User login
Mutations could be therapeutic target for FL
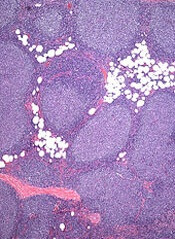
Mutations in the RRAGC gene appear to be an “excellent candidate for therapeutic targeting” in follicular lymphoma (FL), according to investigators.
They analyzed mutations found in tumors with multiple relapses of FL without transformation to diffuse large B-cell lymphoma.
And they found that one commonly mutated gene encodes the protein RagC, which is essential for activating the amino-acid sensing mTORC1 pathway.
Although mutations in genes in the mTORC1 pathway have been associated with various cancers, this is the first time a genetic mutation in any of the 4 Rag proteins has been identified in malignancy.
“One of the mutations that we have identified allows follicular lymphoma tumors to turn on growth signals regardless of whether nutrients are available, thereby evading normal restrictions on its growth,” said study author Jessica Okosun, MB BChir, PhD, of Barts Cancer Institute at Queen Mary University of London in the UK.
“Remarkably, the mutations we have discovered have not been seen in other cancer types. However, drugs that directly target this nutrient-sensing mechanism are currently used to treat other types of cancer and may benefit patients with follicular lymphoma.”
Dr Okosun and her colleagues reported these findings in a letter to Nature Genetics.
In experiments with cell lines, the investigators found that expression of the mutated RagC proteins activate mTORC1 signaling in the absence of amino acids and increase binding to an important part of the mTORC1 complex, consistent with the established role of RagC in the mTORC1 pathway.
Because this research was performed exclusively in cell lines, the investigators have not yet deciphered the mutations’ mechanistic effect in patients. However, study author Rachel Wolfson, of the Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology in Cambridge, Massachusetts, said there are clues to the mutations’ significance.
“mTORC1 is linked to cell growth, so it is not surprising that activation of the pathway could lead to some growth advantage for cancer cells,” Wolfson said. “But it leads to an interesting question: When is it a proliferative advantage versus a disadvantage to no longer be able to accurately sense amino acid levels? That is something we would need to investigate further, likely in vivo.”
The investigators would also like to know how the drug rapamycin affects FL with RagC mutations. Rapamycin binds to mTORC1 and inhibits its activity. If the drug interferes with mTORC1 dysregulation caused by RagC mutations, perhaps the drug could be used in FL treatment.
“If so, maybe these RagC mutations could be used as biomarkers to predict sensitivity to rapamycin treatment in follicular lymphoma patients,” Wolfson said. “That would be very exciting, and it’s something that should be investigated further.” ![]()

Mutations in the RRAGC gene appear to be an “excellent candidate for therapeutic targeting” in follicular lymphoma (FL), according to investigators.
They analyzed mutations found in tumors with multiple relapses of FL without transformation to diffuse large B-cell lymphoma.
And they found that one commonly mutated gene encodes the protein RagC, which is essential for activating the amino-acid sensing mTORC1 pathway.
Although mutations in genes in the mTORC1 pathway have been associated with various cancers, this is the first time a genetic mutation in any of the 4 Rag proteins has been identified in malignancy.
“One of the mutations that we have identified allows follicular lymphoma tumors to turn on growth signals regardless of whether nutrients are available, thereby evading normal restrictions on its growth,” said study author Jessica Okosun, MB BChir, PhD, of Barts Cancer Institute at Queen Mary University of London in the UK.
“Remarkably, the mutations we have discovered have not been seen in other cancer types. However, drugs that directly target this nutrient-sensing mechanism are currently used to treat other types of cancer and may benefit patients with follicular lymphoma.”
Dr Okosun and her colleagues reported these findings in a letter to Nature Genetics.
In experiments with cell lines, the investigators found that expression of the mutated RagC proteins activate mTORC1 signaling in the absence of amino acids and increase binding to an important part of the mTORC1 complex, consistent with the established role of RagC in the mTORC1 pathway.
Because this research was performed exclusively in cell lines, the investigators have not yet deciphered the mutations’ mechanistic effect in patients. However, study author Rachel Wolfson, of the Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology in Cambridge, Massachusetts, said there are clues to the mutations’ significance.
“mTORC1 is linked to cell growth, so it is not surprising that activation of the pathway could lead to some growth advantage for cancer cells,” Wolfson said. “But it leads to an interesting question: When is it a proliferative advantage versus a disadvantage to no longer be able to accurately sense amino acid levels? That is something we would need to investigate further, likely in vivo.”
The investigators would also like to know how the drug rapamycin affects FL with RagC mutations. Rapamycin binds to mTORC1 and inhibits its activity. If the drug interferes with mTORC1 dysregulation caused by RagC mutations, perhaps the drug could be used in FL treatment.
“If so, maybe these RagC mutations could be used as biomarkers to predict sensitivity to rapamycin treatment in follicular lymphoma patients,” Wolfson said. “That would be very exciting, and it’s something that should be investigated further.” ![]()

Mutations in the RRAGC gene appear to be an “excellent candidate for therapeutic targeting” in follicular lymphoma (FL), according to investigators.
They analyzed mutations found in tumors with multiple relapses of FL without transformation to diffuse large B-cell lymphoma.
And they found that one commonly mutated gene encodes the protein RagC, which is essential for activating the amino-acid sensing mTORC1 pathway.
Although mutations in genes in the mTORC1 pathway have been associated with various cancers, this is the first time a genetic mutation in any of the 4 Rag proteins has been identified in malignancy.
“One of the mutations that we have identified allows follicular lymphoma tumors to turn on growth signals regardless of whether nutrients are available, thereby evading normal restrictions on its growth,” said study author Jessica Okosun, MB BChir, PhD, of Barts Cancer Institute at Queen Mary University of London in the UK.
“Remarkably, the mutations we have discovered have not been seen in other cancer types. However, drugs that directly target this nutrient-sensing mechanism are currently used to treat other types of cancer and may benefit patients with follicular lymphoma.”
Dr Okosun and her colleagues reported these findings in a letter to Nature Genetics.
In experiments with cell lines, the investigators found that expression of the mutated RagC proteins activate mTORC1 signaling in the absence of amino acids and increase binding to an important part of the mTORC1 complex, consistent with the established role of RagC in the mTORC1 pathway.
Because this research was performed exclusively in cell lines, the investigators have not yet deciphered the mutations’ mechanistic effect in patients. However, study author Rachel Wolfson, of the Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology in Cambridge, Massachusetts, said there are clues to the mutations’ significance.
“mTORC1 is linked to cell growth, so it is not surprising that activation of the pathway could lead to some growth advantage for cancer cells,” Wolfson said. “But it leads to an interesting question: When is it a proliferative advantage versus a disadvantage to no longer be able to accurately sense amino acid levels? That is something we would need to investigate further, likely in vivo.”
The investigators would also like to know how the drug rapamycin affects FL with RagC mutations. Rapamycin binds to mTORC1 and inhibits its activity. If the drug interferes with mTORC1 dysregulation caused by RagC mutations, perhaps the drug could be used in FL treatment.
“If so, maybe these RagC mutations could be used as biomarkers to predict sensitivity to rapamycin treatment in follicular lymphoma patients,” Wolfson said. “That would be very exciting, and it’s something that should be investigated further.” ![]()
Studies reveal lack of transparency and poor reporting of research

Photo by Bill Branson
Two new studies suggest biomedical research may be hindered by poor reporting and a lack of transparency.
In one study, researchers analyzed more than 400 biomedical science articles and found the papers rarely provided full protocol information, complete data, and the necessary level of transparency to verify or replicate the work.
In the other study, researchers analyzed more than 500 preclinical experiments and found that most didn’t contain sufficient information on the animals used.
Both studies were published in PLOS Biology.
For the first study, Shareen Iqbal, PhD, of Emory University in Atlanta, Georgia, and her colleagues analyzed papers published between 2000 and 2014.
The team set out to determine the extent to which researchers report key information necessary for properly evaluating and replicating published research, including availability of protocols, data, and the frequency of published novel or replication studies.
Out of 441 articles drawn from across the biomedical literature, only 1 paper provided a full protocol, and none of the papers made all the data available. The majority of studies didn’t state funding or conflicts of interest, and replication studies were very rare.
Dr Iqbal and her colleagues said they hope their study will further sensitize scientists, funders, journals, and other science-related stakeholders about the need to improve these indicators.
For the second study, Ulrich Dirnagl, MD, of Charité Universitätsmedizin in Berlin, Germany, and his colleagues examined 100 papers describing preclinical research on stroke and cancer. These papers contained accounts of 316 experiments on infarct volume and 206 experiments on tumor shrinkage.
The vast majority of the reports didn’t contain sufficient information on how many animals were used in the experiments. What’s more, in many papers, animals “vanished” over the course of the study.
Using a computer model, the researchers simulated the effects of such animal loss on the validity of the experiments. They found that the more animals lost or removed, the shakier or more biased the experimental conclusions.
“The study began with an attempt to look at the robustness of findings in a handful of preclinical papers, but the sheer number of missing animals stopped us in our tracks,” said author Constance Holman, a graduate student at Charité Universitätsmedizin.
Researchers from both studies believe their findings add to the list of concerns about bias and reporting in research, but the results also establish ways in which research can become more transparent and potentially more reproducible. ![]()

