as outcomes resulting from an interaction between treatment and behavior.
The investigators used the formula to analyze 6 DBRCTs evaluating the antidepressants imipramine (a tricyclic antidepressant also known as Tofranil) and paroxetine (a selective serotonin reuptake inhibitor sold as Paxil).
First, the researchers wanted to see if there was evidence that patients behave differently when they have a high probability of treatment and when they have a low probability of treatment.
The trials recorded how many patients dropped out of the study, so this was the behavior Dr Snowberg and his colleagues analyzed. They found that, in trials where patients happened to have a relatively high probability of treatment—near 70%—the dropout rate was significantly lower than in trials where patients had a lower probability of treatment—around 50%.
Although the team did not have any specific behaviors to analyze, other than dropping out of the study, they also wanted to determine if behavior in general could have added to the effect of the treatments.
Using their statistical techniques, the investigators found that imipramine seemed to have a pure treatment effect but no effect from the interaction between treatment and behavior. That is, the drug seemed to work fine, regardless of any behavioral differences that may have been present.
The researchers also found that paroxetine seemed to have no effect from the treatment alone or behavior alone. However, an interaction between the treatment and behavior did effectively decrease depression.
Because this study was conducted in the past, the investigators could not determine which specific behavior was responsible for the interaction, but with the mathematical formula, they were able to tell that this behavior was necessary for the drug to be effective.
In their paper, Dr Snowberg and his colleagues speculate how a situation like this might come about.
“Maybe there is a drug, for instance, that makes people feel better in social situations, and if you’re in the high probability group, then maybe you’d be more willing to go out to parties to see if the drug helps you talk to people,” Dr Snowberg explained.
“Your behavior drives you to go to the party, and once you’re at the party, the drug helps you feel comfortable talking to people. That would be an example of an interaction effect. You couldn’t get that if people just took this drug alone at home.”
Although this specific example is just speculation, Dr Snowberg said the researchers’ actual results reveal that some behavior or set of behaviors interact with paroxetine to effectively treat depression. And, without this behavior, the drug appears to be ineffective.
“Normally, what you get when you run a standard blind trial is some sort of mishmash of the treatment effect and the treatment-behavior interaction effect,” Dr Snowberg said. “But knowing the full interaction effect is important.”
“Our work indicates that clinical trials underestimate the efficacy of a drug where behavior matters. It may be the case that 50% probability isn’t high enough for people to change any of their behaviors, especially if it’s a really uncertain new treatment. Then, it’s just going to look like the drug doesn’t work, and that isn’t the case.”
Because the meta-analysis supported the team’s hypothesis—that the interaction between treatment and behavior can have an effect on health outcomes—the next step is incorporating these new ideas into an active clinical trial.
Dr Snowberg said the best fit would be a drug trial for a condition, such as a mental health disorder or an addiction, that is known to be associated with behavior. At the very least, he hopes these results will lead the medical research community to a conversation about ways to improve the DBRCT. ![]()
Modern housing may reduce malaria risk
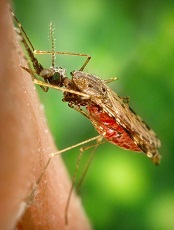
Photo by James Gathany
Living in a “modern” house may reduce a person’s risk of contracting malaria, according to research published in Malaria Journal.
As insecticide and drug resistance are on the rise, researchers wanted to determine how making changes to housing might aid the fight against malaria.
So they reviewed 90 studies conducted in Africa, Asia, and South America, comparing the incidence of malaria among people who live in “traditional” and “modern” houses.
The traditional houses consisted of mud, stone, bamboo, or wood walls; thatched, mud, or wood roofs; and earth or wood floors. The modern houses had closed eaves, ceilings, screened doors, and windows.
The researchers found that residents of modern homes were 47% less likely to be infected with malaria than those living in traditional houses. And residents of modern homes were 45% to 65% less likely to have clinical malaria (fever with infection).
“Housing improvements were traditionally an important pillar of public health, but they remain underexploited in malaria control,” said study author Lucy Tusting, of the London School of Hygiene & Tropical Medicine in the UK.
“Good housing can block mosquitoes from entering homes and prevent them from transmitting malaria to the people who live there. Our study suggests housing could be an important tool in tackling malaria. This is a welcome finding at a time when we are facing increasing resistance to our most effective insecticides and drugs.”
“We now need to pinpoint which housing features can reduce mosquito entry in different settings, to incorporate these into local housing designs, and to assess the impact on malaria in large-scale field trials.”
The researchers noted that the effectiveness of improving housing will vary depending on the location. While many mosquitoes enter homes to bite humans at night, outdoor malaria transmission is more common in some places. So interventions centered on the home will have less of an impact in these areas.
The researchers also conceded that the studies eligible for inclusion in this review were of low quality. However, they said the consistency of the findings indicate that housing is an important risk factor for malaria. ![]()

