User login
Tool identifies CNAs other algorithms miss

Image courtesy of NIGMS
A new tool can detect genetic alterations that have proven difficult to identify, according to research published in Nature Methods.
The tool is an algorithm called CONSERTING (Copy Number Segmentation by Regression Tree in Next-Generation Sequencing).
Researchers created CONSERTING to improve their ability to detect copy number alterations (CNAs) in the information generated by whole-genome sequencing techniques.
The group showed that CONSERTING could detect CNAs with better accuracy and sensitivity than other techniques, including 4 published algorithms used to recognize CNAs in whole-genome sequencing data.
The data the team analyzed encompassed the normal and tumor genomes from 43 children and adults with leukemia, brain tumors, melanoma, and retinoblastoma.
“CONSERTING helped us identify alterations that other algorithms missed, including previously undetected chromosomal rearrangements and copy number alterations present in a small percentage of tumor cells,” said study author Xiang Chen, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee.
“[C]ONSERTING identified copy number alterations in children with 100 times greater precision and 10 times greater precision in adults,” added Jinghui Zhang, PhD, also of St. Jude.
Using CONSERTING, the researchers discovered genetic alterations driving pediatric leukemia, low-grade glioma, glioblastoma, and retinoblastoma.
The algorithm also helped the team identify genetic changes that are present in a small percentage of a tumor’s cells. The alterations may be the key to understanding why tumors sometimes return after treatment, they said.
Dr Zhang said CONSERTING should make it easier to track the evolution of tumors with complex genetic rearrangements.
St. Jude has made CONSERTING available to researchers free of charge. The software, user manual, and related data can be downloaded from http://www.stjuderesearch.org/site/lab/zhang.
St. Jude researchers have also developed a cloud version of CONSERTING and related tools that can be accessed through Amazon Web Services. Instead of downloading CONSERTING, scientists can upload data for analysis.
Creating CONSERTING
Work on CONSERTING began in 2010, shortly after the St. Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome Project was launched. The Pediatric Cancer Genome Project used next-generation, whole-genome sequencing to study some of the most aggressive and least understood childhood cancers.
Early on in the project, researchers realized that existing analytic methods often missed duplications or deletions of DNA segments, particularly small changes that involve a handful of genes and provide insight into the origins of a patient’s cancer.
CONSERTING has now been used to analyze data for the Pediatric Cancer Genome Project. The project includes the normal and cancer genomes of 700 pediatric cancer patients with 21 different cancer subtypes.
CONSERTING combines a method of data analysis called regression tree, which is a machine-learning algorithm, with next-generation, whole-genome sequencing. Machine learning capitalizes on advances in computing to design algorithms that repeatedly and rapidly analyze large, complex sets of data and unearth unexpected insights.
“This combination has provided us with a powerful tool for recognizing copy number alterations, even those present in relatively few cells or in tumor samples that include normal cells along with tumor cells,” Dr Zhang said.
Next-generation, whole-genome sequencing involves breaking the human genome into about 1 billion pieces that are copied and reassembled using the normal genome as a template.
CONSERTING software compensates for gaps and variations in sequencing data. The sequencing data is integrated with information about the chromosomal rearrangements to find CNAs and identify their origins in the genome. ![]()

Image courtesy of NIGMS
A new tool can detect genetic alterations that have proven difficult to identify, according to research published in Nature Methods.
The tool is an algorithm called CONSERTING (Copy Number Segmentation by Regression Tree in Next-Generation Sequencing).
Researchers created CONSERTING to improve their ability to detect copy number alterations (CNAs) in the information generated by whole-genome sequencing techniques.
The group showed that CONSERTING could detect CNAs with better accuracy and sensitivity than other techniques, including 4 published algorithms used to recognize CNAs in whole-genome sequencing data.
The data the team analyzed encompassed the normal and tumor genomes from 43 children and adults with leukemia, brain tumors, melanoma, and retinoblastoma.
“CONSERTING helped us identify alterations that other algorithms missed, including previously undetected chromosomal rearrangements and copy number alterations present in a small percentage of tumor cells,” said study author Xiang Chen, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee.
“[C]ONSERTING identified copy number alterations in children with 100 times greater precision and 10 times greater precision in adults,” added Jinghui Zhang, PhD, also of St. Jude.
Using CONSERTING, the researchers discovered genetic alterations driving pediatric leukemia, low-grade glioma, glioblastoma, and retinoblastoma.
The algorithm also helped the team identify genetic changes that are present in a small percentage of a tumor’s cells. The alterations may be the key to understanding why tumors sometimes return after treatment, they said.
Dr Zhang said CONSERTING should make it easier to track the evolution of tumors with complex genetic rearrangements.
St. Jude has made CONSERTING available to researchers free of charge. The software, user manual, and related data can be downloaded from http://www.stjuderesearch.org/site/lab/zhang.
St. Jude researchers have also developed a cloud version of CONSERTING and related tools that can be accessed through Amazon Web Services. Instead of downloading CONSERTING, scientists can upload data for analysis.
Creating CONSERTING
Work on CONSERTING began in 2010, shortly after the St. Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome Project was launched. The Pediatric Cancer Genome Project used next-generation, whole-genome sequencing to study some of the most aggressive and least understood childhood cancers.
Early on in the project, researchers realized that existing analytic methods often missed duplications or deletions of DNA segments, particularly small changes that involve a handful of genes and provide insight into the origins of a patient’s cancer.
CONSERTING has now been used to analyze data for the Pediatric Cancer Genome Project. The project includes the normal and cancer genomes of 700 pediatric cancer patients with 21 different cancer subtypes.
CONSERTING combines a method of data analysis called regression tree, which is a machine-learning algorithm, with next-generation, whole-genome sequencing. Machine learning capitalizes on advances in computing to design algorithms that repeatedly and rapidly analyze large, complex sets of data and unearth unexpected insights.
“This combination has provided us with a powerful tool for recognizing copy number alterations, even those present in relatively few cells or in tumor samples that include normal cells along with tumor cells,” Dr Zhang said.
Next-generation, whole-genome sequencing involves breaking the human genome into about 1 billion pieces that are copied and reassembled using the normal genome as a template.
CONSERTING software compensates for gaps and variations in sequencing data. The sequencing data is integrated with information about the chromosomal rearrangements to find CNAs and identify their origins in the genome. ![]()

Image courtesy of NIGMS
A new tool can detect genetic alterations that have proven difficult to identify, according to research published in Nature Methods.
The tool is an algorithm called CONSERTING (Copy Number Segmentation by Regression Tree in Next-Generation Sequencing).
Researchers created CONSERTING to improve their ability to detect copy number alterations (CNAs) in the information generated by whole-genome sequencing techniques.
The group showed that CONSERTING could detect CNAs with better accuracy and sensitivity than other techniques, including 4 published algorithms used to recognize CNAs in whole-genome sequencing data.
The data the team analyzed encompassed the normal and tumor genomes from 43 children and adults with leukemia, brain tumors, melanoma, and retinoblastoma.
“CONSERTING helped us identify alterations that other algorithms missed, including previously undetected chromosomal rearrangements and copy number alterations present in a small percentage of tumor cells,” said study author Xiang Chen, PhD, of St. Jude Children’s Research Hospital in Memphis, Tennessee.
“[C]ONSERTING identified copy number alterations in children with 100 times greater precision and 10 times greater precision in adults,” added Jinghui Zhang, PhD, also of St. Jude.
Using CONSERTING, the researchers discovered genetic alterations driving pediatric leukemia, low-grade glioma, glioblastoma, and retinoblastoma.
The algorithm also helped the team identify genetic changes that are present in a small percentage of a tumor’s cells. The alterations may be the key to understanding why tumors sometimes return after treatment, they said.
Dr Zhang said CONSERTING should make it easier to track the evolution of tumors with complex genetic rearrangements.
St. Jude has made CONSERTING available to researchers free of charge. The software, user manual, and related data can be downloaded from http://www.stjuderesearch.org/site/lab/zhang.
St. Jude researchers have also developed a cloud version of CONSERTING and related tools that can be accessed through Amazon Web Services. Instead of downloading CONSERTING, scientists can upload data for analysis.
Creating CONSERTING
Work on CONSERTING began in 2010, shortly after the St. Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome Project was launched. The Pediatric Cancer Genome Project used next-generation, whole-genome sequencing to study some of the most aggressive and least understood childhood cancers.
Early on in the project, researchers realized that existing analytic methods often missed duplications or deletions of DNA segments, particularly small changes that involve a handful of genes and provide insight into the origins of a patient’s cancer.
CONSERTING has now been used to analyze data for the Pediatric Cancer Genome Project. The project includes the normal and cancer genomes of 700 pediatric cancer patients with 21 different cancer subtypes.
CONSERTING combines a method of data analysis called regression tree, which is a machine-learning algorithm, with next-generation, whole-genome sequencing. Machine learning capitalizes on advances in computing to design algorithms that repeatedly and rapidly analyze large, complex sets of data and unearth unexpected insights.
“This combination has provided us with a powerful tool for recognizing copy number alterations, even those present in relatively few cells or in tumor samples that include normal cells along with tumor cells,” Dr Zhang said.
Next-generation, whole-genome sequencing involves breaking the human genome into about 1 billion pieces that are copied and reassembled using the normal genome as a template.
CONSERTING software compensates for gaps and variations in sequencing data. The sequencing data is integrated with information about the chromosomal rearrangements to find CNAs and identify their origins in the genome. ![]()
Simpler, more cost-effective way to grow stem cells