Photo by Bill Branson
Two new studies suggest biomedical research may be hindered by poor reporting and a lack of transparency.
In one study, researchers analyzed more than 400 biomedical science articles and found the papers rarely provided full protocol information, complete data, and the necessary level of transparency to verify or replicate the work.
In the other study, researchers analyzed more than 500 preclinical experiments and found that most didn’t contain sufficient information on the animals used.
Both studies were published in PLOS Biology.
For the first study, Shareen Iqbal, PhD, of Emory University in Atlanta, Georgia, and her colleagues analyzed papers published between 2000 and 2014.
The team set out to determine the extent to which researchers report key information necessary for properly evaluating and replicating published research, including availability of protocols, data, and the frequency of published novel or replication studies.
Out of 441 articles drawn from across the biomedical literature, only 1 paper provided a full protocol, and none of the papers made all the data available. The majority of studies didn’t state funding or conflicts of interest, and replication studies were very rare.
Dr Iqbal and her colleagues said they hope their study will further sensitize scientists, funders, journals, and other science-related stakeholders about the need to improve these indicators.
For the second study, Ulrich Dirnagl, MD, of Charité Universitätsmedizin in Berlin, Germany, and his colleagues examined 100 papers describing preclinical research on stroke and cancer. These papers contained accounts of 316 experiments on infarct volume and 206 experiments on tumor shrinkage.
The vast majority of the reports didn’t contain sufficient information on how many animals were used in the experiments. What’s more, in many papers, animals “vanished” over the course of the study.
Using a computer model, the researchers simulated the effects of such animal loss on the validity of the experiments. They found that the more animals lost or removed, the shakier or more biased the experimental conclusions.
“The study began with an attempt to look at the robustness of findings in a handful of preclinical papers, but the sheer number of missing animals stopped us in our tracks,” said author Constance Holman, a graduate student at Charité Universitätsmedizin.
Researchers from both studies believe their findings add to the list of concerns about bias and reporting in research, but the results also establish ways in which research can become more transparent and potentially more reproducible. ![]()

Photo by Bill Branson
Two new studies suggest biomedical research may be hindered by poor reporting and a lack of transparency.
In one study, researchers analyzed more than 400 biomedical science articles and found the papers rarely provided full protocol information, complete data, and the necessary level of transparency to verify or replicate the work.
In the other study, researchers analyzed more than 500 preclinical experiments and found that most didn’t contain sufficient information on the animals used.
Both studies were published in PLOS Biology.
For the first study, Shareen Iqbal, PhD, of Emory University in Atlanta, Georgia, and her colleagues analyzed papers published between 2000 and 2014.
The team set out to determine the extent to which researchers report key information necessary for properly evaluating and replicating published research, including availability of protocols, data, and the frequency of published novel or replication studies.
Out of 441 articles drawn from across the biomedical literature, only 1 paper provided a full protocol, and none of the papers made all the data available. The majority of studies didn’t state funding or conflicts of interest, and replication studies were very rare.
Dr Iqbal and her colleagues said they hope their study will further sensitize scientists, funders, journals, and other science-related stakeholders about the need to improve these indicators.
For the second study, Ulrich Dirnagl, MD, of Charité Universitätsmedizin in Berlin, Germany, and his colleagues examined 100 papers describing preclinical research on stroke and cancer. These papers contained accounts of 316 experiments on infarct volume and 206 experiments on tumor shrinkage.
The vast majority of the reports didn’t contain sufficient information on how many animals were used in the experiments. What’s more, in many papers, animals “vanished” over the course of the study.
Using a computer model, the researchers simulated the effects of such animal loss on the validity of the experiments. They found that the more animals lost or removed, the shakier or more biased the experimental conclusions.
“The study began with an attempt to look at the robustness of findings in a handful of preclinical papers, but the sheer number of missing animals stopped us in our tracks,” said author Constance Holman, a graduate student at Charité Universitätsmedizin.
Researchers from both studies believe their findings add to the list of concerns about bias and reporting in research, but the results also establish ways in which research can become more transparent and potentially more reproducible. ![]()
Drug granted orphan designation for SCD
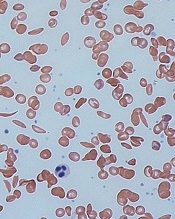
red blood cells
Image by Graham Beards
The US Food and Drug Administration (FDA) has granted orphan drug designation for the small molecule GBT440 to treat patients with sickle cell disease (SCD).
GBT440 is being developed as a potentially disease-modifying therapy for SCD. The drug works by increasing hemoglobin’s affinity for oxygen.
Since oxygenated sickle hemoglobin does not polymerize, it is believed that GBT440 blocks polymerization and the resultant sickling of red blood cells.
If GBT440 can restore normal hemoglobin function and improve oxygen delivery, the drug may be capable of modifying the progression of SCD.
“Receiving orphan drug designation, along with the previously announced fast track designation, are important milestones in our regulatory strategy for GBT440 and highlight the FDA’s agreement that the SCD community faces a critical need for new treatments,” said Ted W. Love, MD, chief executive officer of Global Blood Therapeutics, Inc., the company developing GBT440.
The FDA grants orphan designation to drugs that are intended to treat diseases or conditions affecting fewer than 200,000 patients in the US. The designation provides the drug’s sponsor with various development incentives, including opportunities to apply for research-related tax credits and grant funding, assistance in designing clinical trials, and 7 years of US market exclusivity if the drug is approved.
The FDA grants fast track designation to facilitate and expedite the development and review of new drugs intended to treat serious or life-threatening conditions and address unmet medical need. Through the fast track program, a drug may be eligible for priority review and rolling review, and the company developing the drug may receive additional help from the FDA to expedite development.
GBT440 trial
Early results from an ongoing phase 1/2 study of GBT440 were presented at the 2015 ASH Annual Meeting last month (abstract 542*).
The trial, which includes healthy subjects and patients with SCD, is being conducted in 2 parts: part A (single-dose administration) and part B (multiple-dose administration, daily for 15 days in healthy subjects and 28 days in SCD patients).
As of November 20, 2015, 8 SCD patients completed part A, and 30 SCD patients had either completed or were ongoing in part B.
Of the 30 SCD patients, 16 patients completed 700 mg daily dosing and follow-up (12 on GBT440 and 4 on placebo), and 14 patients completed or were ongoing at 500 mg daily dosing and follow-up (10 on GBT440 and 4 on placebo). A cohort of SCD patients on 1000 mg per day for 28 days is currently enrolling.
Thus far, GBT440 treatment has conferred several improvements from baseline to day 28.
Hemoglobin increases were evident by day 4 of treatment. And the researchers observed absolute hemoglobin increases of 0.5 and 0.7 g/dL with GBT440 at 500 and 700 mg, respectively, compared with a 0.1 g/dL decrease with placebo.
The median reticulocyte count decreased by 31% and 37% with GBT440 at 500 and 700 mg, respectively, compared with a 7% increase with placebo, indicating that the hemoglobin rise is due to decreased hemolysis.
Median erythropoietin levels decreased by 9 and 18 mU/mL with GBT440 at 500 and 700 mg, respectively, compared with an increase of 28 mU/mL with placebo.
Median unconjugated bilirubin levels decrease by 31% and 43% with GBT440 at 500 mg and 700 mg, respectively, compared with an increase of 2% with placebo.
Median lactate dehydrogenase levels decreased by 20% and 12% with GBT440 at 500 and 700 mg, respectively, compared with a decrease of 7% with placebo.
Median sickle cell counts decreased by 56% and 46% with GBT440 at 500 and 700 mg, respectively, compared with a 14% increase with placebo.
The researchers noted high inter- and intra-patient variability in circulating sickle cell counts.
They said inflammatory soluble adhesion molecules for the 700 mg dose cohort showed promising trends in improvement. The median P-selectin decreased 19%, compared with an increase of 20% with placebo. And the median ICAM-1 decreased 6%, compared with an increase of 33% in placebo. Data for the 500 mg dose cohort has not yet been analyzed.
The researchers said pharmacokinetic data demonstrated linear and dose-proportional properties, with a half-life amenable to once-daily dosing.
And GBT440 was well tolerated over the 28 days of dosing. None of the SCD patients discontinued GBT440. The most common adverse event was headache, and there have been no serious adverse events thought to be drug-related.
“We continue to believe that GBT440 has the potential to become the first mechanism-based and disease-modifying therapeutic for this grievous disease and look forward to sharing full results from our phase 1/2 trial and potentially initiating a pivotal trial in adult patients with SCD in 2016,” Dr Love said. ![]()
*Data in the abstract differ from the presentation.

