User login
Interventions reduce bloodstream infections
A bundled intervention can considerably reduce central line-associated bloodstream infections (CLABSIs), according to research published in Infection Control & Hospital Epidemiology.
The intervention focused on evidence-based infection prevention practices, safety culture and teamwork, and scheduled measurement of infection rates.
By implementing these measures, intensive care units (ICUs) in Abu Dhabi achieved an overall 38% reduction in CLABSIs.
“These hospitals were able to show significant improvements in infection rates and have been able to sustain the improvements a year after we finished the project,” said study author Asad Latif, MBBS, MD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
“Our results suggest that ICUs in disparate settings around the world could use this program and achieve similar results, significantly reducing the global morbidity, mortality, and excess costs associated with CLABSIs. In addition, this collaborative could serve as a model for future efforts to reduce other types of preventable medical harms in the Middle East and around the world.”
This study was a collaborative effort by the Armstrong Institute, Johns Hopkins Medicine International, and the Abu Dhabi Health Services Company (SEHA), which operates the government healthcare system in Abu Dhabi.
For the study, ICUs were instructed to assemble a comprehensive unit-based safety program (CUSP) team comprising local physician and nursing leaders, a senior executive, frontline healthcare providers, an infection control provider, and hospital quality and safety leaders.
The ICUs included 10 adult, 5 neonatal, and 3 pediatric ICUs, accounting for 77% of the adult, 74% of the neonatal, and 100% of the pediatric ICU beds in Abu Dhabi.
Starting in May 2012, the SEHA corporate quality team and ICU CUSP teams attended 14 weekly live webinars on CLABSI prevention conducted by Armstrong Institute faculty, followed by content and coaching webinars every 2 weeks for 24 months. The webinars were recorded by SEHA and posted on a local, shared computer drive, along with educational and training materials.
Armstrong faculty also conducted 4 site visits in Abu Dhabi at the beginning of the study, visiting each ICU to meet the CUSP team and tour the units. A year later, they conducted a 3-day patient safety workshop for participating hospitals.
CUSP teams implemented 3 interventions as part of the program: an effort to prevent CLABSIs that targeted clinicians’ use of evidence-based infection prevention recommendations from the Centers for Disease Control and Prevention, a CUSP process to improve safety culture and teamwork, and measurement of monthly CLABSI data and feedback to safety teams, senior leaders, and ICU staff.
The overall mean crude CLABSI rate for participating ICUs decreased from 2.56 infections per 1000 catheter days to 1.79 per 1,000 catheter days by the end of the study, corresponding to a 30% reduction.
By unit type, CLABSI rates decreased by 16% among adult ICUs, 48% among pediatric ICUs, and 47% in neonatal ICUs. The percentage of ICUs that achieved a quarterly CLABSI rate of less than 1 infection per 1000 catheter days increased from 44% to 61% after the interventions.
“Despite growing awareness, many hospitals around the world continue to struggle in their efforts to meaningfully reduce their CLABSI rates in a sustained manner,” said Sean M. Berenholtz, MD, of the Johns Hopkins University School of Medicine.
“In addition, hospitals and healthcare systems in the Middle East have unique barriers to implementing quality improvement programs, such as challenges with staff recruitment and retention, and personnel fearful of punitive repercussions from speaking up regarding patient safety concerns. In our study, bringing all stakeholders to the same table allowed everyone to share their concerns and ensure their voices were heard.” ![]()

A bundled intervention can considerably reduce central line-associated bloodstream infections (CLABSIs), according to research published in Infection Control & Hospital Epidemiology.
The intervention focused on evidence-based infection prevention practices, safety culture and teamwork, and scheduled measurement of infection rates.
By implementing these measures, intensive care units (ICUs) in Abu Dhabi achieved an overall 38% reduction in CLABSIs.
“These hospitals were able to show significant improvements in infection rates and have been able to sustain the improvements a year after we finished the project,” said study author Asad Latif, MBBS, MD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
“Our results suggest that ICUs in disparate settings around the world could use this program and achieve similar results, significantly reducing the global morbidity, mortality, and excess costs associated with CLABSIs. In addition, this collaborative could serve as a model for future efforts to reduce other types of preventable medical harms in the Middle East and around the world.”