Photo courtesy of University
of Texas at El Paso
Researchers say they have developed a protocol to prepare human induced pluripotent stem (hiPS) cells using chemically fixed feeder cells.
This method saves time and money by avoiding the need for colony formation of live feeder cells, which is required by current conventional methods.
The new protocol challenges the theory that live feeder cells are required to provide nutrients to growing stem cells.
“We’ve proved an important phenomenon,” said Binata Joddar, PhD, of the University of Texas at El Paso. “And it suggests that these feeder cells, which are difficult to grow, may not be important at all for stem cell growth.”
Dr Joddar and her colleagues described the phenomenon in Journal of Materials Chemistry B.
Using 2.5% glutaraldehyde (GA) or formaldehyde (FA) for 10 minutes, the researchers prepared a niche matrix from autologus human dermal fibroblast (HDF) feeder cells.
They then introduced hiPS cells to the niche matrix, which adhered to and were maintained as colonies on the fixed feeder cells.
The colony doubling times of the cells grown this way were similar to those of hiPS cells grown on mitomycin-C-treated HDF or SNL feeders. (SNL cells are derived from mouse fibroblast STO cells transformed with a neomycin resistance gene.)
But the colony doubling time for the hiPS cells was shorter with the fixed feeder than for cells cultured on laminin-5.
The researchers also discovered that the average number of colonies per passage was signficiantly higher for hiPS cells cultured on fixed feeder cells compared to those cultured without feeders.
They noted hiPS cells cultured on gelatin did not grow beyond the first passage.
The team concluded that the two types of chemically fixed HDF feeder cells (HDF-glutaraldehyde and HDF-formaldehyde) can be used as substitutes for mitomycin-C-treated HDF feeders to culture hiPS cells.
This new method would not extend the doubling time, would save preparation time, and would avoid labor-intensive protocols to prepare.
In addition, after chemical fixation, the feeder cells are non-viable and cannot release active growth factors or chemokines into the cell culture. Therefore, fixed feeder cells can be refrigerated for long-term storage prior to use.
“Because feeder cells don’t need to stay alive in the process, we can store them at room temperature and spend less time cultivating them,” Dr Joddar said.
“This makes me think that we [could] use a nanomanufacturing approach to grow stem cells. We could mimic feeder cells’ nanotopology with 3-D printing techniques and skip using feeder cells altogether in the future.” ![]()

Photo courtesy of University
of Texas at El Paso
Researchers say they have developed a protocol to prepare human induced pluripotent stem (hiPS) cells using chemically fixed feeder cells.
This method saves time and money by avoiding the need for colony formation of live feeder cells, which is required by current conventional methods.
The new protocol challenges the theory that live feeder cells are required to provide nutrients to growing stem cells.
“We’ve proved an important phenomenon,” said Binata Joddar, PhD, of the University of Texas at El Paso. “And it suggests that these feeder cells, which are difficult to grow, may not be important at all for stem cell growth.”
Dr Joddar and her colleagues described the phenomenon in Journal of Materials Chemistry B.
Using 2.5% glutaraldehyde (GA) or formaldehyde (FA) for 10 minutes, the researchers prepared a niche matrix from autologus human dermal fibroblast (HDF) feeder cells.
They then introduced hiPS cells to the niche matrix, which adhered to and were maintained as colonies on the fixed feeder cells.
The colony doubling times of the cells grown this way were similar to those of hiPS cells grown on mitomycin-C-treated HDF or SNL feeders. (SNL cells are derived from mouse fibroblast STO cells transformed with a neomycin resistance gene.)
But the colony doubling time for the hiPS cells was shorter with the fixed feeder than for cells cultured on laminin-5.
The researchers also discovered that the average number of colonies per passage was signficiantly higher for hiPS cells cultured on fixed feeder cells compared to those cultured without feeders.
They noted hiPS cells cultured on gelatin did not grow beyond the first passage.
The team concluded that the two types of chemically fixed HDF feeder cells (HDF-glutaraldehyde and HDF-formaldehyde) can be used as substitutes for mitomycin-C-treated HDF feeders to culture hiPS cells.
This new method would not extend the doubling time, would save preparation time, and would avoid labor-intensive protocols to prepare.
In addition, after chemical fixation, the feeder cells are non-viable and cannot release active growth factors or chemokines into the cell culture. Therefore, fixed feeder cells can be refrigerated for long-term storage prior to use.
“Because feeder cells don’t need to stay alive in the process, we can store them at room temperature and spend less time cultivating them,” Dr Joddar said.
“This makes me think that we [could] use a nanomanufacturing approach to grow stem cells. We could mimic feeder cells’ nanotopology with 3-D printing techniques and skip using feeder cells altogether in the future.” ![]()

Photo courtesy of University
of Texas at El Paso
Researchers say they have developed a protocol to prepare human induced pluripotent stem (hiPS) cells using chemically fixed feeder cells.
This method saves time and money by avoiding the need for colony formation of live feeder cells, which is required by current conventional methods.
The new protocol challenges the theory that live feeder cells are required to provide nutrients to growing stem cells.
“We’ve proved an important phenomenon,” said Binata Joddar, PhD, of the University of Texas at El Paso. “And it suggests that these feeder cells, which are difficult to grow, may not be important at all for stem cell growth.”
Dr Joddar and her colleagues described the phenomenon in Journal of Materials Chemistry B.
Using 2.5% glutaraldehyde (GA) or formaldehyde (FA) for 10 minutes, the researchers prepared a niche matrix from autologus human dermal fibroblast (HDF) feeder cells.
They then introduced hiPS cells to the niche matrix, which adhered to and were maintained as colonies on the fixed feeder cells.
The colony doubling times of the cells grown this way were similar to those of hiPS cells grown on mitomycin-C-treated HDF or SNL feeders. (SNL cells are derived from mouse fibroblast STO cells transformed with a neomycin resistance gene.)
But the colony doubling time for the hiPS cells was shorter with the fixed feeder than for cells cultured on laminin-5.
The researchers also discovered that the average number of colonies per passage was signficiantly higher for hiPS cells cultured on fixed feeder cells compared to those cultured without feeders.
They noted hiPS cells cultured on gelatin did not grow beyond the first passage.
The team concluded that the two types of chemically fixed HDF feeder cells (HDF-glutaraldehyde and HDF-formaldehyde) can be used as substitutes for mitomycin-C-treated HDF feeders to culture hiPS cells.
This new method would not extend the doubling time, would save preparation time, and would avoid labor-intensive protocols to prepare.
In addition, after chemical fixation, the feeder cells are non-viable and cannot release active growth factors or chemokines into the cell culture. Therefore, fixed feeder cells can be refrigerated for long-term storage prior to use.
“Because feeder cells don’t need to stay alive in the process, we can store them at room temperature and spend less time cultivating them,” Dr Joddar said.
“This makes me think that we [could] use a nanomanufacturing approach to grow stem cells. We could mimic feeder cells’ nanotopology with 3-D printing techniques and skip using feeder cells altogether in the future.” ![]()
Artificial blood vessels give way to the real thing

biocompatible polymer
Photo courtesy of Vienna
University of Technology
Scientists have created implantable artificial blood vessels using a newly developed polymer that is biodegradable.
In experiments with rats, these thin-walled vascular grafts were replaced by endogenous material, ultimately leaving natural, fully functional blood vessels in their place.
Furthermore, 6 months after the artificial vessels were implanted, none of the animals had experienced thromboses, inflammation, or aneurysms.
Helga Bergmeister, MD, DVM, PhD, of the Medical University of Vienna in Austria, and her colleagues conducted this research and described the results in Acta Biomaterialia.
To create artificial blood vessels that are compatible with body tissue, the researchers developed a new polymer—thermoplastic polyurethane.
“By selecting very specific molecular building blocks, we have succeeded in synthesizing a polymer with the desired properties,” said Robert Liska, PhD, of the Vienna University of Technology.
To produce the grafts, the researchers spun polymer solutions in an electrical field to form very fine threads and wound these threads onto a spool.
“The wall of these artificial blood vessels is very similar to that of natural ones,” said Heinrich Schima, PhD, of the Medical University of Vienna.
The polymer fabric is slightly porous. So, initially, it allows a small amount of blood to seep through, which enriches the wall with growth factors. And this encourages the migration of endogenous cells.
The researchers implanted these artificial blood vessels in rats and found them to be safe and functional long-term.
“The rats’ blood vessels were examined 6 months after insertion of the vascular prostheses,” Dr Bergmeister said.
“We did not find any aneurysms, thrombosis, or inflammation. Endogenous cells had colonized the vascular prostheses and turned the artificial constructs into natural body tissue.”
In fact, natural body tissue regrew much faster than expected.
The researchers said their thin-walled grafts “offer a new and desirable form of biodegradable vascular implant.” ![]()