red blood cells
Image by Graham Beards
The US Food and Drug Administration (FDA) has granted orphan drug designation for the small molecule GBT440 to treat patients with sickle cell disease (SCD).
GBT440 is being developed as a potentially disease-modifying therapy for SCD. The drug works by increasing hemoglobin’s affinity for oxygen.
Since oxygenated sickle hemoglobin does not polymerize, it is believed that GBT440 blocks polymerization and the resultant sickling of red blood cells.
If GBT440 can restore normal hemoglobin function and improve oxygen delivery, the drug may be capable of modifying the progression of SCD.
“Receiving orphan drug designation, along with the previously announced fast track designation, are important milestones in our regulatory strategy for GBT440 and highlight the FDA’s agreement that the SCD community faces a critical need for new treatments,” said Ted W. Love, MD, chief executive officer of Global Blood Therapeutics, Inc., the company developing GBT440.
The FDA grants orphan designation to drugs that are intended to treat diseases or conditions affecting fewer than 200,000 patients in the US. The designation provides the drug’s sponsor with various development incentives, including opportunities to apply for research-related tax credits and grant funding, assistance in designing clinical trials, and 7 years of US market exclusivity if the drug is approved.
The FDA grants fast track designation to facilitate and expedite the development and review of new drugs intended to treat serious or life-threatening conditions and address unmet medical need. Through the fast track program, a drug may be eligible for priority review and rolling review, and the company developing the drug may receive additional help from the FDA to expedite development.
GBT440 trial
Early results from an ongoing phase 1/2 study of GBT440 were presented at the 2015 ASH Annual Meeting last month (abstract 542*).
The trial, which includes healthy subjects and patients with SCD, is being conducted in 2 parts: part A (single-dose administration) and part B (multiple-dose administration, daily for 15 days in healthy subjects and 28 days in SCD patients).
As of November 20, 2015, 8 SCD patients completed part A, and 30 SCD patients had either completed or were ongoing in part B.
Of the 30 SCD patients, 16 patients completed 700 mg daily dosing and follow-up (12 on GBT440 and 4 on placebo), and 14 patients completed or were ongoing at 500 mg daily dosing and follow-up (10 on GBT440 and 4 on placebo). A cohort of SCD patients on 1000 mg per day for 28 days is currently enrolling.
Thus far, GBT440 treatment has conferred several improvements from baseline to day 28.
Hemoglobin increases were evident by day 4 of treatment. And the researchers observed absolute hemoglobin increases of 0.5 and 0.7 g/dL with GBT440 at 500 and 700 mg, respectively, compared with a 0.1 g/dL decrease with placebo.
The median reticulocyte count decreased by 31% and 37% with GBT440 at 500 and 700 mg, respectively, compared with a 7% increase with placebo, indicating that the hemoglobin rise is due to decreased hemolysis.
Median erythropoietin levels decreased by 9 and 18 mU/mL with GBT440 at 500 and 700 mg, respectively, compared with an increase of 28 mU/mL with placebo.
Median unconjugated bilirubin levels decrease by 31% and 43% with GBT440 at 500 mg and 700 mg, respectively, compared with an increase of 2% with placebo.
Median lactate dehydrogenase levels decreased by 20% and 12% with GBT440 at 500 and 700 mg, respectively, compared with a decrease of 7% with placebo.
Median sickle cell counts decreased by 56% and 46% with GBT440 at 500 and 700 mg, respectively, compared with a 14% increase with placebo.
The researchers noted high inter- and intra-patient variability in circulating sickle cell counts.
They said inflammatory soluble adhesion molecules for the 700 mg dose cohort showed promising trends in improvement. The median P-selectin decreased 19%, compared with an increase of 20% with placebo. And the median ICAM-1 decreased 6%, compared with an increase of 33% in placebo. Data for the 500 mg dose cohort has not yet been analyzed.
The researchers said pharmacokinetic data demonstrated linear and dose-proportional properties, with a half-life amenable to once-daily dosing.
And GBT440 was well tolerated over the 28 days of dosing. None of the SCD patients discontinued GBT440. The most common adverse event was headache, and there have been no serious adverse events thought to be drug-related.
“We continue to believe that GBT440 has the potential to become the first mechanism-based and disease-modifying therapeutic for this grievous disease and look forward to sharing full results from our phase 1/2 trial and potentially initiating a pivotal trial in adult patients with SCD in 2016,” Dr Love said. ![]()
*Data in the abstract differ from the presentation.

red blood cells
Image by Graham Beards
The US Food and Drug Administration (FDA) has granted orphan drug designation for the small molecule GBT440 to treat patients with sickle cell disease (SCD).
GBT440 is being developed as a potentially disease-modifying therapy for SCD. The drug works by increasing hemoglobin’s affinity for oxygen.
Since oxygenated sickle hemoglobin does not polymerize, it is believed that GBT440 blocks polymerization and the resultant sickling of red blood cells.
If GBT440 can restore normal hemoglobin function and improve oxygen delivery, the drug may be capable of modifying the progression of SCD.
“Receiving orphan drug designation, along with the previously announced fast track designation, are important milestones in our regulatory strategy for GBT440 and highlight the FDA’s agreement that the SCD community faces a critical need for new treatments,” said Ted W. Love, MD, chief executive officer of Global Blood Therapeutics, Inc., the company developing GBT440.
The FDA grants orphan designation to drugs that are intended to treat diseases or conditions affecting fewer than 200,000 patients in the US. The designation provides the drug’s sponsor with various development incentives, including opportunities to apply for research-related tax credits and grant funding, assistance in designing clinical trials, and 7 years of US market exclusivity if the drug is approved.
The FDA grants fast track designation to facilitate and expedite the development and review of new drugs intended to treat serious or life-threatening conditions and address unmet medical need. Through the fast track program, a drug may be eligible for priority review and rolling review, and the company developing the drug may receive additional help from the FDA to expedite development.
GBT440 trial
Early results from an ongoing phase 1/2 study of GBT440 were presented at the 2015 ASH Annual Meeting last month (abstract 542*).
The trial, which includes healthy subjects and patients with SCD, is being conducted in 2 parts: part A (single-dose administration) and part B (multiple-dose administration, daily for 15 days in healthy subjects and 28 days in SCD patients).
As of November 20, 2015, 8 SCD patients completed part A, and 30 SCD patients had either completed or were ongoing in part B.
Of the 30 SCD patients, 16 patients completed 700 mg daily dosing and follow-up (12 on GBT440 and 4 on placebo), and 14 patients completed or were ongoing at 500 mg daily dosing and follow-up (10 on GBT440 and 4 on placebo). A cohort of SCD patients on 1000 mg per day for 28 days is currently enrolling.
Thus far, GBT440 treatment has conferred several improvements from baseline to day 28.
Hemoglobin increases were evident by day 4 of treatment. And the researchers observed absolute hemoglobin increases of 0.5 and 0.7 g/dL with GBT440 at 500 and 700 mg, respectively, compared with a 0.1 g/dL decrease with placebo.
The median reticulocyte count decreased by 31% and 37% with GBT440 at 500 and 700 mg, respectively, compared with a 7% increase with placebo, indicating that the hemoglobin rise is due to decreased hemolysis.
Median erythropoietin levels decreased by 9 and 18 mU/mL with GBT440 at 500 and 700 mg, respectively, compared with an increase of 28 mU/mL with placebo.
Median unconjugated bilirubin levels decrease by 31% and 43% with GBT440 at 500 mg and 700 mg, respectively, compared with an increase of 2% with placebo.
Median lactate dehydrogenase levels decreased by 20% and 12% with GBT440 at 500 and 700 mg, respectively, compared with a decrease of 7% with placebo.
Median sickle cell counts decreased by 56% and 46% with GBT440 at 500 and 700 mg, respectively, compared with a 14% increase with placebo.
The researchers noted high inter- and intra-patient variability in circulating sickle cell counts.
They said inflammatory soluble adhesion molecules for the 700 mg dose cohort showed promising trends in improvement. The median P-selectin decreased 19%, compared with an increase of 20% with placebo. And the median ICAM-1 decreased 6%, compared with an increase of 33% in placebo. Data for the 500 mg dose cohort has not yet been analyzed.
The researchers said pharmacokinetic data demonstrated linear and dose-proportional properties, with a half-life amenable to once-daily dosing.
And GBT440 was well tolerated over the 28 days of dosing. None of the SCD patients discontinued GBT440. The most common adverse event was headache, and there have been no serious adverse events thought to be drug-related.
“We continue to believe that GBT440 has the potential to become the first mechanism-based and disease-modifying therapeutic for this grievous disease and look forward to sharing full results from our phase 1/2 trial and potentially initiating a pivotal trial in adult patients with SCD in 2016,” Dr Love said. ![]()
*Data in the abstract differ from the presentation.
Docs’ body language may convey racial bias