Photo by James Gathany
Living in a “modern” house may reduce a person’s risk of contracting malaria, according to research published in Malaria Journal.
As insecticide and drug resistance are on the rise, researchers wanted to determine how making changes to housing might aid the fight against malaria.
So they reviewed 90 studies conducted in Africa, Asia, and South America, comparing the incidence of malaria among people who live in “traditional” and “modern” houses.
The traditional houses consisted of mud, stone, bamboo, or wood walls; thatched, mud, or wood roofs; and earth or wood floors. The modern houses had closed eaves, ceilings, screened doors, and windows.
The researchers found that residents of modern homes were 47% less likely to be infected with malaria than those living in traditional houses. And residents of modern homes were 45% to 65% less likely to have clinical malaria (fever with infection).
“Housing improvements were traditionally an important pillar of public health, but they remain underexploited in malaria control,” said study author Lucy Tusting, of the London School of Hygiene & Tropical Medicine in the UK.
“Good housing can block mosquitoes from entering homes and prevent them from transmitting malaria to the people who live there. Our study suggests housing could be an important tool in tackling malaria. This is a welcome finding at a time when we are facing increasing resistance to our most effective insecticides and drugs.”
“We now need to pinpoint which housing features can reduce mosquito entry in different settings, to incorporate these into local housing designs, and to assess the impact on malaria in large-scale field trials.”
The researchers noted that the effectiveness of improving housing will vary depending on the location. While many mosquitoes enter homes to bite humans at night, outdoor malaria transmission is more common in some places. So interventions centered on the home will have less of an impact in these areas.
The researchers also conceded that the studies eligible for inclusion in this review were of low quality. However, they said the consistency of the findings indicate that housing is an important risk factor for malaria. ![]()

Photo by James Gathany
Living in a “modern” house may reduce a person’s risk of contracting malaria, according to research published in Malaria Journal.
As insecticide and drug resistance are on the rise, researchers wanted to determine how making changes to housing might aid the fight against malaria.
So they reviewed 90 studies conducted in Africa, Asia, and South America, comparing the incidence of malaria among people who live in “traditional” and “modern” houses.
The traditional houses consisted of mud, stone, bamboo, or wood walls; thatched, mud, or wood roofs; and earth or wood floors. The modern houses had closed eaves, ceilings, screened doors, and windows.
The researchers found that residents of modern homes were 47% less likely to be infected with malaria than those living in traditional houses. And residents of modern homes were 45% to 65% less likely to have clinical malaria (fever with infection).
“Housing improvements were traditionally an important pillar of public health, but they remain underexploited in malaria control,” said study author Lucy Tusting, of the London School of Hygiene & Tropical Medicine in the UK.
“Good housing can block mosquitoes from entering homes and prevent them from transmitting malaria to the people who live there. Our study suggests housing could be an important tool in tackling malaria. This is a welcome finding at a time when we are facing increasing resistance to our most effective insecticides and drugs.”
“We now need to pinpoint which housing features can reduce mosquito entry in different settings, to incorporate these into local housing designs, and to assess the impact on malaria in large-scale field trials.”
The researchers noted that the effectiveness of improving housing will vary depending on the location. While many mosquitoes enter homes to bite humans at night, outdoor malaria transmission is more common in some places. So interventions centered on the home will have less of an impact in these areas.
The researchers also conceded that the studies eligible for inclusion in this review were of low quality. However, they said the consistency of the findings indicate that housing is an important risk factor for malaria. ![]()
Viral protein protects EBV-infected B cells
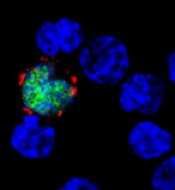
expresses the ligand (red)
that activates NKG2D, while
uninfected cells (blue) do not
Benjamin Chaigne-Delalande
A study published in PLOS Pathogens sheds new light on why the immune system cannot eliminate Epstein-Barr virus (EBV) or the risk of cancer associated with the virus.
Researchers investigated the immune system’s response against EBV, focusing on the role of LMP2A.
This viral protein is present in latently infected B cells and in many EBV-associated cancers, which have somehow escaped detection and elimination by the immune system.
Andreas Moosmann, PhD, of the Helmholtz-Zentrum in Munich, Germany, and his colleagues studied an engineered EBV virus that cannot make LMP2A and compared this mutant virus with the normal one.
The researchers infected human B cells with normal and LMP2A-deficient EBV. Because EBV transforms these cells, the team was able to examine lymphoblastic cell lines that contained either virus.
They found that LMP2A counteracts the recognition of EBV-infected B cells by EBV-specific, CD8+ killer T cells. In contrast, EBV-transformed cells without LMP2A are more efficiently identified, and the T cells’ ability to recognize and kill the EBV-infected B cells is enhanced.
The researchers examined the mechanism underlying the LMP2A-mediated evasion and found several ways in which it interferes with the recognition of EBV-infected cells.
First, LMP2A reduced the levels of several EBV proteins whose fragments are recognized by CD8+ T cells on the surface of the cell targeted for killing.
Second, LMP2A disturbs the expression of cellular molecules on infected B cells that interact with NKG2D, a host molecule on the surface of CD8+ T cells that aids their activation, thereby weakening the immune response against EBV-infected cells.
The researchers said these results suggest a functional immunomodulatory effect for the EBV protein LMP2A and show that LMP2A mediates the partial escape of infected B cells from recognition by CD8+ T cells.
The team also said similar immune evasion mechanisms may operate in different types of LMP2A-expressing cancers caused by EBV. ![]()