biocompatible polymer
Photo courtesy of Vienna
University of Technology
Scientists have created implantable artificial blood vessels using a newly developed polymer that is biodegradable.
In experiments with rats, these thin-walled vascular grafts were replaced by endogenous material, ultimately leaving natural, fully functional blood vessels in their place.
Furthermore, 6 months after the artificial vessels were implanted, none of the animals had experienced thromboses, inflammation, or aneurysms.
Helga Bergmeister, MD, DVM, PhD, of the Medical University of Vienna in Austria, and her colleagues conducted this research and described the results in Acta Biomaterialia.
To create artificial blood vessels that are compatible with body tissue, the researchers developed a new polymer—thermoplastic polyurethane.
“By selecting very specific molecular building blocks, we have succeeded in synthesizing a polymer with the desired properties,” said Robert Liska, PhD, of the Vienna University of Technology.
To produce the grafts, the researchers spun polymer solutions in an electrical field to form very fine threads and wound these threads onto a spool.
“The wall of these artificial blood vessels is very similar to that of natural ones,” said Heinrich Schima, PhD, of the Medical University of Vienna.
The polymer fabric is slightly porous. So, initially, it allows a small amount of blood to seep through, which enriches the wall with growth factors. And this encourages the migration of endogenous cells.
The researchers implanted these artificial blood vessels in rats and found them to be safe and functional long-term.
“The rats’ blood vessels were examined 6 months after insertion of the vascular prostheses,” Dr Bergmeister said.
“We did not find any aneurysms, thrombosis, or inflammation. Endogenous cells had colonized the vascular prostheses and turned the artificial constructs into natural body tissue.”
In fact, natural body tissue regrew much faster than expected.
The researchers said their thin-walled grafts “offer a new and desirable form of biodegradable vascular implant.” ![]()

biocompatible polymer
Photo courtesy of Vienna
University of Technology
Scientists have created implantable artificial blood vessels using a newly developed polymer that is biodegradable.
In experiments with rats, these thin-walled vascular grafts were replaced by endogenous material, ultimately leaving natural, fully functional blood vessels in their place.
Furthermore, 6 months after the artificial vessels were implanted, none of the animals had experienced thromboses, inflammation, or aneurysms.
Helga Bergmeister, MD, DVM, PhD, of the Medical University of Vienna in Austria, and her colleagues conducted this research and described the results in Acta Biomaterialia.
To create artificial blood vessels that are compatible with body tissue, the researchers developed a new polymer—thermoplastic polyurethane.
“By selecting very specific molecular building blocks, we have succeeded in synthesizing a polymer with the desired properties,” said Robert Liska, PhD, of the Vienna University of Technology.
To produce the grafts, the researchers spun polymer solutions in an electrical field to form very fine threads and wound these threads onto a spool.
“The wall of these artificial blood vessels is very similar to that of natural ones,” said Heinrich Schima, PhD, of the Medical University of Vienna.
The polymer fabric is slightly porous. So, initially, it allows a small amount of blood to seep through, which enriches the wall with growth factors. And this encourages the migration of endogenous cells.
The researchers implanted these artificial blood vessels in rats and found them to be safe and functional long-term.
“The rats’ blood vessels were examined 6 months after insertion of the vascular prostheses,” Dr Bergmeister said.
“We did not find any aneurysms, thrombosis, or inflammation. Endogenous cells had colonized the vascular prostheses and turned the artificial constructs into natural body tissue.”
In fact, natural body tissue regrew much faster than expected.
The researchers said their thin-walled grafts “offer a new and desirable form of biodegradable vascular implant.” ![]()
Enzyme could enable creation of universal blood type
Photo by Elise Amendola
Chemists have generated an enzyme that shows the potential for converting type A or B blood into a universal blood type.
The enzyme works by snipping off the antigens found in blood types A and B, making these blood types more like O, which can be given to patients of all blood types.
The enzyme was able to remove most of the antigens in type A and B blood. Before it can be used in clinical settings, however, all of the antigens would need to be removed.
David Kwan, PhD, of the University of British Columbia in Vancouver, Canada, and his colleagues described their work with this enzyme in the Journal of the American Chemical Society.
“We produced a mutant enzyme that is very efficient at cutting off the sugars in A and B blood and is much more proficient at removing the subtypes of the A antigen that the parent enzyme struggles with,” Dr Kwan said.
To create the enzyme, Dr Kwan and his colleagues used a technology called directed evolution. It involves inserting mutations into the gene that codes for the enzyme and selecting mutants that are more effective at cutting the antigens.
The team started with the family 98 glycoside hydrolase from Streptococcus pneumoniae SP3-BS71 (Sp3GH98), which cleaves the entire terminal trisaccharide antigenic determinants of both A and B antigens from some of the linkages on red blood cell surface glycans.
Through directed evolution, the researchers developed variants of Sp3GH98 that showed improved activity toward some of the linkages that are resistant to cleavage by the wild-type enzyme.
In 5 generations, the enzyme became 170 times more effective. This Sp3GH98 variant could remove the majority of the antigens in type A and B blood.
The researchers said the enzyme must be able to remove all of the antigens before it can be used in the clinic. The immune system is highly sensitive to blood groups, and even small amounts of residual antigens could trigger an immune response.
The concept of using an enzyme to change blood types is not new, said study author Steve Withers, PhD, also from the University of British Columbia.
“But, until now, we needed so much of the enzyme to make it work that it was impractical,” he said. “Now, I’m confident that we can take this a whole lot further.” ![]()

Photo by Elise Amendola
Chemists have generated an enzyme that shows the potential for converting type A or B blood into a universal blood type.
The enzyme works by snipping off the antigens found in blood types A and B, making these blood types more like O, which can be given to patients of all blood types.
The enzyme was able to remove most of the antigens in type A and B blood. Before it can be used in clinical settings, however, all of the antigens would need to be removed.
David Kwan, PhD, of the University of British Columbia in Vancouver, Canada, and his colleagues described their work with this enzyme in the Journal of the American Chemical Society.
“We produced a mutant enzyme that is very efficient at cutting off the sugars in A and B blood and is much more proficient at removing the subtypes of the A antigen that the parent enzyme struggles with,” Dr Kwan said.
To create the enzyme, Dr Kwan and his colleagues used a technology called directed evolution. It involves inserting mutations into the gene that codes for the enzyme and selecting mutants that are more effective at cutting the antigens.
The team started with the family 98 glycoside hydrolase from Streptococcus pneumoniae SP3-BS71 (Sp3GH98), which cleaves the entire terminal trisaccharide antigenic determinants of both A and B antigens from some of the linkages on red blood cell surface glycans.
Through directed evolution, the researchers developed variants of Sp3GH98 that showed improved activity toward some of the linkages that are resistant to cleavage by the wild-type enzyme.
In 5 generations, the enzyme became 170 times more effective. This Sp3GH98 variant could remove the majority of the antigens in type A and B blood.
The researchers said the enzyme must be able to remove all of the antigens before it can be used in the clinic. The immune system is highly sensitive to blood groups, and even small amounts of residual antigens could trigger an immune response.
The concept of using an enzyme to change blood types is not new, said study author Steve Withers, PhD, also from the University of British Columbia.
“But, until now, we needed so much of the enzyme to make it work that it was impractical,” he said. “Now, I’m confident that we can take this a whole lot further.” ![]()

Photo by Elise Amendola
Chemists have generated an enzyme that shows the potential for converting type A or B blood into a universal blood type.
The enzyme works by snipping off the antigens found in blood types A and B, making these blood types more like O, which can be given to patients of all blood types.
The enzyme was able to remove most of the antigens in type A and B blood. Before it can be used in clinical settings, however, all of the antigens would need to be removed.
David Kwan, PhD, of the University of British Columbia in Vancouver, Canada, and his colleagues described their work with this enzyme in the Journal of the American Chemical Society.
“We produced a mutant enzyme that is very efficient at cutting off the sugars in A and B blood and is much more proficient at removing the subtypes of the A antigen that the parent enzyme struggles with,” Dr Kwan said.
To create the enzyme, Dr Kwan and his colleagues used a technology called directed evolution. It involves inserting mutations into the gene that codes for the enzyme and selecting mutants that are more effective at cutting the antigens.
The team started with the family 98 glycoside hydrolase from Streptococcus pneumoniae SP3-BS71 (Sp3GH98), which cleaves the entire terminal trisaccharide antigenic determinants of both A and B antigens from some of the linkages on red blood cell surface glycans.
Through directed evolution, the researchers developed variants of Sp3GH98 that showed improved activity toward some of the linkages that are resistant to cleavage by the wild-type enzyme.
In 5 generations, the enzyme became 170 times more effective. This Sp3GH98 variant could remove the majority of the antigens in type A and B blood.
The researchers said the enzyme must be able to remove all of the antigens before it can be used in the clinic. The immune system is highly sensitive to blood groups, and even small amounts of residual antigens could trigger an immune response.
The concept of using an enzyme to change blood types is not new, said study author Steve Withers, PhD, also from the University of British Columbia.
“But, until now, we needed so much of the enzyme to make it work that it was impractical,” he said. “Now, I’m confident that we can take this a whole lot further.” ![]()
Identifying artemisinin resistance not so straightforward
Photo by Juan D. Alfonso
The current method used to identify resistance to the antimalarial drug artemisinin is not entirely accurate, according to research published in PLOS Medicine.
Artemisinin rapidly clears malaria parasites from the blood of infected patients. When parasites develop resistance, clearance takes longer.
The best measure of parasite clearance is the parasite half-life in a patient’s blood, and a common cutoff used to denote artemisinin resistance is 5 hours.
Study author Lisa White, of Mahidol University in Bangkok, Thailand, and her colleagues found that parasite half-life predicts the likelihood of an artemisinin-resistant infection for individual patients. But the half-life is influenced by how common resistance is in the particular area.
The critical half-life varied between 3.5 hours in areas where resistance is rare to 5.5 hours in areas where resistance is common. This means there is no universal cutoff value in parasite half-life that can determine whether a particular infection is “sensitive” or “resistant” to artemisinin-based combination (ACT) therapy.
Because measuring the parasite half-life requires frequent blood sampling that is difficult to do in resource-limited settings, the World Health Organization (WHO) uses the following working definition for surveillance. Artemisinin resistance in a population is suspected if more than 10% of patients are still carrying parasites 3 days after the start of ACT.
Arguing that the cutoff used in the WHO’s working definition is based on limited data, the researchers examined how well the definition matches actual data from patients in areas with artemisinin-resistant parasites.
Applying a model specifically developed for this purpose, the team found that the WHO’s day-3 cutoff value of 10% is useful, but it would be more informative if the parasite load at the start of ACT was taken into account.
The researchers concluded that the WHO definition alone cannot be used to accurately predict the real proportion of artemisinin-resistant parasites, so a more detailed assessment is needed. ![]()