Doctors may convey racial bias with their body language, according to research published in The Journal of Pain and Symptom Management.
In this small study, a group of physicians, most of whom were white males, gave less compassionate nonverbal cues when interacting with black actors portraying seriously ill patients than when interacting with white actors portraying seriously ill patients.
“Although we found that physicians said the same things to their black and white patients, communication is not just the spoken word. It also involves nonverbal cues, such as eye contact, body positioning, and touch,” said study author Amber Barnato, MD, of the University of Pittsburg in Pennsylvania.
“Poor nonverbal communication—something the physician may not even be aware he or she is doing—could explain why many black patients perceive discrimination in the healthcare setting.”
For this study, Dr Barnato and her colleagues recruited 33 hospital-based attending emergency medicine physicians, hospitalists, and intensivists from Allegheny County, Pennsylvania, and put them in realistic simulations where actors portrayed dying black and white patients accompanied by a family member.
The actors portrayed comparable medical conditions—plummeting vital signs related to either metastatic gastric or pancreatic cancer—and read from matching scripts. The physicians were unaware of what the trial was testing.
The majority of the physicians were white men, so the researchers could not derive any statistically significant conclusions about whether a physician’s race impacted his or her actions.
Physicians were scored on a point system for both their verbal and nonverbal communication skills when interacting with the patient and family member. The physicians averaged 7% lower scores for their nonverbal interactions with the black patients than with the white patients.
“When explaining what was happening and what the next steps for care could be, with the white patients, the physicians were more likely to stand right at the patient’s bedside and touch them in a sympathetic manner,” Dr Barnato said.
She explained that something as simple as a physician staying near the door and holding a binder in front of his body could be perceived by the patient and family as defensive or disengaged. This could lead to a cascade of misunderstandings that result in patients and their families requesting extraordinary life-saving measures because they don’t trust the doctor has their best interests in mind when suggesting gentler, end-of-life care options.
“When you survey people in the community about their feelings on end-of-life care, blacks are only slightly more likely than whites to say they want aggressive, life-sustaining measures when terminally ill,” Dr Barnato said.
“However, blacks are much more likely than whites to request such care when they are faced with making the decision in the hospital. Body language is a significant tool in building trust—or mistrust—and physicians need to ensure that their body language isn’t contributing to that decision.”
“To help black patients and their families feel welcome and encouraged to be partners in medical decision-making, it is critical that doctors be aware of their verbal and nonverbal communication and any unintentional biases.” ![]()

Doctors may convey racial bias with their body language, according to research published in The Journal of Pain and Symptom Management.
In this small study, a group of physicians, most of whom were white males, gave less compassionate nonverbal cues when interacting with black actors portraying seriously ill patients than when interacting with white actors portraying seriously ill patients.
“Although we found that physicians said the same things to their black and white patients, communication is not just the spoken word. It also involves nonverbal cues, such as eye contact, body positioning, and touch,” said study author Amber Barnato, MD, of the University of Pittsburg in Pennsylvania.
“Poor nonverbal communication—something the physician may not even be aware he or she is doing—could explain why many black patients perceive discrimination in the healthcare setting.”
For this study, Dr Barnato and her colleagues recruited 33 hospital-based attending emergency medicine physicians, hospitalists, and intensivists from Allegheny County, Pennsylvania, and put them in realistic simulations where actors portrayed dying black and white patients accompanied by a family member.
The actors portrayed comparable medical conditions—plummeting vital signs related to either metastatic gastric or pancreatic cancer—and read from matching scripts. The physicians were unaware of what the trial was testing.
The majority of the physicians were white men, so the researchers could not derive any statistically significant conclusions about whether a physician’s race impacted his or her actions.
Physicians were scored on a point system for both their verbal and nonverbal communication skills when interacting with the patient and family member. The physicians averaged 7% lower scores for their nonverbal interactions with the black patients than with the white patients.
“When explaining what was happening and what the next steps for care could be, with the white patients, the physicians were more likely to stand right at the patient’s bedside and touch them in a sympathetic manner,” Dr Barnato said.
She explained that something as simple as a physician staying near the door and holding a binder in front of his body could be perceived by the patient and family as defensive or disengaged. This could lead to a cascade of misunderstandings that result in patients and their families requesting extraordinary life-saving measures because they don’t trust the doctor has their best interests in mind when suggesting gentler, end-of-life care options.
“When you survey people in the community about their feelings on end-of-life care, blacks are only slightly more likely than whites to say they want aggressive, life-sustaining measures when terminally ill,” Dr Barnato said.
“However, blacks are much more likely than whites to request such care when they are faced with making the decision in the hospital. Body language is a significant tool in building trust—or mistrust—and physicians need to ensure that their body language isn’t contributing to that decision.”
“To help black patients and their families feel welcome and encouraged to be partners in medical decision-making, it is critical that doctors be aware of their verbal and nonverbal communication and any unintentional biases.” ![]()

Doctors may convey racial bias with their body language, according to research published in The Journal of Pain and Symptom Management.
In this small study, a group of physicians, most of whom were white males, gave less compassionate nonverbal cues when interacting with black actors portraying seriously ill patients than when interacting with white actors portraying seriously ill patients.
“Although we found that physicians said the same things to their black and white patients, communication is not just the spoken word. It also involves nonverbal cues, such as eye contact, body positioning, and touch,” said study author Amber Barnato, MD, of the University of Pittsburg in Pennsylvania.
“Poor nonverbal communication—something the physician may not even be aware he or she is doing—could explain why many black patients perceive discrimination in the healthcare setting.”
For this study, Dr Barnato and her colleagues recruited 33 hospital-based attending emergency medicine physicians, hospitalists, and intensivists from Allegheny County, Pennsylvania, and put them in realistic simulations where actors portrayed dying black and white patients accompanied by a family member.
The actors portrayed comparable medical conditions—plummeting vital signs related to either metastatic gastric or pancreatic cancer—and read from matching scripts. The physicians were unaware of what the trial was testing.
The majority of the physicians were white men, so the researchers could not derive any statistically significant conclusions about whether a physician’s race impacted his or her actions.
Physicians were scored on a point system for both their verbal and nonverbal communication skills when interacting with the patient and family member. The physicians averaged 7% lower scores for their nonverbal interactions with the black patients than with the white patients.
“When explaining what was happening and what the next steps for care could be, with the white patients, the physicians were more likely to stand right at the patient’s bedside and touch them in a sympathetic manner,” Dr Barnato said.
She explained that something as simple as a physician staying near the door and holding a binder in front of his body could be perceived by the patient and family as defensive or disengaged. This could lead to a cascade of misunderstandings that result in patients and their families requesting extraordinary life-saving measures because they don’t trust the doctor has their best interests in mind when suggesting gentler, end-of-life care options.
“When you survey people in the community about their feelings on end-of-life care, blacks are only slightly more likely than whites to say they want aggressive, life-sustaining measures when terminally ill,” Dr Barnato said.
“However, blacks are much more likely than whites to request such care when they are faced with making the decision in the hospital. Body language is a significant tool in building trust—or mistrust—and physicians need to ensure that their body language isn’t contributing to that decision.”
“To help black patients and their families feel welcome and encouraged to be partners in medical decision-making, it is critical that doctors be aware of their verbal and nonverbal communication and any unintentional biases.” ![]()
Cancer drug discovery database goes 3D

Photo by Rhoda Baer
Researchers have updated the canSAR database, a tool designed to aid cancer drug discovery, by adding 3D structures of faulty proteins and maps of cancer’s communication networks.
The canSAR database brings together biological, chemical, and pharmacological data.
The goal of the database is to make these data accessible to researchers worldwide to help with hypothesis generation and support drug discovery decisions.
Users can search canSAR using text queries, protein/gene name searches, any keyword searches, chemical structure searches, and sequence similarity searches. Users can also explore and filter chemical compound sets, view experimental data, and produce summary plots.
The canSAR database was launched in 2011 with the goal of using Big Data approaches to build a detailed picture of how the majority of known human molecules behave.
The database has already collated billions of experimental measurements, mapping the actions of 1 million drugs and chemicals on human proteins, and it has combined these data with genetic information and results from clinical trials.
The updated version of canSAR uses artificial intelligence to identify nooks and crannies on the surface of faulty cancer-causing molecules as a key step in designing new drugs to block them. It also allows researchers to identify communication lines that can be intercepted within tumor cells, opening up potential new approaches for cancer treatment.
The growing database now holds the 3D structures of almost 3 million cavities on the surface of nearly 110,000 molecules.
“Our database is constantly growing with information and is the largest of its kind, with more than 140,000 users from over 175 countries,” said Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK.
“And we regularly develop new artificial intelligence technologies that help scientists make predictions and design experiments. Our aim is that cancer scientists will be armed with the data they need to carry out life-saving research into the most exciting drugs of the future.”
“Scientists need to find all the information there is about a faulty gene or protein to understand whether a new drug might work. These data are vast and scattered, but the canSAR database brings them together and adds value by identifying hidden links and presenting the key information easily.”
Details on the updates to canSAR have been published in Nucleic Acid Research. The database is available online at https://cansar.icr.ac.uk/. ![]()