expresses the ligand (red)
that activates NKG2D, while
uninfected cells (blue) do not
Benjamin Chaigne-Delalande
A study published in PLOS Pathogens sheds new light on why the immune system cannot eliminate Epstein-Barr virus (EBV) or the risk of cancer associated with the virus.
Researchers investigated the immune system’s response against EBV, focusing on the role of LMP2A.
This viral protein is present in latently infected B cells and in many EBV-associated cancers, which have somehow escaped detection and elimination by the immune system.
Andreas Moosmann, PhD, of the Helmholtz-Zentrum in Munich, Germany, and his colleagues studied an engineered EBV virus that cannot make LMP2A and compared this mutant virus with the normal one.
The researchers infected human B cells with normal and LMP2A-deficient EBV. Because EBV transforms these cells, the team was able to examine lymphoblastic cell lines that contained either virus.
They found that LMP2A counteracts the recognition of EBV-infected B cells by EBV-specific, CD8+ killer T cells. In contrast, EBV-transformed cells without LMP2A are more efficiently identified, and the T cells’ ability to recognize and kill the EBV-infected B cells is enhanced.
The researchers examined the mechanism underlying the LMP2A-mediated evasion and found several ways in which it interferes with the recognition of EBV-infected cells.
First, LMP2A reduced the levels of several EBV proteins whose fragments are recognized by CD8+ T cells on the surface of the cell targeted for killing.
Second, LMP2A disturbs the expression of cellular molecules on infected B cells that interact with NKG2D, a host molecule on the surface of CD8+ T cells that aids their activation, thereby weakening the immune response against EBV-infected cells.
The researchers said these results suggest a functional immunomodulatory effect for the EBV protein LMP2A and show that LMP2A mediates the partial escape of infected B cells from recognition by CD8+ T cells.
The team also said similar immune evasion mechanisms may operate in different types of LMP2A-expressing cancers caused by EBV. ![]()