Photo by Juan D. Alfonso
The current method used to identify resistance to the antimalarial drug artemisinin is not entirely accurate, according to research published in PLOS Medicine.
Artemisinin rapidly clears malaria parasites from the blood of infected patients. When parasites develop resistance, clearance takes longer.
The best measure of parasite clearance is the parasite half-life in a patient’s blood, and a common cutoff used to denote artemisinin resistance is 5 hours.
Study author Lisa White, of Mahidol University in Bangkok, Thailand, and her colleagues found that parasite half-life predicts the likelihood of an artemisinin-resistant infection for individual patients. But the half-life is influenced by how common resistance is in the particular area.
The critical half-life varied between 3.5 hours in areas where resistance is rare to 5.5 hours in areas where resistance is common. This means there is no universal cutoff value in parasite half-life that can determine whether a particular infection is “sensitive” or “resistant” to artemisinin-based combination (ACT) therapy.
Because measuring the parasite half-life requires frequent blood sampling that is difficult to do in resource-limited settings, the World Health Organization (WHO) uses the following working definition for surveillance. Artemisinin resistance in a population is suspected if more than 10% of patients are still carrying parasites 3 days after the start of ACT.
Arguing that the cutoff used in the WHO’s working definition is based on limited data, the researchers examined how well the definition matches actual data from patients in areas with artemisinin-resistant parasites.
Applying a model specifically developed for this purpose, the team found that the WHO’s day-3 cutoff value of 10% is useful, but it would be more informative if the parasite load at the start of ACT was taken into account.
The researchers concluded that the WHO definition alone cannot be used to accurately predict the real proportion of artemisinin-resistant parasites, so a more detailed assessment is needed. ![]()

Photo by Juan D. Alfonso
The current method used to identify resistance to the antimalarial drug artemisinin is not entirely accurate, according to research published in PLOS Medicine.
Artemisinin rapidly clears malaria parasites from the blood of infected patients. When parasites develop resistance, clearance takes longer.
The best measure of parasite clearance is the parasite half-life in a patient’s blood, and a common cutoff used to denote artemisinin resistance is 5 hours.
Study author Lisa White, of Mahidol University in Bangkok, Thailand, and her colleagues found that parasite half-life predicts the likelihood of an artemisinin-resistant infection for individual patients. But the half-life is influenced by how common resistance is in the particular area.
The critical half-life varied between 3.5 hours in areas where resistance is rare to 5.5 hours in areas where resistance is common. This means there is no universal cutoff value in parasite half-life that can determine whether a particular infection is “sensitive” or “resistant” to artemisinin-based combination (ACT) therapy.
Because measuring the parasite half-life requires frequent blood sampling that is difficult to do in resource-limited settings, the World Health Organization (WHO) uses the following working definition for surveillance. Artemisinin resistance in a population is suspected if more than 10% of patients are still carrying parasites 3 days after the start of ACT.
Arguing that the cutoff used in the WHO’s working definition is based on limited data, the researchers examined how well the definition matches actual data from patients in areas with artemisinin-resistant parasites.
Applying a model specifically developed for this purpose, the team found that the WHO’s day-3 cutoff value of 10% is useful, but it would be more informative if the parasite load at the start of ACT was taken into account.
The researchers concluded that the WHO definition alone cannot be used to accurately predict the real proportion of artemisinin-resistant parasites, so a more detailed assessment is needed. ![]()
Team discovers mechanism behind malaria progression
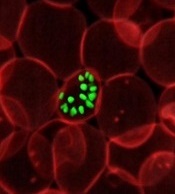
a red blood cell
Photo courtesy of St. Jude
Children’s Research Hospital
Researchers say they have pinpointed one of the mechanisms responsible for the progression of malaria.
Computer modeling showed that the nanoscale knobs that form at the membrane of infected red blood cells cause the cell stiffening that is, in part, responsible for the reduced blood flow that can turn malaria deadly.
Subra Suresh, ScD, of Carnegie Mellon University in Pittsburgh, Pennsylvania, and his colleagues reported this finding in PNAS.
“Many of malaria’s symptoms are the result of impeded blood flow, which is directly tied to structural changes in infected red blood cells,” Dr Suresh said.
When a red blood cell is infected with malaria, the parasite releases proteins that interact with the cell membrane. The cell membrane undergoes a series of changes that result in stiffness and stickiness.
While researchers are fairly certain that the stickiness is caused by nanoscale knobs that protrude from the cell membrane, they were uncertain as to what caused the stiffness.
They hypothesized that the parasite protein/cell membrane interaction caused spectrin, a cytoskeletal protein that provides a scaffold for the cell membrane, to rearrange its networked structure to be more rigid.
However, the complexity of the cell membrane made it difficult for researchers to study and prove this hypothesis experimentally.
To visualize what happens at the cell membrane during malarial infection, Dr Suresh and his colleagues turned to a computer simulation technique called coarse-grained molecular dynamics (CGMD). CGMD has proven valuable for studying what happens at the cell membrane because it represents the membrane’s complex proteins and lipids with larger, simplified components rather than atom by atom.
Doing this requires less computing time and power than standard atomistic models, which allows scientists to run simulations for longer periods of time while still accurately recreating the behavior of the cell membrane.
Typically, researchers introduce different variables into the simulation and observe how the membrane reacts. In the current study, the researchers seeded the model membrane with proteins released by the malaria parasite Plasmodium falciparum.
From their simulation, the researchers found that the stiffening of the red blood cell membrane had little to do with the remodeling of spectrin.
Instead, the nanoscale knobs that cause the red blood cells to stick to the vein’s walls also cause the membrane to stiffen through a number of different mechanisms, including composite strengthening, strain hardening, and density-dependent vertical coupling effects.
According to the researchers, the discovery of this mechanism could provide a promising target for new antimalarial therapies. ![]()

a red blood cell
Photo courtesy of St. Jude
Children’s Research Hospital
Researchers say they have pinpointed one of the mechanisms responsible for the progression of malaria.
Computer modeling showed that the nanoscale knobs that form at the membrane of infected red blood cells cause the cell stiffening that is, in part, responsible for the reduced blood flow that can turn malaria deadly.
Subra Suresh, ScD, of Carnegie Mellon University in Pittsburgh, Pennsylvania, and his colleagues reported this finding in PNAS.
“Many of malaria’s symptoms are the result of impeded blood flow, which is directly tied to structural changes in infected red blood cells,” Dr Suresh said.
When a red blood cell is infected with malaria, the parasite releases proteins that interact with the cell membrane. The cell membrane undergoes a series of changes that result in stiffness and stickiness.
While researchers are fairly certain that the stickiness is caused by nanoscale knobs that protrude from the cell membrane, they were uncertain as to what caused the stiffness.
They hypothesized that the parasite protein/cell membrane interaction caused spectrin, a cytoskeletal protein that provides a scaffold for the cell membrane, to rearrange its networked structure to be more rigid.
However, the complexity of the cell membrane made it difficult for researchers to study and prove this hypothesis experimentally.
To visualize what happens at the cell membrane during malarial infection, Dr Suresh and his colleagues turned to a computer simulation technique called coarse-grained molecular dynamics (CGMD). CGMD has proven valuable for studying what happens at the cell membrane because it represents the membrane’s complex proteins and lipids with larger, simplified components rather than atom by atom.
Doing this requires less computing time and power than standard atomistic models, which allows scientists to run simulations for longer periods of time while still accurately recreating the behavior of the cell membrane.
Typically, researchers introduce different variables into the simulation and observe how the membrane reacts. In the current study, the researchers seeded the model membrane with proteins released by the malaria parasite Plasmodium falciparum.
From their simulation, the researchers found that the stiffening of the red blood cell membrane had little to do with the remodeling of spectrin.
Instead, the nanoscale knobs that cause the red blood cells to stick to the vein’s walls also cause the membrane to stiffen through a number of different mechanisms, including composite strengthening, strain hardening, and density-dependent vertical coupling effects.
According to the researchers, the discovery of this mechanism could provide a promising target for new antimalarial therapies. ![]()