Photo by Rhoda Baer
Researchers have updated the canSAR database, a tool designed to aid cancer drug discovery, by adding 3D structures of faulty proteins and maps of cancer’s communication networks.
The canSAR database brings together biological, chemical, and pharmacological data.
The goal of the database is to make these data accessible to researchers worldwide to help with hypothesis generation and support drug discovery decisions.
Users can search canSAR using text queries, protein/gene name searches, any keyword searches, chemical structure searches, and sequence similarity searches. Users can also explore and filter chemical compound sets, view experimental data, and produce summary plots.
The canSAR database was launched in 2011 with the goal of using Big Data approaches to build a detailed picture of how the majority of known human molecules behave.
The database has already collated billions of experimental measurements, mapping the actions of 1 million drugs and chemicals on human proteins, and it has combined these data with genetic information and results from clinical trials.
The updated version of canSAR uses artificial intelligence to identify nooks and crannies on the surface of faulty cancer-causing molecules as a key step in designing new drugs to block them. It also allows researchers to identify communication lines that can be intercepted within tumor cells, opening up potential new approaches for cancer treatment.
The growing database now holds the 3D structures of almost 3 million cavities on the surface of nearly 110,000 molecules.
“Our database is constantly growing with information and is the largest of its kind, with more than 140,000 users from over 175 countries,” said Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK.
“And we regularly develop new artificial intelligence technologies that help scientists make predictions and design experiments. Our aim is that cancer scientists will be armed with the data they need to carry out life-saving research into the most exciting drugs of the future.”
“Scientists need to find all the information there is about a faulty gene or protein to understand whether a new drug might work. These data are vast and scattered, but the canSAR database brings them together and adds value by identifying hidden links and presenting the key information easily.”
Details on the updates to canSAR have been published in Nucleic Acid Research. The database is available online at https://cansar.icr.ac.uk/. ![]()

Photo by Rhoda Baer
Researchers have updated the canSAR database, a tool designed to aid cancer drug discovery, by adding 3D structures of faulty proteins and maps of cancer’s communication networks.
The canSAR database brings together biological, chemical, and pharmacological data.
The goal of the database is to make these data accessible to researchers worldwide to help with hypothesis generation and support drug discovery decisions.
Users can search canSAR using text queries, protein/gene name searches, any keyword searches, chemical structure searches, and sequence similarity searches. Users can also explore and filter chemical compound sets, view experimental data, and produce summary plots.
The canSAR database was launched in 2011 with the goal of using Big Data approaches to build a detailed picture of how the majority of known human molecules behave.
The database has already collated billions of experimental measurements, mapping the actions of 1 million drugs and chemicals on human proteins, and it has combined these data with genetic information and results from clinical trials.
The updated version of canSAR uses artificial intelligence to identify nooks and crannies on the surface of faulty cancer-causing molecules as a key step in designing new drugs to block them. It also allows researchers to identify communication lines that can be intercepted within tumor cells, opening up potential new approaches for cancer treatment.
The growing database now holds the 3D structures of almost 3 million cavities on the surface of nearly 110,000 molecules.
“Our database is constantly growing with information and is the largest of its kind, with more than 140,000 users from over 175 countries,” said Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK.
“And we regularly develop new artificial intelligence technologies that help scientists make predictions and design experiments. Our aim is that cancer scientists will be armed with the data they need to carry out life-saving research into the most exciting drugs of the future.”
“Scientists need to find all the information there is about a faulty gene or protein to understand whether a new drug might work. These data are vast and scattered, but the canSAR database brings them together and adds value by identifying hidden links and presenting the key information easily.”
Details on the updates to canSAR have been published in Nucleic Acid Research. The database is available online at https://cansar.icr.ac.uk/. ![]()
Team discovers virus linked to HCV
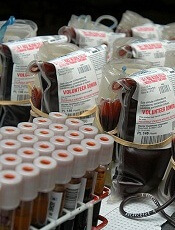
Photo by Daniel Gay
Researchers have discovered a bloodborne virus, known as human pegivirus 2 (HPgV-2), in patients with hepatitis C virus (HCV).
The team identified 8 complete strains of HPgV-2 and noted that the virus was only found in patients who tested positive for HCV RNA. However, it’s not clear if HPgV-2 causes hepatitis.
Charles Chiu, MD, PhD, of the University of California San Francisco, and his colleagues described their discovery of HPgV-2 in PLOS Pathogens.
The team identified the virus by sequencing plasma from an HCV-infected patient with multiple bloodborne exposures who died from sepsis of unknown etiology.
They said HPgV-2 is “highly divergent,” sharing less than 32% amino acid identity with its nearest relatives, rodent and bat pegiviruses.
After their initial discovery, the researchers screened an additional 2440 plasma samples and found 11 HPgV-2 RNA-positive samples.
All 12 HPgV-2 RNA-positive cases were found in patients who tested positive for HCV RNA, including 2 patients who were also infected with HIV.
The researchers performed longitudinal sampling in 2 patients and found that active HPgV-2 infection can persist in blood for at least 7 weeks, despite the presence of virus-specific antibodies.
The team also identified 1 patient with HPgV-2 and HCV RNA who was seronegative for both viruses. They said this suggests a high likelihood of simultaneous acquisition of HCV and HPgV-2 infection from an acute co-transmission event.
“Based on our findings, our team used the genetic makeup of the virus to develop both a molecular test for detecting it in the bloodstream and an antibody test for determining an immune response to the virus,” said John Hackett Jr, PhD, of Abbott Laboratories, Inc. in Abbott Park, Illinois.
“Our next step is to explore whether this new virus can cause disease, and if so, work with blood banks to continue to help safeguard the world’s blood supply against these types of new viruses. Research such as this is ultimately focused on unlocking new technologies that hold the potential for significant improvements to the practice of healthcare.” ![]()

Photo by Daniel Gay
Researchers have discovered a bloodborne virus, known as human pegivirus 2 (HPgV-2), in patients with hepatitis C virus (HCV).
The team identified 8 complete strains of HPgV-2 and noted that the virus was only found in patients who tested positive for HCV RNA. However, it’s not clear if HPgV-2 causes hepatitis.
Charles Chiu, MD, PhD, of the University of California San Francisco, and his colleagues described their discovery of HPgV-2 in PLOS Pathogens.
The team identified the virus by sequencing plasma from an HCV-infected patient with multiple bloodborne exposures who died from sepsis of unknown etiology.
They said HPgV-2 is “highly divergent,” sharing less than 32% amino acid identity with its nearest relatives, rodent and bat pegiviruses.
After their initial discovery, the researchers screened an additional 2440 plasma samples and found 11 HPgV-2 RNA-positive samples.
All 12 HPgV-2 RNA-positive cases were found in patients who tested positive for HCV RNA, including 2 patients who were also infected with HIV.
The researchers performed longitudinal sampling in 2 patients and found that active HPgV-2 infection can persist in blood for at least 7 weeks, despite the presence of virus-specific antibodies.
The team also identified 1 patient with HPgV-2 and HCV RNA who was seronegative for both viruses. They said this suggests a high likelihood of simultaneous acquisition of HCV and HPgV-2 infection from an acute co-transmission event.
“Based on our findings, our team used the genetic makeup of the virus to develop both a molecular test for detecting it in the bloodstream and an antibody test for determining an immune response to the virus,” said John Hackett Jr, PhD, of Abbott Laboratories, Inc. in Abbott Park, Illinois.
“Our next step is to explore whether this new virus can cause disease, and if so, work with blood banks to continue to help safeguard the world’s blood supply against these types of new viruses. Research such as this is ultimately focused on unlocking new technologies that hold the potential for significant improvements to the practice of healthcare.” ![]()

Photo by Daniel Gay
Researchers have discovered a bloodborne virus, known as human pegivirus 2 (HPgV-2), in patients with hepatitis C virus (HCV).
The team identified 8 complete strains of HPgV-2 and noted that the virus was only found in patients who tested positive for HCV RNA. However, it’s not clear if HPgV-2 causes hepatitis.
Charles Chiu, MD, PhD, of the University of California San Francisco, and his colleagues described their discovery of HPgV-2 in PLOS Pathogens.
The team identified the virus by sequencing plasma from an HCV-infected patient with multiple bloodborne exposures who died from sepsis of unknown etiology.
They said HPgV-2 is “highly divergent,” sharing less than 32% amino acid identity with its nearest relatives, rodent and bat pegiviruses.
After their initial discovery, the researchers screened an additional 2440 plasma samples and found 11 HPgV-2 RNA-positive samples.
All 12 HPgV-2 RNA-positive cases were found in patients who tested positive for HCV RNA, including 2 patients who were also infected with HIV.
The researchers performed longitudinal sampling in 2 patients and found that active HPgV-2 infection can persist in blood for at least 7 weeks, despite the presence of virus-specific antibodies.
The team also identified 1 patient with HPgV-2 and HCV RNA who was seronegative for both viruses. They said this suggests a high likelihood of simultaneous acquisition of HCV and HPgV-2 infection from an acute co-transmission event.
“Based on our findings, our team used the genetic makeup of the virus to develop both a molecular test for detecting it in the bloodstream and an antibody test for determining an immune response to the virus,” said John Hackett Jr, PhD, of Abbott Laboratories, Inc. in Abbott Park, Illinois.
“Our next step is to explore whether this new virus can cause disease, and if so, work with blood banks to continue to help safeguard the world’s blood supply against these types of new viruses. Research such as this is ultimately focused on unlocking new technologies that hold the potential for significant improvements to the practice of healthcare.” ![]()
Tool may provide new insight into pediatric cancers