expresses the ligand (red)
that activates NKG2D, while
uninfected cells (blue) do not
Benjamin Chaigne-Delalande
A study published in PLOS Pathogens sheds new light on why the immune system cannot eliminate Epstein-Barr virus (EBV) or the risk of cancer associated with the virus.
Researchers investigated the immune system’s response against EBV, focusing on the role of LMP2A.
This viral protein is present in latently infected B cells and in many EBV-associated cancers, which have somehow escaped detection and elimination by the immune system.
Andreas Moosmann, PhD, of the Helmholtz-Zentrum in Munich, Germany, and his colleagues studied an engineered EBV virus that cannot make LMP2A and compared this mutant virus with the normal one.
The researchers infected human B cells with normal and LMP2A-deficient EBV. Because EBV transforms these cells, the team was able to examine lymphoblastic cell lines that contained either virus.
They found that LMP2A counteracts the recognition of EBV-infected B cells by EBV-specific, CD8+ killer T cells. In contrast, EBV-transformed cells without LMP2A are more efficiently identified, and the T cells’ ability to recognize and kill the EBV-infected B cells is enhanced.
The researchers examined the mechanism underlying the LMP2A-mediated evasion and found several ways in which it interferes with the recognition of EBV-infected cells.
First, LMP2A reduced the levels of several EBV proteins whose fragments are recognized by CD8+ T cells on the surface of the cell targeted for killing.
Second, LMP2A disturbs the expression of cellular molecules on infected B cells that interact with NKG2D, a host molecule on the surface of CD8+ T cells that aids their activation, thereby weakening the immune response against EBV-infected cells.
The researchers said these results suggest a functional immunomodulatory effect for the EBV protein LMP2A and show that LMP2A mediates the partial escape of infected B cells from recognition by CD8+ T cells.
The team also said similar immune evasion mechanisms may operate in different types of LMP2A-expressing cancers caused by EBV. ![]()
Harnessing immune defense to treat Candida infection
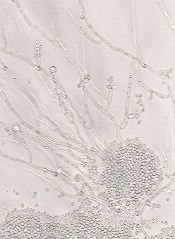
An upset in the body’s natural balance of gut bacteria that may lead to life-threatening bloodstream infections can be reversed by enhancing an
immune response, according to research published in Nature Medicine.
Researchers found that a transcription factor known as HIF-1α works with LL-37, a naturally occurring antibiotic, to kill the infection-causing fungi Candida albicans.
And this response can be enhanced with a drug called L-mimosine.
The researchers noted that Candida albicans can be lethal if it invades the bloodstream from the gut. And stem cell transplant recipients and immunosuppressed cancer patients have a high risk for this type of infection.
“For a cancer patient with a Candida bloodstream infection, the fatality rate is about 30%, [and] Candida is the number 1 fungal pathogen,” said study author Andrew Koh, MD, of the University of Texas Southwestern Medical Center in Dallas.
With that in mind, he and his colleagues set out to determine how the body’s natural immune defense system might be enhanced to fight a Candida infection. By studying how mice infected with Candida responded in different scenarios, the team found their answer.
“The commensal bacteria stimulate gut tissue to make a transcription factor and a natural antibiotic, which then kills the Candida fungus,” Dr Koh explained.
“When we gave the mice a pharmacologic agent called L-mimosine that stimulates the transcription factor, the agent knocked down Candida 100-fold, which translated into a 50% reduction in mortality from invasive Candida infection.”
Specifically, the researchers found that enhancing the transcription factor HIF-1α with L-mimosine led to increased production of the natural antibiotic peptide LL-37, which, in turn, killed the fungi. L-mimosine is a natural product derived from seeds of the koa haole tree that is known to boost HIF-1α activity.
The study also suggested that certain gut bacteria—Clostridial Firmicutes and Bacteroidetes—may be important in producing short-chain fatty acids that help fight infection.
The researchers said more work is needed to pinpoint the optimal method of inducing the body’s gut defense system, whether through use of an agent like L-mimosine or by administering short-chain fatty acids such as vinegar.
“Can we modulate the gut system to maintain balance so that it never gets to the point of pathogens invading the bloodstream?” Dr Koh asked. “Boosting [gastrointestinal] mucosal immune effectors to reduce fungal burden may be the key to tipping the balance back toward normal and preventing invasive fungal disease.” ![]()

An upset in the body’s natural balance of gut bacteria that may lead to life-threatening bloodstream infections can be reversed by enhancing an
immune response, according to research published in Nature Medicine.
Researchers found that a transcription factor known as HIF-1α works with LL-37, a naturally occurring antibiotic, to kill the infection-causing fungi Candida albicans.
And this response can be enhanced with a drug called L-mimosine.
The researchers noted that Candida albicans can be lethal if it invades the bloodstream from the gut. And stem cell transplant recipients and immunosuppressed cancer patients have a high risk for this type of infection.
“For a cancer patient with a Candida bloodstream infection, the fatality rate is about 30%, [and] Candida is the number 1 fungal pathogen,” said study author Andrew Koh, MD, of the University of Texas Southwestern Medical Center in Dallas.
With that in mind, he and his colleagues set out to determine how the body’s natural immune defense system might be enhanced to fight a Candida infection. By studying how mice infected with Candida responded in different scenarios, the team found their answer.
“The commensal bacteria stimulate gut tissue to make a transcription factor and a natural antibiotic, which then kills the Candida fungus,” Dr Koh explained.
“When we gave the mice a pharmacologic agent called L-mimosine that stimulates the transcription factor, the agent knocked down Candida 100-fold, which translated into a 50% reduction in mortality from invasive Candida infection.”
Specifically, the researchers found that enhancing the transcription factor HIF-1α with L-mimosine led to increased production of the natural antibiotic peptide LL-37, which, in turn, killed the fungi. L-mimosine is a natural product derived from seeds of the koa haole tree that is known to boost HIF-1α activity.
The study also suggested that certain gut bacteria—Clostridial Firmicutes and Bacteroidetes—may be important in producing short-chain fatty acids that help fight infection.
The researchers said more work is needed to pinpoint the optimal method of inducing the body’s gut defense system, whether through use of an agent like L-mimosine or by administering short-chain fatty acids such as vinegar.
“Can we modulate the gut system to maintain balance so that it never gets to the point of pathogens invading the bloodstream?” Dr Koh asked. “Boosting [gastrointestinal] mucosal immune effectors to reduce fungal burden may be the key to tipping the balance back toward normal and preventing invasive fungal disease.” ![]()