a red blood cell
Photo courtesy of St. Jude
Children’s Research Hospital
Researchers say they have pinpointed one of the mechanisms responsible for the progression of malaria.
Computer modeling showed that the nanoscale knobs that form at the membrane of infected red blood cells cause the cell stiffening that is, in part, responsible for the reduced blood flow that can turn malaria deadly.
Subra Suresh, ScD, of Carnegie Mellon University in Pittsburgh, Pennsylvania, and his colleagues reported this finding in PNAS.
“Many of malaria’s symptoms are the result of impeded blood flow, which is directly tied to structural changes in infected red blood cells,” Dr Suresh said.
When a red blood cell is infected with malaria, the parasite releases proteins that interact with the cell membrane. The cell membrane undergoes a series of changes that result in stiffness and stickiness.
While researchers are fairly certain that the stickiness is caused by nanoscale knobs that protrude from the cell membrane, they were uncertain as to what caused the stiffness.
They hypothesized that the parasite protein/cell membrane interaction caused spectrin, a cytoskeletal protein that provides a scaffold for the cell membrane, to rearrange its networked structure to be more rigid.
However, the complexity of the cell membrane made it difficult for researchers to study and prove this hypothesis experimentally.
To visualize what happens at the cell membrane during malarial infection, Dr Suresh and his colleagues turned to a computer simulation technique called coarse-grained molecular dynamics (CGMD). CGMD has proven valuable for studying what happens at the cell membrane because it represents the membrane’s complex proteins and lipids with larger, simplified components rather than atom by atom.
Doing this requires less computing time and power than standard atomistic models, which allows scientists to run simulations for longer periods of time while still accurately recreating the behavior of the cell membrane.
Typically, researchers introduce different variables into the simulation and observe how the membrane reacts. In the current study, the researchers seeded the model membrane with proteins released by the malaria parasite Plasmodium falciparum.
From their simulation, the researchers found that the stiffening of the red blood cell membrane had little to do with the remodeling of spectrin.
Instead, the nanoscale knobs that cause the red blood cells to stick to the vein’s walls also cause the membrane to stiffen through a number of different mechanisms, including composite strengthening, strain hardening, and density-dependent vertical coupling effects.
According to the researchers, the discovery of this mechanism could provide a promising target for new antimalarial therapies. ![]()
Malaria vaccine candidate proves somewhat effective
Photo by Caitlin Kleiboer
The malaria vaccine candidate RTS,S/AS01 is somewhat effective in young African children for up to 4 years after vaccination, according to final data from a phase 3 trial.
The vaccine proved more effective against clinical and severe malaria in children than in young infants, but efficacy waned over time in both age groups.
On the other hand, a booster dose of RTS,S/AS01 increased the average number of malaria cases prevented in children and infants.
“Despite the falling efficacy over time, there is still a clear benefit from RTS,S/AS01,” said Brian Greenwood, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“An average 1363 cases of clinical malaria were prevented over 4 years of follow-up for every 1000 children vaccinated, and 1774 cases in those who also received a booster shot. Over 3 years of follow-up, an average 558 cases were averted for every 1000 infants vaccinated, and 983 cases in those also given a booster dose.”
Dr Greenwood and his colleagues disclosed these data in The Lancet. The research was funded by GlaxoSmithKline Biologicals SA, the company developing RTS,S/AS01, and the PATH Malaria Vaccine Initiative.
The trial included 15,459 young infants (aged 6 weeks to 12 weeks at first vaccination) and children (5 months to 17 months at first vaccination) from 11 sites across 7 sub-Saharan African countries (Burkina Faso, Gabon, Ghana, Kenya, Malawi, Mozambique, and United Republic of Tanzania) with varying levels of malaria transmission.
Earlier results from this trial, at 18 months of follow-up, showed efficacy of about 46% against clinical malaria in children and around 27% among young infants. Vaccine efficacy is defined as the reduction in the incidence of disease among participants who receive the vaccine compared to the incidence among participants who do not.
Dr Greenwood and his colleagues followed the infants and children for a further 20 to 30 months, respectively, and assessed the impact of a fourth booster dose.
Participants were each vaccinated 3 times with RTS,S/AS01, with or without a booster dose 18 months later, or given 4 doses of a comparator vaccine (control group).
In children who received 3 doses of RTS,S/AS01 plus a booster, the number of clinical episodes of malaria at 4 years was reduced by just over a third (36%). This is a drop in efficacy from the 50% protection against malaria seen in the first year.
Without a booster dose, the vaccine was not significantly effective against severe malaria in this age group. However, in children given a booster dose, the overall protective efficacy against severe malaria was 32% and 35% against malaria-associated hospitalizations.
In infants who received 3 doses of RTS,S/AS01 plus a booster, the vaccine reduced the risk of clinical episodes of malaria by 26% over 3 years of follow-up. There was no significant protection against severe disease in infants.
Meningitis occurred more frequently in children given RTS,S/AS01 than in children given the control vaccine. There were 11 cases of meningitis among children who received a booster, 10 cases among children who did not receive a booster, and 1 case among children in the control group.
RTS,S/AS02 produced more adverse reactions than the control vaccines. Convulsions following vaccination, although uncommon, occurred more frequently in children who received RTS,S/AS01. The incidence of other serious adverse events was similar in all the groups.
“The European Medicines Agency (EMA) will assess the quality, safety, and efficacy of the vaccine based on these final data,” Dr Greenwood said. “If the EMA gives a favorable opinion, WHO could recommend the use of RTS,S/AS01 as early as October this year. If licensed, RTS,S/AS01 would be the first licensed human vaccine against a parasitic disease.” ![]()

Photo by Caitlin Kleiboer
The malaria vaccine candidate RTS,S/AS01 is somewhat effective in young African children for up to 4 years after vaccination, according to final data from a phase 3 trial.
The vaccine proved more effective against clinical and severe malaria in children than in young infants, but efficacy waned over time in both age groups.
On the other hand, a booster dose of RTS,S/AS01 increased the average number of malaria cases prevented in children and infants.
“Despite the falling efficacy over time, there is still a clear benefit from RTS,S/AS01,” said Brian Greenwood, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“An average 1363 cases of clinical malaria were prevented over 4 years of follow-up for every 1000 children vaccinated, and 1774 cases in those who also received a booster shot. Over 3 years of follow-up, an average 558 cases were averted for every 1000 infants vaccinated, and 983 cases in those also given a booster dose.”
Dr Greenwood and his colleagues disclosed these data in The Lancet. The research was funded by GlaxoSmithKline Biologicals SA, the company developing RTS,S/AS01, and the PATH Malaria Vaccine Initiative.
The trial included 15,459 young infants (aged 6 weeks to 12 weeks at first vaccination) and children (5 months to 17 months at first vaccination) from 11 sites across 7 sub-Saharan African countries (Burkina Faso, Gabon, Ghana, Kenya, Malawi, Mozambique, and United Republic of Tanzania) with varying levels of malaria transmission.
Earlier results from this trial, at 18 months of follow-up, showed efficacy of about 46% against clinical malaria in children and around 27% among young infants. Vaccine efficacy is defined as the reduction in the incidence of disease among participants who receive the vaccine compared to the incidence among participants who do not.
Dr Greenwood and his colleagues followed the infants and children for a further 20 to 30 months, respectively, and assessed the impact of a fourth booster dose.
Participants were each vaccinated 3 times with RTS,S/AS01, with or without a booster dose 18 months later, or given 4 doses of a comparator vaccine (control group).
In children who received 3 doses of RTS,S/AS01 plus a booster, the number of clinical episodes of malaria at 4 years was reduced by just over a third (36%). This is a drop in efficacy from the 50% protection against malaria seen in the first year.
Without a booster dose, the vaccine was not significantly effective against severe malaria in this age group. However, in children given a booster dose, the overall protective efficacy against severe malaria was 32% and 35% against malaria-associated hospitalizations.
In infants who received 3 doses of RTS,S/AS01 plus a booster, the vaccine reduced the risk of clinical episodes of malaria by 26% over 3 years of follow-up. There was no significant protection against severe disease in infants.
Meningitis occurred more frequently in children given RTS,S/AS01 than in children given the control vaccine. There were 11 cases of meningitis among children who received a booster, 10 cases among children who did not receive a booster, and 1 case among children in the control group.
RTS,S/AS02 produced more adverse reactions than the control vaccines. Convulsions following vaccination, although uncommon, occurred more frequently in children who received RTS,S/AS01. The incidence of other serious adverse events was similar in all the groups.
“The European Medicines Agency (EMA) will assess the quality, safety, and efficacy of the vaccine based on these final data,” Dr Greenwood said. “If the EMA gives a favorable opinion, WHO could recommend the use of RTS,S/AS01 as early as October this year. If licensed, RTS,S/AS01 would be the first licensed human vaccine against a parasitic disease.” ![]()