and Xin Zhou, PhD
Photo by Peter Barta/St. Jude
Children’s Research Hospital
Researchers say they have developed a tool that may advance our understanding of the mutations that drive pediatric cancers.
The tool, called ProteinPaint, is a web application that allows the user to visualize genetic lesions and RNA expression in pediatric cancers.
ProteinPaint’s infographics let users see all mutations in individual genes and their corresponding proteins, including detailed information about mutation type, frequency in cancer subtype, and location in the protein domain.
That information provides clues about how a change might contribute to cancer’s start, progression, or relapse.
Jinghui Zhang, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and her colleagues described ProteinPaint in a letter to Nature Genetics.
ProteinPaint currently integrates information from 5 studies, but Dr Zhang and her colleagues said the data will be updated as new studies are published.
ProteinPaint now includes information on almost 27,500 mutations discovered in more than 1000 pediatric patients with 21 cancer subtypes. The application also includes RNA-sequencing data from 928 pediatric tumors belonging to 36 different subtypes.
Xin Zhou, PhD, also of St. Jude, developed ProteinPaint’s infographics to display the genomic information in an interactive format. A click of the mouse gives users additional details about the mutations listed, including the pediatric cancer subtype where the change has been validated and a link to the publication.
“ProteinPaint’s focus on pediatric cancer and presentation of mutations at the gene level complements existing cancer genome data portals,” Dr Zhang said. “For St. Jude, the application is the foundation for developing a global reference database for information about pediatric cancer.”
Dr Zhou added that the ProteinPaint software has the potential to help researchers studying other disorders, including sickle cell disease, that involve a mutation that affects protein function.
ProteinPaint is available at no cost to academic researchers who are free to use the tool to analyze their own data. The application also lets researchers compare information about pediatric and adult cancer genomes by providing a parallel view of data from COSMIC, the world’s largest database of somatic mutations, primarily from adult cancer.
ProteinPaint has already been used to study the role played by germline mutations in pediatric cancers. That research was published in NEJM in November.
More information about ProteinPaint is available on the St. Jude PeCan Data Portal. ![]()

and Xin Zhou, PhD
Photo by Peter Barta/St. Jude
Children’s Research Hospital
Researchers say they have developed a tool that may advance our understanding of the mutations that drive pediatric cancers.
The tool, called ProteinPaint, is a web application that allows the user to visualize genetic lesions and RNA expression in pediatric cancers.
ProteinPaint’s infographics let users see all mutations in individual genes and their corresponding proteins, including detailed information about mutation type, frequency in cancer subtype, and location in the protein domain.
That information provides clues about how a change might contribute to cancer’s start, progression, or relapse.
Jinghui Zhang, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and her colleagues described ProteinPaint in a letter to Nature Genetics.
ProteinPaint currently integrates information from 5 studies, but Dr Zhang and her colleagues said the data will be updated as new studies are published.
ProteinPaint now includes information on almost 27,500 mutations discovered in more than 1000 pediatric patients with 21 cancer subtypes. The application also includes RNA-sequencing data from 928 pediatric tumors belonging to 36 different subtypes.
Xin Zhou, PhD, also of St. Jude, developed ProteinPaint’s infographics to display the genomic information in an interactive format. A click of the mouse gives users additional details about the mutations listed, including the pediatric cancer subtype where the change has been validated and a link to the publication.
“ProteinPaint’s focus on pediatric cancer and presentation of mutations at the gene level complements existing cancer genome data portals,” Dr Zhang said. “For St. Jude, the application is the foundation for developing a global reference database for information about pediatric cancer.”
Dr Zhou added that the ProteinPaint software has the potential to help researchers studying other disorders, including sickle cell disease, that involve a mutation that affects protein function.
ProteinPaint is available at no cost to academic researchers who are free to use the tool to analyze their own data. The application also lets researchers compare information about pediatric and adult cancer genomes by providing a parallel view of data from COSMIC, the world’s largest database of somatic mutations, primarily from adult cancer.
ProteinPaint has already been used to study the role played by germline mutations in pediatric cancers. That research was published in NEJM in November.
More information about ProteinPaint is available on the St. Jude PeCan Data Portal. ![]()

and Xin Zhou, PhD
Photo by Peter Barta/St. Jude
Children’s Research Hospital
Researchers say they have developed a tool that may advance our understanding of the mutations that drive pediatric cancers.
The tool, called ProteinPaint, is a web application that allows the user to visualize genetic lesions and RNA expression in pediatric cancers.
ProteinPaint’s infographics let users see all mutations in individual genes and their corresponding proteins, including detailed information about mutation type, frequency in cancer subtype, and location in the protein domain.
That information provides clues about how a change might contribute to cancer’s start, progression, or relapse.
Jinghui Zhang, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee, and her colleagues described ProteinPaint in a letter to Nature Genetics.
ProteinPaint currently integrates information from 5 studies, but Dr Zhang and her colleagues said the data will be updated as new studies are published.
ProteinPaint now includes information on almost 27,500 mutations discovered in more than 1000 pediatric patients with 21 cancer subtypes. The application also includes RNA-sequencing data from 928 pediatric tumors belonging to 36 different subtypes.
Xin Zhou, PhD, also of St. Jude, developed ProteinPaint’s infographics to display the genomic information in an interactive format. A click of the mouse gives users additional details about the mutations listed, including the pediatric cancer subtype where the change has been validated and a link to the publication.
“ProteinPaint’s focus on pediatric cancer and presentation of mutations at the gene level complements existing cancer genome data portals,” Dr Zhang said. “For St. Jude, the application is the foundation for developing a global reference database for information about pediatric cancer.”
Dr Zhou added that the ProteinPaint software has the potential to help researchers studying other disorders, including sickle cell disease, that involve a mutation that affects protein function.
ProteinPaint is available at no cost to academic researchers who are free to use the tool to analyze their own data. The application also lets researchers compare information about pediatric and adult cancer genomes by providing a parallel view of data from COSMIC, the world’s largest database of somatic mutations, primarily from adult cancer.
ProteinPaint has already been used to study the role played by germline mutations in pediatric cancers. That research was published in NEJM in November.
More information about ProteinPaint is available on the St. Jude PeCan Data Portal.
Identifying druggable proteins

Photo by Darren Baker
A computer model that employs techniques used to analyze social networks could aid the development of new cancer treatments, according to researchers.
The model analyzes the unique behaviors of cancer-causing proteins, spotting what makes them different from normal proteins and mapping out molecular targets for drugs that could potentially be developed to treat cancers.
The researchers described this model in PLOS Computational Biology.
Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK, and her colleagues compared proteins to members of an enormous social network, mapping the ways they interact. This allowed the team to predict which proteins might be most effectively targeted with drugs.
Cancer-causing proteins that have already been successfully targeted tended to have particular “social” characteristics that differed from non-cancer proteins. “Hub-like” proteins that were shown to “communicate” with lots of other proteins were more likely to cause cancers.
The researchers said this suggests that previously unexplored cancer proteins with similar characteristics could make good drug targets.
“Our study is the first to identify the rules of social behavior of cancer proteins and use it to predict new targets for potential cancer drugs,” Dr Al-Lazikani said.
“It shows that cancer drug targets behave very differently from normal proteins and often have a complex web of social interactions. Finding new targets is one of the most important steps in drug discovery, but it can be a lengthy, expensive process.”
“The map that we’ve made will help researchers design better new drugs, more quickly, saving time and money. It also sheds light on how resistance to treatments may occur and, in just a few years, could help doctors choose the best drug combinations to suit individual patients.”
All of the researchers’ target predictions are available on the canSAR website. The underlying data and tools are also available on the site.

Photo by Darren Baker
A computer model that employs techniques used to analyze social networks could aid the development of new cancer treatments, according to researchers.
The model analyzes the unique behaviors of cancer-causing proteins, spotting what makes them different from normal proteins and mapping out molecular targets for drugs that could potentially be developed to treat cancers.
The researchers described this model in PLOS Computational Biology.
Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK, and her colleagues compared proteins to members of an enormous social network, mapping the ways they interact. This allowed the team to predict which proteins might be most effectively targeted with drugs.
Cancer-causing proteins that have already been successfully targeted tended to have particular “social” characteristics that differed from non-cancer proteins. “Hub-like” proteins that were shown to “communicate” with lots of other proteins were more likely to cause cancers.
The researchers said this suggests that previously unexplored cancer proteins with similar characteristics could make good drug targets.
“Our study is the first to identify the rules of social behavior of cancer proteins and use it to predict new targets for potential cancer drugs,” Dr Al-Lazikani said.
“It shows that cancer drug targets behave very differently from normal proteins and often have a complex web of social interactions. Finding new targets is one of the most important steps in drug discovery, but it can be a lengthy, expensive process.”
“The map that we’ve made will help researchers design better new drugs, more quickly, saving time and money. It also sheds light on how resistance to treatments may occur and, in just a few years, could help doctors choose the best drug combinations to suit individual patients.”
All of the researchers’ target predictions are available on the canSAR website. The underlying data and tools are also available on the site.