An upset in the body’s natural balance of gut bacteria that may lead to life-threatening bloodstream infections can be reversed by enhancing an
immune response, according to research published in Nature Medicine.
Researchers found that a transcription factor known as HIF-1α works with LL-37, a naturally occurring antibiotic, to kill the infection-causing fungi Candida albicans.
And this response can be enhanced with a drug called L-mimosine.
The researchers noted that Candida albicans can be lethal if it invades the bloodstream from the gut. And stem cell transplant recipients and immunosuppressed cancer patients have a high risk for this type of infection.
“For a cancer patient with a Candida bloodstream infection, the fatality rate is about 30%, [and] Candida is the number 1 fungal pathogen,” said study author Andrew Koh, MD, of the University of Texas Southwestern Medical Center in Dallas.
With that in mind, he and his colleagues set out to determine how the body’s natural immune defense system might be enhanced to fight a Candida infection. By studying how mice infected with Candida responded in different scenarios, the team found their answer.
“The commensal bacteria stimulate gut tissue to make a transcription factor and a natural antibiotic, which then kills the Candida fungus,” Dr Koh explained.
“When we gave the mice a pharmacologic agent called L-mimosine that stimulates the transcription factor, the agent knocked down Candida 100-fold, which translated into a 50% reduction in mortality from invasive Candida infection.”
Specifically, the researchers found that enhancing the transcription factor HIF-1α with L-mimosine led to increased production of the natural antibiotic peptide LL-37, which, in turn, killed the fungi. L-mimosine is a natural product derived from seeds of the koa haole tree that is known to boost HIF-1α activity.
The study also suggested that certain gut bacteria—Clostridial Firmicutes and Bacteroidetes—may be important in producing short-chain fatty acids that help fight infection.
The researchers said more work is needed to pinpoint the optimal method of inducing the body’s gut defense system, whether through use of an agent like L-mimosine or by administering short-chain fatty acids such as vinegar.
“Can we modulate the gut system to maintain balance so that it never gets to the point of pathogens invading the bloodstream?” Dr Koh asked. “Boosting [gastrointestinal] mucosal immune effectors to reduce fungal burden may be the key to tipping the balance back toward normal and preventing invasive fungal disease.” ![]()
Host cell type may impact malaria treatment
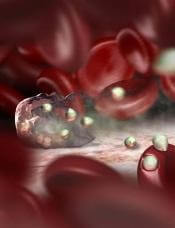
Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism. ![]()

Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism. ![]()

Image by Peter H. Seeberger
A study published in PLOS Pathogens suggests the different metabolic states of reticulocytes and erythrocytes provide different growth conditions for the malaria parasites Plasmodium vivax and Plasmodium falciparum.
As P vivax grows exclusively in reticulocytes, and P falciparum grows primarily in erythrocytes, the research suggests drugs that work against one species might fail to be effective against the other.
Andrew Waters, PhD, of the University of Glasgow in the UK, and his colleagues set out to determine whether the 2 classes of host red blood cells offer different resources for parasite survival and whether these resources could influence antimalarial drug efficacy.
To do that, the team analyzed the metabolites present in reticulocytes and erythrocytes. They found that reticulocytes contain elevated levels of many metabolites that could potentially be scavenged by the invading and growing malaria parasite.
And there was a marked overlap in metabolic pathways observed in the reticulocyte and those predicted in the parasite. The researchers thought these common pathways might be uniquely dispensable to Plasmodium during its growth in reticulocytes but essential—and therefore a good drug target—for growth in erythrocytes.
To test this hypothesis, the team disrupted some of the overlapping pathways in P berghei, a species that causes malaria in mice and, similar to P vivax, has a strong preference for growth in reticulocytes.
They found the mutant P berghei strains could grow in mouse reticulocytes (utilizing the host’s metabolic products).
The researchers also compared the sensitivity of P berghei and P falciparum to a drug known to target one of the overlapping pathways, the pyrimidine biosynthesis inhibitor 5-fluoroorotate (5FOA).
They found that P berghei was considerably less sensitive to 5FOA than P falciparum. The IC50 value of 5FOA in vitro was almost 90-fold higher in P berghei than in P falciparum.
This was presumably because P berghei was able to scavenge the metabolites from their reticulocyte host environment, but no such external sources were available in the erythrocyte host cells invaded by P falciparum.
The researchers said their results indicate that reticulocytes provide a highly enriched host cell environment for Plasmodium parasites, and the availability of the reticulocyte metabolome might reduce or block the efficacy of antimalarial drugs that target parasite metabolism.
Blood test screens for many viruses simultaneously
Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.

Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.

Photo by Максим Кукушкин
Scientists have reported that a new test can screen patients for current and past infection with more than 1000 viral strains, using a single drop of blood and for a cost of about $25 per blood sample.
The investigators used the test, known as VirScan, to screen more than 500 people from around the world and found that, on average, participants had been exposed to about 10 viral species over their lifetimes.
“Using this method, we can take a tiny drop of blood and determine what viruses a person has been infected with over the course of many years,” said Stephen Elledge, PhD, of Harvard Medical School in Boston, Massachusetts.
“What makes this so unique is the scale. Right now, a physician needs to guess what virus might be at play and individually test for it. With VirScan, we can look for virtually all viruses, even rare ones, with a single test.”
Dr Elledge and his colleagues described their work with VirScan in Science.
To develop VirScan, the group synthesized more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacteriophages.
Each bacteriophage manufactured a peptide and displayed it on the bacteriophage surface. As a group, the bacteriophages displayed all of the protein sequences found in the more than 1000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophages are allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes within the displayed peptides.
The scientists then retrieve the antibodies and wash away everything except the few bacteriophages that cling to them. By sequencing the DNA of those bacteriophages, the team can identify which viral protein pieces are bound to antibodies in the blood sample.
That reveals which viruses a person’s immune system has previously encountered, either through infection or vaccination.
Dr Elledge estimated that it would take about 2 to 3 days to process 100 samples with VirScan, assuming sequencing is working optimally. He said he is optimistic the speed of the assay will increase with further development.
Putting VirScan to the test
To test VirScan, Dr Elledge and his colleagues used it to analyze blood samples from patients known to be infected with particular viruses, including HIV and hepatitis C.
“It turns out that it works really well,” Dr Elledge said. “We were in the sensitivity range of 95% to 100% for those, and the specificity was good. We didn’t falsely identify people who were negative. That gave us confidence that we could detect other viruses, and when we did see them, we would know they were real.”
So the investigators used VirScan to analyze the antibodies in 569 people from 4 countries (Peru, South Africa, Thailand, and the US), examining about 100 million potential antibody/epitope interactions.
The team found that, on average, each person had antibodies to 10 different species of viruses. As expected, antibodies against certain viruses were common among adults but not in children, suggesting that children had not yet been exposed to those viruses.
Individuals residing in South Africa, Peru, and Thailand tended to have antibodies against more viruses than people in the US. And people infected with HIV had antibodies against many more viruses than people without HIV.
Dr Elledge and his colleagues were surprised to find that antibody responses against specific viruses were similar between individuals, with different people’s antibodies recognizing identical amino acids in the viral peptides.
“In this paper alone, we identified more antibody/peptide interactions to viral proteins than had been identified in the previous history of all viral exploration,” Dr Elledge said.
The reproducibility of those interactions allowed the scientists to refine their analysis and improve the sensitivity of VirScan, Dr Elledge said, adding that the method will continue to improve as his team analyzes more samples.
The investigators also noted that their work is not limited to antiviral antibodies. They are now using it to look for antibodies that attack the body’s tissue in autoimmune diseases associated with cancers. A similar approach could be used to screen for antibodies against other types of pathogens as well.
Biochemist Irwin Rose dies at 88

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.