Photo by Caitlin Kleiboer
The malaria vaccine candidate RTS,S/AS01 is somewhat effective in young African children for up to 4 years after vaccination, according to final data from a phase 3 trial.
The vaccine proved more effective against clinical and severe malaria in children than in young infants, but efficacy waned over time in both age groups.
On the other hand, a booster dose of RTS,S/AS01 increased the average number of malaria cases prevented in children and infants.
“Despite the falling efficacy over time, there is still a clear benefit from RTS,S/AS01,” said Brian Greenwood, MD, of the London School of Hygiene & Tropical Medicine in the UK.
“An average 1363 cases of clinical malaria were prevented over 4 years of follow-up for every 1000 children vaccinated, and 1774 cases in those who also received a booster shot. Over 3 years of follow-up, an average 558 cases were averted for every 1000 infants vaccinated, and 983 cases in those also given a booster dose.”
Dr Greenwood and his colleagues disclosed these data in The Lancet. The research was funded by GlaxoSmithKline Biologicals SA, the company developing RTS,S/AS01, and the PATH Malaria Vaccine Initiative.
The trial included 15,459 young infants (aged 6 weeks to 12 weeks at first vaccination) and children (5 months to 17 months at first vaccination) from 11 sites across 7 sub-Saharan African countries (Burkina Faso, Gabon, Ghana, Kenya, Malawi, Mozambique, and United Republic of Tanzania) with varying levels of malaria transmission.
Earlier results from this trial, at 18 months of follow-up, showed efficacy of about 46% against clinical malaria in children and around 27% among young infants. Vaccine efficacy is defined as the reduction in the incidence of disease among participants who receive the vaccine compared to the incidence among participants who do not.
Dr Greenwood and his colleagues followed the infants and children for a further 20 to 30 months, respectively, and assessed the impact of a fourth booster dose.
Participants were each vaccinated 3 times with RTS,S/AS01, with or without a booster dose 18 months later, or given 4 doses of a comparator vaccine (control group).
In children who received 3 doses of RTS,S/AS01 plus a booster, the number of clinical episodes of malaria at 4 years was reduced by just over a third (36%). This is a drop in efficacy from the 50% protection against malaria seen in the first year.
Without a booster dose, the vaccine was not significantly effective against severe malaria in this age group. However, in children given a booster dose, the overall protective efficacy against severe malaria was 32% and 35% against malaria-associated hospitalizations.
In infants who received 3 doses of RTS,S/AS01 plus a booster, the vaccine reduced the risk of clinical episodes of malaria by 26% over 3 years of follow-up. There was no significant protection against severe disease in infants.
Meningitis occurred more frequently in children given RTS,S/AS01 than in children given the control vaccine. There were 11 cases of meningitis among children who received a booster, 10 cases among children who did not receive a booster, and 1 case among children in the control group.
RTS,S/AS02 produced more adverse reactions than the control vaccines. Convulsions following vaccination, although uncommon, occurred more frequently in children who received RTS,S/AS01. The incidence of other serious adverse events was similar in all the groups.
“The European Medicines Agency (EMA) will assess the quality, safety, and efficacy of the vaccine based on these final data,” Dr Greenwood said. “If the EMA gives a favorable opinion, WHO could recommend the use of RTS,S/AS01 as early as October this year. If licensed, RTS,S/AS01 would be the first licensed human vaccine against a parasitic disease.”
CCSs more likely to claim social security support

Photo courtesy of
Huntsman Cancer Institute
A new study indicates that childhood cancer survivors (CCSs) are more likely than individuals without a cancer history to enroll on federal programs that provide disability benefits.
CCSs diagnosed between 1970 and 1986 were about 2 to 5 times as likely as control subjects to utilize such a program.
“The long-term impact of cancer can affect other issues besides health outcomes,” said study author Anne Kirchhoff, PhD, of the Huntsman Cancer Institute at the University of Utah.
“We need to do a better job of helping people function throughout their lives, not just when they’re finishing their cancer therapy.”
Dr Kirchhoff and her colleagues conducted this research and detailed the results in the Journal of the National Cancer Institute.
The researchers looked at health insurance surveys completed in 2011 and 2012 by a random sample of 698 CCSs who were diagnosed between the ages of 0 and 20 years. Today, they range in age from 20s to early 60s.
The patients are part of a National Cancer Institute initiative, called the Childhood Cancer Survivor Study, which has followed more than 14,000 children and adolescents since 1994 who were diagnosed with cancer and survived at least 5 years after diagnosis. A comparison group of 210 siblings without cancer also responded to the survey and were used as controls.
Dr Kirchhoff and her colleagues looked at current or former enrollment on 2 federal disability programs:
- Supplemental security income (SSI), which is for people with limited income who have no prior work history
- Social security disability insurance (SSDI), which pays disability benefits to adults ages 18 years and older who have worked and paid social security taxes.
In all, 13.5% of CCSs reported being enrolled on SSI in the past or present, and 10% of survivors reported being enrolled on SSDI at some point. This was substantially higher than for the comparison group, in which 2.6% of patients reported SSI enrollment and 5.4% reported SSDI enrollment.
In addition, CCSs reported current enrollment in SSI more frequently than the US population, at rates of 7.3% and 2.5%, respectively.
Dr Kirchoff and her colleagues also identified survivor socio-demographic and treatment characteristics that were associated with a higher rate of enrollment in federal support programs.
“Survivors that were younger at diagnosis, age 4 or under, were about 7 times more likely to be on SSI than we see with survivors that were diagnosed in their adolescence,” she said.
SSI enrollment was more likely for female CCSs and for survivors with a history of cranial radiation treatment as well.
Dr Kirchhoff noted that, over the years, research on CCSs has caused hospitals to rethink how to better care for cancer survivors.
“There’s really a growing strategy to support survivors in the long-term,” she said. “For example, here at Huntsman Cancer Institute, we have a pediatric cancer late-effects clinic, which helps manage issues that might come up with childhood cancer survivors in the long term, including health-management support, health-behavior support, and access to providers to help them with other issues.”

Photo courtesy of
Huntsman Cancer Institute
A new study indicates that childhood cancer survivors (CCSs) are more likely than individuals without a cancer history to enroll on federal programs that provide disability benefits.
CCSs diagnosed between 1970 and 1986 were about 2 to 5 times as likely as control subjects to utilize such a program.
“The long-term impact of cancer can affect other issues besides health outcomes,” said study author Anne Kirchhoff, PhD, of the Huntsman Cancer Institute at the University of Utah.
“We need to do a better job of helping people function throughout their lives, not just when they’re finishing their cancer therapy.”
Dr Kirchhoff and her colleagues conducted this research and detailed the results in the Journal of the National Cancer Institute.
The researchers looked at health insurance surveys completed in 2011 and 2012 by a random sample of 698 CCSs who were diagnosed between the ages of 0 and 20 years. Today, they range in age from 20s to early 60s.
The patients are part of a National Cancer Institute initiative, called the Childhood Cancer Survivor Study, which has followed more than 14,000 children and adolescents since 1994 who were diagnosed with cancer and survived at least 5 years after diagnosis. A comparison group of 210 siblings without cancer also responded to the survey and were used as controls.
Dr Kirchhoff and her colleagues looked at current or former enrollment on 2 federal disability programs:
- Supplemental security income (SSI), which is for people with limited income who have no prior work history
- Social security disability insurance (SSDI), which pays disability benefits to adults ages 18 years and older who have worked and paid social security taxes.
In all, 13.5% of CCSs reported being enrolled on SSI in the past or present, and 10% of survivors reported being enrolled on SSDI at some point. This was substantially higher than for the comparison group, in which 2.6% of patients reported SSI enrollment and 5.4% reported SSDI enrollment.
In addition, CCSs reported current enrollment in SSI more frequently than the US population, at rates of 7.3% and 2.5%, respectively.
Dr Kirchoff and her colleagues also identified survivor socio-demographic and treatment characteristics that were associated with a higher rate of enrollment in federal support programs.
“Survivors that were younger at diagnosis, age 4 or under, were about 7 times more likely to be on SSI than we see with survivors that were diagnosed in their adolescence,” she said.
SSI enrollment was more likely for female CCSs and for survivors with a history of cranial radiation treatment as well.
Dr Kirchhoff noted that, over the years, research on CCSs has caused hospitals to rethink how to better care for cancer survivors.
“There’s really a growing strategy to support survivors in the long-term,” she said. “For example, here at Huntsman Cancer Institute, we have a pediatric cancer late-effects clinic, which helps manage issues that might come up with childhood cancer survivors in the long term, including health-management support, health-behavior support, and access to providers to help them with other issues.”

Photo courtesy of
Huntsman Cancer Institute
A new study indicates that childhood cancer survivors (CCSs) are more likely than individuals without a cancer history to enroll on federal programs that provide disability benefits.
CCSs diagnosed between 1970 and 1986 were about 2 to 5 times as likely as control subjects to utilize such a program.
“The long-term impact of cancer can affect other issues besides health outcomes,” said study author Anne Kirchhoff, PhD, of the Huntsman Cancer Institute at the University of Utah.
“We need to do a better job of helping people function throughout their lives, not just when they’re finishing their cancer therapy.”
Dr Kirchhoff and her colleagues conducted this research and detailed the results in the Journal of the National Cancer Institute.
The researchers looked at health insurance surveys completed in 2011 and 2012 by a random sample of 698 CCSs who were diagnosed between the ages of 0 and 20 years. Today, they range in age from 20s to early 60s.
The patients are part of a National Cancer Institute initiative, called the Childhood Cancer Survivor Study, which has followed more than 14,000 children and adolescents since 1994 who were diagnosed with cancer and survived at least 5 years after diagnosis. A comparison group of 210 siblings without cancer also responded to the survey and were used as controls.
Dr Kirchhoff and her colleagues looked at current or former enrollment on 2 federal disability programs:
- Supplemental security income (SSI), which is for people with limited income who have no prior work history
- Social security disability insurance (SSDI), which pays disability benefits to adults ages 18 years and older who have worked and paid social security taxes.
In all, 13.5% of CCSs reported being enrolled on SSI in the past or present, and 10% of survivors reported being enrolled on SSDI at some point. This was substantially higher than for the comparison group, in which 2.6% of patients reported SSI enrollment and 5.4% reported SSDI enrollment.
In addition, CCSs reported current enrollment in SSI more frequently than the US population, at rates of 7.3% and 2.5%, respectively.
Dr Kirchoff and her colleagues also identified survivor socio-demographic and treatment characteristics that were associated with a higher rate of enrollment in federal support programs.
“Survivors that were younger at diagnosis, age 4 or under, were about 7 times more likely to be on SSI than we see with survivors that were diagnosed in their adolescence,” she said.
SSI enrollment was more likely for female CCSs and for survivors with a history of cranial radiation treatment as well.
Dr Kirchhoff noted that, over the years, research on CCSs has caused hospitals to rethink how to better care for cancer survivors.
“There’s really a growing strategy to support survivors in the long-term,” she said. “For example, here at Huntsman Cancer Institute, we have a pediatric cancer late-effects clinic, which helps manage issues that might come up with childhood cancer survivors in the long term, including health-management support, health-behavior support, and access to providers to help them with other issues.”
Team images immune response