Photo by Darren Baker
A computer model that employs techniques used to analyze social networks could aid the development of new cancer treatments, according to researchers.
The model analyzes the unique behaviors of cancer-causing proteins, spotting what makes them different from normal proteins and mapping out molecular targets for drugs that could potentially be developed to treat cancers.
The researchers described this model in PLOS Computational Biology.
Bissan Al-Lazikani, PhD, of The Institute of Cancer Research in London, UK, and her colleagues compared proteins to members of an enormous social network, mapping the ways they interact. This allowed the team to predict which proteins might be most effectively targeted with drugs.
Cancer-causing proteins that have already been successfully targeted tended to have particular “social” characteristics that differed from non-cancer proteins. “Hub-like” proteins that were shown to “communicate” with lots of other proteins were more likely to cause cancers.
The researchers said this suggests that previously unexplored cancer proteins with similar characteristics could make good drug targets.
“Our study is the first to identify the rules of social behavior of cancer proteins and use it to predict new targets for potential cancer drugs,” Dr Al-Lazikani said.
“It shows that cancer drug targets behave very differently from normal proteins and often have a complex web of social interactions. Finding new targets is one of the most important steps in drug discovery, but it can be a lengthy, expensive process.”
“The map that we’ve made will help researchers design better new drugs, more quickly, saving time and money. It also sheds light on how resistance to treatments may occur and, in just a few years, could help doctors choose the best drug combinations to suit individual patients.”
All of the researchers’ target predictions are available on the canSAR website. The underlying data and tools are also available on the site.
Factors predict low accrual in cancer clinical trials

for a clinical trial
Photo by Esther Dyson
Twelve factors may predict low patient accrual in cancer clinical trials, according to research published in JNCI.
Many studies have been conducted to investigate the perceived barriers to clinical trial accrual from the patient or provider perspective.
However, researchers have rarely taken a trial-level view and investigated why certain trials are able to accrue patients faster than expected while others fail to attract even a fraction of the intended number of participants.
Caroline S. Bennette, PhD, of the University of Washington in Seattle, and her colleagues conducted their study to do just that.
They analyzed information on 787 phase 2/3 clinical trials sponsored by the National Clinical Trials Network (NCTN; formerly the Cooperative Group Program) launched between 2000 and 2011.
After excluding trials that closed because of toxicity or interim results, the researchers found that 145 (18%) NCTN trials closed with low accrual or were accruing at less than 50% of target accrual 3 years or more after opening.
The team identified potential risk factors from the literature and interviews with clinical trial experts and found multiple trial-level factors that were associated with poor accrual to NCTN trials, such as increased competition for patients from currently ongoing trials, planning to enroll a higher proportion of the available patient population, and not evaluating a new investigational agent or targeted therapy.
The researchers then developed a multivariable prediction model of low accrual using 12 trial-level risk factors. The team said these factors had good agreement between predicted and observed risks of low accrual in a preliminary validation using 46 trials opened between 2012 and 2013.
Those 12 risk factors are:
- The number of competing trials per 10,000 eligible patients per year (odds ratio [OR]=1.88)
- Phase 3 vs phase 2 trial (OR=1.86)
- Enrollment as percentage of eligible population for targeted therapy (OR=0.57)
- Enrollment as percentage of eligible population for radiation therapy (OR=1.81)
- Annual incidence of clinical condition(s) per 10,000 (OR=0.99)
- Tissue sample required to assess eligibility (OR=1.26)
- Investigational new drug (OR=0.34)
- Metastatic setting (OR=1.46)
- Sample size per 100 (OR=0.95)
- More than one condition evaluated (OR=1.98)
- Common solid cancer (prostate, breast, lung, or colon) vs liquid or rare solid cancers (OR=2.32)
- Interaction term (phase 3 x investigational new drug, OR=2.47).
The researchers concluded that systematically considering the overall influence of these risk factors could aid in the design and prioritization of future clinical trials.

for a clinical trial
Photo by Esther Dyson
Twelve factors may predict low patient accrual in cancer clinical trials, according to research published in JNCI.
Many studies have been conducted to investigate the perceived barriers to clinical trial accrual from the patient or provider perspective.
However, researchers have rarely taken a trial-level view and investigated why certain trials are able to accrue patients faster than expected while others fail to attract even a fraction of the intended number of participants.
Caroline S. Bennette, PhD, of the University of Washington in Seattle, and her colleagues conducted their study to do just that.
They analyzed information on 787 phase 2/3 clinical trials sponsored by the National Clinical Trials Network (NCTN; formerly the Cooperative Group Program) launched between 2000 and 2011.
After excluding trials that closed because of toxicity or interim results, the researchers found that 145 (18%) NCTN trials closed with low accrual or were accruing at less than 50% of target accrual 3 years or more after opening.
The team identified potential risk factors from the literature and interviews with clinical trial experts and found multiple trial-level factors that were associated with poor accrual to NCTN trials, such as increased competition for patients from currently ongoing trials, planning to enroll a higher proportion of the available patient population, and not evaluating a new investigational agent or targeted therapy.
The researchers then developed a multivariable prediction model of low accrual using 12 trial-level risk factors. The team said these factors had good agreement between predicted and observed risks of low accrual in a preliminary validation using 46 trials opened between 2012 and 2013.
Those 12 risk factors are:
- The number of competing trials per 10,000 eligible patients per year (odds ratio [OR]=1.88)
- Phase 3 vs phase 2 trial (OR=1.86)
- Enrollment as percentage of eligible population for targeted therapy (OR=0.57)
- Enrollment as percentage of eligible population for radiation therapy (OR=1.81)
- Annual incidence of clinical condition(s) per 10,000 (OR=0.99)
- Tissue sample required to assess eligibility (OR=1.26)
- Investigational new drug (OR=0.34)
- Metastatic setting (OR=1.46)
- Sample size per 100 (OR=0.95)
- More than one condition evaluated (OR=1.98)
- Common solid cancer (prostate, breast, lung, or colon) vs liquid or rare solid cancers (OR=2.32)
- Interaction term (phase 3 x investigational new drug, OR=2.47).
The researchers concluded that systematically considering the overall influence of these risk factors could aid in the design and prioritization of future clinical trials.

for a clinical trial
Photo by Esther Dyson
Twelve factors may predict low patient accrual in cancer clinical trials, according to research published in JNCI.
Many studies have been conducted to investigate the perceived barriers to clinical trial accrual from the patient or provider perspective.
However, researchers have rarely taken a trial-level view and investigated why certain trials are able to accrue patients faster than expected while others fail to attract even a fraction of the intended number of participants.
Caroline S. Bennette, PhD, of the University of Washington in Seattle, and her colleagues conducted their study to do just that.
They analyzed information on 787 phase 2/3 clinical trials sponsored by the National Clinical Trials Network (NCTN; formerly the Cooperative Group Program) launched between 2000 and 2011.
After excluding trials that closed because of toxicity or interim results, the researchers found that 145 (18%) NCTN trials closed with low accrual or were accruing at less than 50% of target accrual 3 years or more after opening.
The team identified potential risk factors from the literature and interviews with clinical trial experts and found multiple trial-level factors that were associated with poor accrual to NCTN trials, such as increased competition for patients from currently ongoing trials, planning to enroll a higher proportion of the available patient population, and not evaluating a new investigational agent or targeted therapy.
The researchers then developed a multivariable prediction model of low accrual using 12 trial-level risk factors. The team said these factors had good agreement between predicted and observed risks of low accrual in a preliminary validation using 46 trials opened between 2012 and 2013.
Those 12 risk factors are:
- The number of competing trials per 10,000 eligible patients per year (odds ratio [OR]=1.88)
- Phase 3 vs phase 2 trial (OR=1.86)
- Enrollment as percentage of eligible population for targeted therapy (OR=0.57)
- Enrollment as percentage of eligible population for radiation therapy (OR=1.81)
- Annual incidence of clinical condition(s) per 10,000 (OR=0.99)
- Tissue sample required to assess eligibility (OR=1.26)
- Investigational new drug (OR=0.34)
- Metastatic setting (OR=1.46)
- Sample size per 100 (OR=0.95)
- More than one condition evaluated (OR=1.98)
- Common solid cancer (prostate, breast, lung, or colon) vs liquid or rare solid cancers (OR=2.32)
- Interaction term (phase 3 x investigational new drug, OR=2.47).
The researchers concluded that systematically considering the overall influence of these risk factors could aid in the design and prioritization of future clinical trials.
Why it’s hard to develop immunity against malaria