Photo courtesy of UCI
Biochemist and Nobel laureate Irwin “Ernie” Rose, PhD, has passed away at the age of 88.
Dr Rose and colleagues from Israel won the Nobel Prize in Chemistry in 2004 for their discovery of ubiquitin-mediated protein degradation.
This research has wide-ranging implications for medicine and led to the development of anticancer drugs such as bortezomib, which is approved in the US to treat multiple myeloma and mantle cell lymphoma.
According to his friends and colleagues, Dr Rose was humble, generous, and endlessly curious.
“Ernie was not interested in personal fame and was oblivious to the politics of science,” said Ann Skalka, PhD, of Fox Chase Cancer Center in Philadelphia, Pennsylvania.
“His total satisfaction came from solving intricate biochemical puzzles. Although Ernie was an intellectual leader on the project that ultimately won him the Nobel, he took no personal credit. He was rather surprised at being recognized, but all of us at Fox Chase knew that the Nobel Committee had gotten it right.”
Dr Rose was born in Brooklyn, New York, on July 16, 1926. His scientific ambitions began to take shape after he moved to Spokane, Washington, at 13. While in high school, he spent summers working at a local hospital. And this inspired him to pursue a career that involved “solving medical problems.”
Dr Rose attended Washington State College for his undergraduate work and went on to earn a doctoral degree at the University of Chicago, after a brief stint in the Navy. He spent the better part of his career as a research scientist at the Fox Chase Cancer Center.
There, during the late 1970s and early 1980s, Dr Rose helped reveal how ubiquitin molecules facilitate the breakdown of old and damaged proteins. The discovery of this process fostered a new understanding of the molecular activity present in cancers and other diseases.
For the work, Dr Rose shared the 2004 Nobel Prize in Chemistry with Avram Hershko, MD, PhD, and Aaron Ciechanover, MD, PhD, of the Israel Institute of Technology.
“Ernie had a genius for asking the right questions,” said Jonathan Chernoff, MD, PhD, of Fox Chase Cancer Center.
“In the mid-1950s, when many scientists were interested in how proteins are synthesized, Ernie became fascinated with the opposite issue—how are proteins degraded? With the collaboration of his Israeli colleagues, he cracked that problem with the discovery of the ubiquitin conjugating system.”
After retiring to Laguna Woods, California, in 1997, Dr Rose accepted a special research position with the University of California Irvine (UCI).
There, he studied the mechanisms of fumarase, an enzyme involved in the citric acid cycle, the cellular pathway by which higher organisms convert food into energy. And he quickly became a beloved colleague and mentor to students and faculty.
“[B]oth prior to and after winning the Nobel Prize, he would help any student or young postdoctoral researcher who was having a hard time with an experiment,” said Ralph Bradshaw, PhD, a former professor at UCI.
“It was a lot of fun working with him,” said James Nowick, PhD, of UCI. “He worked with his own hands, not relying on others, with old instrumentation, and was able to do literally superb science.”
“He was the quintessential scientist—perseverant, soft-spoken, and interested in science for science’s sake,” Dr Chernoff said. “We will miss him very much.”
Dr Rose died in his sleep on June 2 in Deerfield, Massachusetts. He is survived by his wife, Zelda; their sons, Howard, Frederic, and Robert; and 5 grandchildren. Dr Rose’s daughter, Sarah, died in 2005.
Nutrition source could improve platinum-based nanodrugs

A parenteral nutrition source can reduce the toxicity and increase the bioavailability of platinum-based anticancer nanodrugs, according to preclinical research published in Scientific Reports.
Many of the side effects of platinum-based drugs occur when they settle in healthy tissue.
To deliver these drugs in a more targeted way, researchers have created nanoscale delivery systems engineered to make the drugs accumulate at tumor sites.
However, tests of these nanodrugs show that between 1% and 10% of the drugs are delivered to the tumor site, with most of the remainder being diverted to the liver and spleen.
“The body’s immune system, especially the liver and spleen, has been one of the biggest stumbling blocks in developing nanoscale chemotherapy drug delivery systems,” said Chien Ho, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“When the drugs collect in those organs, they become less available to treat the cancer and can also cause toxicity.”
But Dr Ho and his colleagues have found evidence to suggest that Intralipid, a fat emulsion used as a parenteral nutrition source, can help prevent that.
While developing cellular nanotags to help detect organ rejection, Dr Ho noticed that Intralipid reduced the amount of nanoparticles that were being cleared by the liver and spleen by about 50%. As a result, the nanoparticles remained in the bloodstream for longer periods of time.
So he and his colleagues decided to see if Intralipid had the same effect on platinum-based anticancer nanodrugs.
In the newly published study, the researchers administered a single, clinical dose of Intralipid to Sprague Dawley rats. One hour later, they administered a dose of a platinum-based chemotherapy drug that had been incorporated into a nanoparticle to both Intralipid-treated rats and controls.
Twenty-four hours after the drug was administered, rats pretreated with Intralipid had experienced reduced accumulation of the platinum-based drug compared to controls.
Drug accumulation decreased by 20.4% in the liver, 42.5% in the spleen, and 31.2% in the kidney. Consequently, in these organs, the toxic side effects of the nanodrug were significantly decreased compared to controls.
Furthermore, Intralipid pretreatment allowed more of the drug to remain available and active in the body for longer periods of time.
After 5 hours, the drug’s bioavailability increased by 18.7% in Intralipid-treated mice compared to controls. After 24 hours, bioavailability was 9.4% higher in Intralipid-treated mice than in controls.
The researchers believe this increased bioavailability will allow more of the drug to reach the tumor site and could perhaps allow clinicians to reduce the dosage needed to treat a patient. The team is now investigating the possibility of bringing this research to a clinical trial.