Photo by Aaron Logan
Researchers say they have devised an approach that allows for real-time imaging of the immune system’s response to tumors, without the need for blood draws or biopsies.
The method harnesses PET to identify areas of immune cell activity associated with inflammation or tumor development.
The researchers believe the approach offers a potential breakthrough in diagnostics and the ability to monitor the efficacy of cancer therapies.
The team described their method in PNAS.
Study author Hidde Ploegh, PhD, of the Whitehead Institute for Biomedical Research in Cambridge, Massachusetts, said that every experimental immunologist wants to monitor an ongoing immune response, but options are limited.
“One can look at blood, but blood is a vehicle of transport for immune cells and is not where immune responses occur,” he said. “Surgical biopsies are invasive and non-random, so, for example, a fine-needle aspirate of a tumor could miss a significant feature of that condition.”
In search of a better monitoring approach, Dr Ploegh and his colleagues leveraged two research tools that have become staples in his lab in recent years.
The first exploits single-domain antibodies known as VHHs, derived from the heavy chain-only antibodies made by the immune systems of animals in the camelid family. Dr Ploegh’s lab immunizes alpacas—his camelid of choice—to generate VHHs specific to immune cells of interest.
The second tool, sortagging, labels the VHHs in a site-specific fashion so that researchers can track the VHHs and their targets in a living animal.
Knowing that the tissue around tumors often contains immune cells such as neutrophils and macrophages, Dr Ploegh and his colleagues hypothesized that appropriately labeled VHHs might allow them to pinpoint tumor locations by finding the tumor-associated immune cells.
Dr Ploegh noted that VHHs’ extremely small size—approximately one-tenth that of conventional antibodies—are likely responsible for their superior tissue penetration and, thus, makes them particularly well suited for such use.
So the researchers generated VHHs that recognize mouse immune cells, then labeled these VHHs with radioisotopes and injected them into tumor-bearing mice. Subsequent PET imaging detected the location of immune cells around the tumor quickly and accurately.
“We were able to image tumors as small as 1 mm in size and within just a few days of their starting to grow,” said Mohammad Rashidian, PhD, a researcher in Dr Ploegh’s lab.
“We’re very excited about this because it’s a powerful approach to pick up inflammation in and around the tumor.”
Drs Rashidian and Ploegh believe that, with further refinement, the method could be used to monitor response to—and perhaps modify—cancer immunotherapy.
“To succeed with immunotherapy, we need more information about the tumor microenvironment,” Dr Rashidian said. “With this method, you could perhaps start immunotherapy and then, a few weeks later, image with VHHs to figure out progress and success of treatment.”
“PET imaging should allow a much more comprehensive look at the entire tumor in its environment,” Dr Ploegh added. “Then we can ask, ‘Did the tumor grow? Did immune cells invade? What has happened to the tumor?’ And to be able to see this without going in invasively is a significant achievement.”

Photo by Aaron Logan
Researchers say they have devised an approach that allows for real-time imaging of the immune system’s response to tumors, without the need for blood draws or biopsies.
The method harnesses PET to identify areas of immune cell activity associated with inflammation or tumor development.
The researchers believe the approach offers a potential breakthrough in diagnostics and the ability to monitor the efficacy of cancer therapies.
The team described their method in PNAS.
Study author Hidde Ploegh, PhD, of the Whitehead Institute for Biomedical Research in Cambridge, Massachusetts, said that every experimental immunologist wants to monitor an ongoing immune response, but options are limited.
“One can look at blood, but blood is a vehicle of transport for immune cells and is not where immune responses occur,” he said. “Surgical biopsies are invasive and non-random, so, for example, a fine-needle aspirate of a tumor could miss a significant feature of that condition.”
In search of a better monitoring approach, Dr Ploegh and his colleagues leveraged two research tools that have become staples in his lab in recent years.
The first exploits single-domain antibodies known as VHHs, derived from the heavy chain-only antibodies made by the immune systems of animals in the camelid family. Dr Ploegh’s lab immunizes alpacas—his camelid of choice—to generate VHHs specific to immune cells of interest.
The second tool, sortagging, labels the VHHs in a site-specific fashion so that researchers can track the VHHs and their targets in a living animal.
Knowing that the tissue around tumors often contains immune cells such as neutrophils and macrophages, Dr Ploegh and his colleagues hypothesized that appropriately labeled VHHs might allow them to pinpoint tumor locations by finding the tumor-associated immune cells.
Dr Ploegh noted that VHHs’ extremely small size—approximately one-tenth that of conventional antibodies—are likely responsible for their superior tissue penetration and, thus, makes them particularly well suited for such use.
So the researchers generated VHHs that recognize mouse immune cells, then labeled these VHHs with radioisotopes and injected them into tumor-bearing mice. Subsequent PET imaging detected the location of immune cells around the tumor quickly and accurately.
“We were able to image tumors as small as 1 mm in size and within just a few days of their starting to grow,” said Mohammad Rashidian, PhD, a researcher in Dr Ploegh’s lab.
“We’re very excited about this because it’s a powerful approach to pick up inflammation in and around the tumor.”
Drs Rashidian and Ploegh believe that, with further refinement, the method could be used to monitor response to—and perhaps modify—cancer immunotherapy.
“To succeed with immunotherapy, we need more information about the tumor microenvironment,” Dr Rashidian said. “With this method, you could perhaps start immunotherapy and then, a few weeks later, image with VHHs to figure out progress and success of treatment.”
“PET imaging should allow a much more comprehensive look at the entire tumor in its environment,” Dr Ploegh added. “Then we can ask, ‘Did the tumor grow? Did immune cells invade? What has happened to the tumor?’ And to be able to see this without going in invasively is a significant achievement.”