Diana Hansen, PhD
Photo courtesy of the Walter
and Eliza Hall Institute
Results of preclinical research appear to explain how the malaria parasite Plasmodium falciparum causes an inflammatory reaction that sabotages the body’s ability to protect itself from malaria.
Researchers found evidence to suggest that the same inflammatory molecules that drive the immune response in clinical and severe malaria also prevent the body from developing protective antibodies against the parasite.
They said this discovery opens up the possibility of improving new or existing malaria vaccines by boosting immune cells needed for long-lasting immunity.
Diana Hansen, PhD, of the Walter and Eliza Hall Institute of Medical Research in Parkville, Victoria, Australia, and her colleagues described this work in Cell Reports.
Dr Hansen said this is the first time scientists have pinpointed why the immune system fails to develop immunity during malaria infection.
“With many infections, a single exposure to the pathogen is enough to induce production of antibodies that will protect you for the rest of your life,” she explained. “However, with malaria, it can take up to 20 years for someone to build up sufficient immunity to be protected.”
The fact that the body is not good at developing long-lasting immunity to the parasite has hampered vaccine development, Dr Hansen added.
“This was complicated by the fact that we didn’t know whether it was the malaria parasite itself or the inflammatory reaction to malaria that was actually inhibiting the ability to develop protective immunity,” she said.
“We have now shown that it was a double-edged sword. The strong inflammatory reaction that accompanies and, in fact, drives severe clinical malaria is also responsible for silencing the key immune cells needed for long-term protection against the parasite.”
Dr Hansen and her colleagues conducted experiments in mouse models of malaria and found that inflammatory molecules released by the body to fight the infection were preventing protective antibodies from being made.
“Specialized immune cells called helper T cells join forces with B cells to generate these protective antibodies,” said Axel Kallies, PhD, of the Walter and Eliza Hall Institute of Medical Research.
“However, we showed that, during malaria infection, critical inflammatory molecules actually arrest development of helper T cells, and, therefore, the B cells don’t get the necessary instructions to make antibodies.”
Specifically, the researchers found that severe malaria infection inhibited the establishment of germinal centers in the spleens of the mice. And malaria infection induced high frequencies of T-follicular-helper cell precursors but resulted in impaired T-follicular-helper cell differentiation.
Precursor T-follicular-helper cells induced during infection had low levels of PD-1 and CXCR5 and co-expressed Th1-associated molecules such as T-bet and CXCR3.
However, when the researchers blocked the inflammatory cytokines TNF and IFN-γ or deleted T-bet, they were able to restore T-follicular-helper cell differentiation and germinal center responses to infection.
Dr Hansen said these findings could lead to new avenues in the search for effective malaria vaccines.
“This research opens the door to therapeutic approaches to accelerate development of protective immunity to malaria and improve efficacy of malaria vaccines,” she said.
“Until now, malaria vaccines have had disappointing results. We can now see a way of improving these responses by tailoring or augmenting the vaccine to boost development of helper T cells that will enable the body to make protective antibodies that target the malaria parasites.”

Diana Hansen, PhD
Photo courtesy of the Walter
and Eliza Hall Institute
Results of preclinical research appear to explain how the malaria parasite Plasmodium falciparum causes an inflammatory reaction that sabotages the body’s ability to protect itself from malaria.
Researchers found evidence to suggest that the same inflammatory molecules that drive the immune response in clinical and severe malaria also prevent the body from developing protective antibodies against the parasite.
They said this discovery opens up the possibility of improving new or existing malaria vaccines by boosting immune cells needed for long-lasting immunity.
Diana Hansen, PhD, of the Walter and Eliza Hall Institute of Medical Research in Parkville, Victoria, Australia, and her colleagues described this work in Cell Reports.
Dr Hansen said this is the first time scientists have pinpointed why the immune system fails to develop immunity during malaria infection.
“With many infections, a single exposure to the pathogen is enough to induce production of antibodies that will protect you for the rest of your life,” she explained. “However, with malaria, it can take up to 20 years for someone to build up sufficient immunity to be protected.”
The fact that the body is not good at developing long-lasting immunity to the parasite has hampered vaccine development, Dr Hansen added.
“This was complicated by the fact that we didn’t know whether it was the malaria parasite itself or the inflammatory reaction to malaria that was actually inhibiting the ability to develop protective immunity,” she said.
“We have now shown that it was a double-edged sword. The strong inflammatory reaction that accompanies and, in fact, drives severe clinical malaria is also responsible for silencing the key immune cells needed for long-term protection against the parasite.”
Dr Hansen and her colleagues conducted experiments in mouse models of malaria and found that inflammatory molecules released by the body to fight the infection were preventing protective antibodies from being made.
“Specialized immune cells called helper T cells join forces with B cells to generate these protective antibodies,” said Axel Kallies, PhD, of the Walter and Eliza Hall Institute of Medical Research.
“However, we showed that, during malaria infection, critical inflammatory molecules actually arrest development of helper T cells, and, therefore, the B cells don’t get the necessary instructions to make antibodies.”
Specifically, the researchers found that severe malaria infection inhibited the establishment of germinal centers in the spleens of the mice. And malaria infection induced high frequencies of T-follicular-helper cell precursors but resulted in impaired T-follicular-helper cell differentiation.
Precursor T-follicular-helper cells induced during infection had low levels of PD-1 and CXCR5 and co-expressed Th1-associated molecules such as T-bet and CXCR3.
However, when the researchers blocked the inflammatory cytokines TNF and IFN-γ or deleted T-bet, they were able to restore T-follicular-helper cell differentiation and germinal center responses to infection.
Dr Hansen said these findings could lead to new avenues in the search for effective malaria vaccines.
“This research opens the door to therapeutic approaches to accelerate development of protective immunity to malaria and improve efficacy of malaria vaccines,” she said.
“Until now, malaria vaccines have had disappointing results. We can now see a way of improving these responses by tailoring or augmenting the vaccine to boost development of helper T cells that will enable the body to make protective antibodies that target the malaria parasites.”

Diana Hansen, PhD
Photo courtesy of the Walter
and Eliza Hall Institute
Results of preclinical research appear to explain how the malaria parasite Plasmodium falciparum causes an inflammatory reaction that sabotages the body’s ability to protect itself from malaria.
Researchers found evidence to suggest that the same inflammatory molecules that drive the immune response in clinical and severe malaria also prevent the body from developing protective antibodies against the parasite.
They said this discovery opens up the possibility of improving new or existing malaria vaccines by boosting immune cells needed for long-lasting immunity.
Diana Hansen, PhD, of the Walter and Eliza Hall Institute of Medical Research in Parkville, Victoria, Australia, and her colleagues described this work in Cell Reports.
Dr Hansen said this is the first time scientists have pinpointed why the immune system fails to develop immunity during malaria infection.
“With many infections, a single exposure to the pathogen is enough to induce production of antibodies that will protect you for the rest of your life,” she explained. “However, with malaria, it can take up to 20 years for someone to build up sufficient immunity to be protected.”
The fact that the body is not good at developing long-lasting immunity to the parasite has hampered vaccine development, Dr Hansen added.
“This was complicated by the fact that we didn’t know whether it was the malaria parasite itself or the inflammatory reaction to malaria that was actually inhibiting the ability to develop protective immunity,” she said.
“We have now shown that it was a double-edged sword. The strong inflammatory reaction that accompanies and, in fact, drives severe clinical malaria is also responsible for silencing the key immune cells needed for long-term protection against the parasite.”
Dr Hansen and her colleagues conducted experiments in mouse models of malaria and found that inflammatory molecules released by the body to fight the infection were preventing protective antibodies from being made.
“Specialized immune cells called helper T cells join forces with B cells to generate these protective antibodies,” said Axel Kallies, PhD, of the Walter and Eliza Hall Institute of Medical Research.
“However, we showed that, during malaria infection, critical inflammatory molecules actually arrest development of helper T cells, and, therefore, the B cells don’t get the necessary instructions to make antibodies.”
Specifically, the researchers found that severe malaria infection inhibited the establishment of germinal centers in the spleens of the mice. And malaria infection induced high frequencies of T-follicular-helper cell precursors but resulted in impaired T-follicular-helper cell differentiation.
Precursor T-follicular-helper cells induced during infection had low levels of PD-1 and CXCR5 and co-expressed Th1-associated molecules such as T-bet and CXCR3.
However, when the researchers blocked the inflammatory cytokines TNF and IFN-γ or deleted T-bet, they were able to restore T-follicular-helper cell differentiation and germinal center responses to infection.
Dr Hansen said these findings could lead to new avenues in the search for effective malaria vaccines.
“This research opens the door to therapeutic approaches to accelerate development of protective immunity to malaria and improve efficacy of malaria vaccines,” she said.
“Until now, malaria vaccines have had disappointing results. We can now see a way of improving these responses by tailoring or augmenting the vaccine to boost development of helper T cells that will enable the body to make protective antibodies that target the malaria parasites.”