A parenteral nutrition source can reduce the toxicity and increase the bioavailability of platinum-based anticancer nanodrugs, according to preclinical research published in Scientific Reports.
Many of the side effects of platinum-based drugs occur when they settle in healthy tissue.
To deliver these drugs in a more targeted way, researchers have created nanoscale delivery systems engineered to make the drugs accumulate at tumor sites.
However, tests of these nanodrugs show that between 1% and 10% of the drugs are delivered to the tumor site, with most of the remainder being diverted to the liver and spleen.
“The body’s immune system, especially the liver and spleen, has been one of the biggest stumbling blocks in developing nanoscale chemotherapy drug delivery systems,” said Chien Ho, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“When the drugs collect in those organs, they become less available to treat the cancer and can also cause toxicity.”
But Dr Ho and his colleagues have found evidence to suggest that Intralipid, a fat emulsion used as a parenteral nutrition source, can help prevent that.
While developing cellular nanotags to help detect organ rejection, Dr Ho noticed that Intralipid reduced the amount of nanoparticles that were being cleared by the liver and spleen by about 50%. As a result, the nanoparticles remained in the bloodstream for longer periods of time.
So he and his colleagues decided to see if Intralipid had the same effect on platinum-based anticancer nanodrugs.
In the newly published study, the researchers administered a single, clinical dose of Intralipid to Sprague Dawley rats. One hour later, they administered a dose of a platinum-based chemotherapy drug that had been incorporated into a nanoparticle to both Intralipid-treated rats and controls.
Twenty-four hours after the drug was administered, rats pretreated with Intralipid had experienced reduced accumulation of the platinum-based drug compared to controls.
Drug accumulation decreased by 20.4% in the liver, 42.5% in the spleen, and 31.2% in the kidney. Consequently, in these organs, the toxic side effects of the nanodrug were significantly decreased compared to controls.
Furthermore, Intralipid pretreatment allowed more of the drug to remain available and active in the body for longer periods of time.
After 5 hours, the drug’s bioavailability increased by 18.7% in Intralipid-treated mice compared to controls. After 24 hours, bioavailability was 9.4% higher in Intralipid-treated mice than in controls.
The researchers believe this increased bioavailability will allow more of the drug to reach the tumor site and could perhaps allow clinicians to reduce the dosage needed to treat a patient. The team is now investigating the possibility of bringing this research to a clinical trial.

A parenteral nutrition source can reduce the toxicity and increase the bioavailability of platinum-based anticancer nanodrugs, according to preclinical research published in Scientific Reports.
Many of the side effects of platinum-based drugs occur when they settle in healthy tissue.
To deliver these drugs in a more targeted way, researchers have created nanoscale delivery systems engineered to make the drugs accumulate at tumor sites.
However, tests of these nanodrugs show that between 1% and 10% of the drugs are delivered to the tumor site, with most of the remainder being diverted to the liver and spleen.
“The body’s immune system, especially the liver and spleen, has been one of the biggest stumbling blocks in developing nanoscale chemotherapy drug delivery systems,” said Chien Ho, PhD, of Carnegie Mellon University in Pittsburg, Pennsylvania.
“When the drugs collect in those organs, they become less available to treat the cancer and can also cause toxicity.”
But Dr Ho and his colleagues have found evidence to suggest that Intralipid, a fat emulsion used as a parenteral nutrition source, can help prevent that.
While developing cellular nanotags to help detect organ rejection, Dr Ho noticed that Intralipid reduced the amount of nanoparticles that were being cleared by the liver and spleen by about 50%. As a result, the nanoparticles remained in the bloodstream for longer periods of time.
So he and his colleagues decided to see if Intralipid had the same effect on platinum-based anticancer nanodrugs.
In the newly published study, the researchers administered a single, clinical dose of Intralipid to Sprague Dawley rats. One hour later, they administered a dose of a platinum-based chemotherapy drug that had been incorporated into a nanoparticle to both Intralipid-treated rats and controls.
Twenty-four hours after the drug was administered, rats pretreated with Intralipid had experienced reduced accumulation of the platinum-based drug compared to controls.
Drug accumulation decreased by 20.4% in the liver, 42.5% in the spleen, and 31.2% in the kidney. Consequently, in these organs, the toxic side effects of the nanodrug were significantly decreased compared to controls.
Furthermore, Intralipid pretreatment allowed more of the drug to remain available and active in the body for longer periods of time.
After 5 hours, the drug’s bioavailability increased by 18.7% in Intralipid-treated mice compared to controls. After 24 hours, bioavailability was 9.4% higher in Intralipid-treated mice than in controls.
The researchers believe this increased bioavailability will allow more of the drug to reach the tumor site and could perhaps allow clinicians to reduce the dosage needed to treat a patient. The team is now investigating the possibility of bringing this research to a clinical trial.