Photo by Aaron Logan
Researchers say they have devised an approach that allows for real-time imaging of the immune system’s response to tumors, without the need for blood draws or biopsies.
The method harnesses PET to identify areas of immune cell activity associated with inflammation or tumor development.
The researchers believe the approach offers a potential breakthrough in diagnostics and the ability to monitor the efficacy of cancer therapies.
The team described their method in PNAS.
Study author Hidde Ploegh, PhD, of the Whitehead Institute for Biomedical Research in Cambridge, Massachusetts, said that every experimental immunologist wants to monitor an ongoing immune response, but options are limited.
“One can look at blood, but blood is a vehicle of transport for immune cells and is not where immune responses occur,” he said. “Surgical biopsies are invasive and non-random, so, for example, a fine-needle aspirate of a tumor could miss a significant feature of that condition.”
In search of a better monitoring approach, Dr Ploegh and his colleagues leveraged two research tools that have become staples in his lab in recent years.
The first exploits single-domain antibodies known as VHHs, derived from the heavy chain-only antibodies made by the immune systems of animals in the camelid family. Dr Ploegh’s lab immunizes alpacas—his camelid of choice—to generate VHHs specific to immune cells of interest.
The second tool, sortagging, labels the VHHs in a site-specific fashion so that researchers can track the VHHs and their targets in a living animal.
Knowing that the tissue around tumors often contains immune cells such as neutrophils and macrophages, Dr Ploegh and his colleagues hypothesized that appropriately labeled VHHs might allow them to pinpoint tumor locations by finding the tumor-associated immune cells.
Dr Ploegh noted that VHHs’ extremely small size—approximately one-tenth that of conventional antibodies—are likely responsible for their superior tissue penetration and, thus, makes them particularly well suited for such use.
So the researchers generated VHHs that recognize mouse immune cells, then labeled these VHHs with radioisotopes and injected them into tumor-bearing mice. Subsequent PET imaging detected the location of immune cells around the tumor quickly and accurately.
“We were able to image tumors as small as 1 mm in size and within just a few days of their starting to grow,” said Mohammad Rashidian, PhD, a researcher in Dr Ploegh’s lab.
“We’re very excited about this because it’s a powerful approach to pick up inflammation in and around the tumor.”
Drs Rashidian and Ploegh believe that, with further refinement, the method could be used to monitor response to—and perhaps modify—cancer immunotherapy.
“To succeed with immunotherapy, we need more information about the tumor microenvironment,” Dr Rashidian said. “With this method, you could perhaps start immunotherapy and then, a few weeks later, image with VHHs to figure out progress and success of treatment.”
“PET imaging should allow a much more comprehensive look at the entire tumor in its environment,” Dr Ploegh added. “Then we can ask, ‘Did the tumor grow? Did immune cells invade? What has happened to the tumor?’ And to be able to see this without going in invasively is a significant achievement.”
Decreased RBC clearance predicts ill health
New research suggests that increased red blood cell distribution width (RDW) is caused by reduced clearance of aging red blood cells (RBCs) from the bloodstream.
And previous studies showed that elevations in RDW predict the development, progression, and risk of death from many conditions.
“It appears that the human body slightly slows down the production and destruction of red blood cells in just about every major disease,” said John Higgins, MD, of Massachusetts General Hospital in Boston.
“If we can accurately measure the production or destruction rates, we might be able to identify many of these diseases in their earlier stages when they are most treatable. Existing measures of the production rate are far too imprecise to detect these subtle changes, but this paper shows how the destruction rate can be estimated using existing blood count data and a mathematical model.”
The paper has been published in the American Journal of Hematology.
Healthy adults generate RBCs at a rate of more than 2 million per second, and the cells circulate in the bloodstream for 100 to 120 days, during which their size decreases by around 30%. Aged RBCs are then cleared at about the same rate of 2 million per second.
Prior to the reports linking elevated RDW with an increased risk for many diseases, that measure had only been used to help distinguish between different forms of anemia.
To understand the correlation between RDW and disease prognosis, Dr Higgins and his colleagues analyzed raw data from more than 60,000 randomly selected blood samples. They used a mathematical model to replicate how RBC populations behave differently in health and in disease.
The researchers measured the extent to which RBCs in different phases of their life cycle contribute to increased RDW and found that the variance in size was strongly determined by mild increases in the numbers of the smallest and oldest cells.
Since the most important mechanism controlling the number of the oldest cells is the rate at which they are cleared from the bloodstream, the team made several predictions, which were validated by applying their model to clinical data from the blood samples and to data from several published studies.
The researchers found that increased RDW was associated with delayed RBC clearance, and increased RDW was associated with increased average age of RBCs.
Delayed RBC clearance was as strongly associated with overall risk of death, as was increased RDW, and delayed RBC clearance was associated with the presence of early signs of hidden diseases associated with increased RDW.
Patients with delayed RBC clearance had a greater risk of developing signs of those diseases in the future than patients with a typical clearance rate. In healthy patients, the rate of RBC clearance varied less than any other traditional blood-count characteristic.
Dr Higgins said there are many potential clinical applications of these findings, which need to be validated by future studies.
“Finding a reduced clearance rate in an apparently healthy person would likely mean that an underlying disease process was developing—such as the early stages of iron deficiency, kidney disease, colon cancer, or congestive heart failure—and would warrant further diagnostic evaluation,” Dr Higgins said.
“Based on this analysis of routine blood tests, a primary care physician could immediately consider appropriate follow-up diagnostic testing, instead of waiting for other signs and symptoms to appear as the condition progresses. In a patient with established disease, a reduced clearance rate could mean progression of disease or treatment failure, and imminent complications could be avoided or reduced by adjusting treatment right away or at least by more frequent monitoring.”
“In addition to confirming our findings in prospective studies that would follow a group of patients over time, we hope to identify the diseases for which an early warning provided by delayed clearance would lead to the most significant improvements in patient outcomes. We’d also like to understand more about the processes controlling red blood cell clearance and are actively developing similar models for populations of white blood cells and platelets.”

New research suggests that increased red blood cell distribution width (RDW) is caused by reduced clearance of aging red blood cells (RBCs) from the bloodstream.
And previous studies showed that elevations in RDW predict the development, progression, and risk of death from many conditions.
“It appears that the human body slightly slows down the production and destruction of red blood cells in just about every major disease,” said John Higgins, MD, of Massachusetts General Hospital in Boston.
“If we can accurately measure the production or destruction rates, we might be able to identify many of these diseases in their earlier stages when they are most treatable. Existing measures of the production rate are far too imprecise to detect these subtle changes, but this paper shows how the destruction rate can be estimated using existing blood count data and a mathematical model.”
The paper has been published in the American Journal of Hematology.
Healthy adults generate RBCs at a rate of more than 2 million per second, and the cells circulate in the bloodstream for 100 to 120 days, during which their size decreases by around 30%. Aged RBCs are then cleared at about the same rate of 2 million per second.
Prior to the reports linking elevated RDW with an increased risk for many diseases, that measure had only been used to help distinguish between different forms of anemia.
To understand the correlation between RDW and disease prognosis, Dr Higgins and his colleagues analyzed raw data from more than 60,000 randomly selected blood samples. They used a mathematical model to replicate how RBC populations behave differently in health and in disease.
The researchers measured the extent to which RBCs in different phases of their life cycle contribute to increased RDW and found that the variance in size was strongly determined by mild increases in the numbers of the smallest and oldest cells.
Since the most important mechanism controlling the number of the oldest cells is the rate at which they are cleared from the bloodstream, the team made several predictions, which were validated by applying their model to clinical data from the blood samples and to data from several published studies.
The researchers found that increased RDW was associated with delayed RBC clearance, and increased RDW was associated with increased average age of RBCs.
Delayed RBC clearance was as strongly associated with overall risk of death, as was increased RDW, and delayed RBC clearance was associated with the presence of early signs of hidden diseases associated with increased RDW.
Patients with delayed RBC clearance had a greater risk of developing signs of those diseases in the future than patients with a typical clearance rate. In healthy patients, the rate of RBC clearance varied less than any other traditional blood-count characteristic.
Dr Higgins said there are many potential clinical applications of these findings, which need to be validated by future studies.
“Finding a reduced clearance rate in an apparently healthy person would likely mean that an underlying disease process was developing—such as the early stages of iron deficiency, kidney disease, colon cancer, or congestive heart failure—and would warrant further diagnostic evaluation,” Dr Higgins said.
“Based on this analysis of routine blood tests, a primary care physician could immediately consider appropriate follow-up diagnostic testing, instead of waiting for other signs and symptoms to appear as the condition progresses. In a patient with established disease, a reduced clearance rate could mean progression of disease or treatment failure, and imminent complications could be avoided or reduced by adjusting treatment right away or at least by more frequent monitoring.”
“In addition to confirming our findings in prospective studies that would follow a group of patients over time, we hope to identify the diseases for which an early warning provided by delayed clearance would lead to the most significant improvements in patient outcomes. We’d also like to understand more about the processes controlling red blood cell clearance and are actively developing similar models for populations of white blood cells and platelets.”

New research suggests that increased red blood cell distribution width (RDW) is caused by reduced clearance of aging red blood cells (RBCs) from the bloodstream.
And previous studies showed that elevations in RDW predict the development, progression, and risk of death from many conditions.
“It appears that the human body slightly slows down the production and destruction of red blood cells in just about every major disease,” said John Higgins, MD, of Massachusetts General Hospital in Boston.
“If we can accurately measure the production or destruction rates, we might be able to identify many of these diseases in their earlier stages when they are most treatable. Existing measures of the production rate are far too imprecise to detect these subtle changes, but this paper shows how the destruction rate can be estimated using existing blood count data and a mathematical model.”
The paper has been published in the American Journal of Hematology.
Healthy adults generate RBCs at a rate of more than 2 million per second, and the cells circulate in the bloodstream for 100 to 120 days, during which their size decreases by around 30%. Aged RBCs are then cleared at about the same rate of 2 million per second.
Prior to the reports linking elevated RDW with an increased risk for many diseases, that measure had only been used to help distinguish between different forms of anemia.
To understand the correlation between RDW and disease prognosis, Dr Higgins and his colleagues analyzed raw data from more than 60,000 randomly selected blood samples. They used a mathematical model to replicate how RBC populations behave differently in health and in disease.
The researchers measured the extent to which RBCs in different phases of their life cycle contribute to increased RDW and found that the variance in size was strongly determined by mild increases in the numbers of the smallest and oldest cells.
Since the most important mechanism controlling the number of the oldest cells is the rate at which they are cleared from the bloodstream, the team made several predictions, which were validated by applying their model to clinical data from the blood samples and to data from several published studies.
The researchers found that increased RDW was associated with delayed RBC clearance, and increased RDW was associated with increased average age of RBCs.
Delayed RBC clearance was as strongly associated with overall risk of death, as was increased RDW, and delayed RBC clearance was associated with the presence of early signs of hidden diseases associated with increased RDW.
Patients with delayed RBC clearance had a greater risk of developing signs of those diseases in the future than patients with a typical clearance rate. In healthy patients, the rate of RBC clearance varied less than any other traditional blood-count characteristic.
Dr Higgins said there are many potential clinical applications of these findings, which need to be validated by future studies.
“Finding a reduced clearance rate in an apparently healthy person would likely mean that an underlying disease process was developing—such as the early stages of iron deficiency, kidney disease, colon cancer, or congestive heart failure—and would warrant further diagnostic evaluation,” Dr Higgins said.
“Based on this analysis of routine blood tests, a primary care physician could immediately consider appropriate follow-up diagnostic testing, instead of waiting for other signs and symptoms to appear as the condition progresses. In a patient with established disease, a reduced clearance rate could mean progression of disease or treatment failure, and imminent complications could be avoided or reduced by adjusting treatment right away or at least by more frequent monitoring.”
“In addition to confirming our findings in prospective studies that would follow a group of patients over time, we hope to identify the diseases for which an early warning provided by delayed clearance would lead to the most significant improvements in patient outcomes. We’d also like to understand more about the processes controlling red blood cell clearance and are actively developing similar models for populations of white blood cells and platelets.”