This study was a collaborative effort by the Armstrong Institute, Johns Hopkins Medicine International, and the Abu Dhabi Health Services Company (SEHA), which operates the government healthcare system in Abu Dhabi.
For the study, ICUs were instructed to assemble a comprehensive unit-based safety program (CUSP) team comprising local physician and nursing leaders, a senior executive, frontline healthcare providers, an infection control provider, and hospital quality and safety leaders.
The ICUs included 10 adult, 5 neonatal, and 3 pediatric ICUs, accounting for 77% of the adult, 74% of the neonatal, and 100% of the pediatric ICU beds in Abu Dhabi.
Starting in May 2012, the SEHA corporate quality team and ICU CUSP teams attended 14 weekly live webinars on CLABSI prevention conducted by Armstrong Institute faculty, followed by content and coaching webinars every 2 weeks for 24 months. The webinars were recorded by SEHA and posted on a local, shared computer drive, along with educational and training materials.
Armstrong faculty also conducted 4 site visits in Abu Dhabi at the beginning of the study, visiting each ICU to meet the CUSP team and tour the units. A year later, they conducted a 3-day patient safety workshop for participating hospitals.
CUSP teams implemented 3 interventions as part of the program: an effort to prevent CLABSIs that targeted clinicians’ use of evidence-based infection prevention recommendations from the Centers for Disease Control and Prevention, a CUSP process to improve safety culture and teamwork, and measurement of monthly CLABSI data and feedback to safety teams, senior leaders, and ICU staff.
The overall mean crude CLABSI rate for participating ICUs decreased from 2.56 infections per 1000 catheter days to 1.79 per 1,000 catheter days by the end of the study, corresponding to a 30% reduction.
By unit type, CLABSI rates decreased by 16% among adult ICUs, 48% among pediatric ICUs, and 47% in neonatal ICUs. The percentage of ICUs that achieved a quarterly CLABSI rate of less than 1 infection per 1000 catheter days increased from 44% to 61% after the interventions.
“Despite growing awareness, many hospitals around the world continue to struggle in their efforts to meaningfully reduce their CLABSI rates in a sustained manner,” said Sean M. Berenholtz, MD, of the Johns Hopkins University School of Medicine.
“In addition, hospitals and healthcare systems in the Middle East have unique barriers to implementing quality improvement programs, such as challenges with staff recruitment and retention, and personnel fearful of punitive repercussions from speaking up regarding patient safety concerns. In our study, bringing all stakeholders to the same table allowed everyone to share their concerns and ensure their voices were heard.” ![]()

A bundled intervention can considerably reduce central line-associated bloodstream infections (CLABSIs), according to research published in Infection Control & Hospital Epidemiology.
The intervention focused on evidence-based infection prevention practices, safety culture and teamwork, and scheduled measurement of infection rates.
By implementing these measures, intensive care units (ICUs) in Abu Dhabi achieved an overall 38% reduction in CLABSIs.
“These hospitals were able to show significant improvements in infection rates and have been able to sustain the improvements a year after we finished the project,” said study author Asad Latif, MBBS, MD, of the Johns Hopkins University School of Medicine in Baltimore, Maryland.
“Our results suggest that ICUs in disparate settings around the world could use this program and achieve similar results, significantly reducing the global morbidity, mortality, and excess costs associated with CLABSIs. In addition, this collaborative could serve as a model for future efforts to reduce other types of preventable medical harms in the Middle East and around the world.”
This study was a collaborative effort by the Armstrong Institute, Johns Hopkins Medicine International, and the Abu Dhabi Health Services Company (SEHA), which operates the government healthcare system in Abu Dhabi.
For the study, ICUs were instructed to assemble a comprehensive unit-based safety program (CUSP) team comprising local physician and nursing leaders, a senior executive, frontline healthcare providers, an infection control provider, and hospital quality and safety leaders.
The ICUs included 10 adult, 5 neonatal, and 3 pediatric ICUs, accounting for 77% of the adult, 74% of the neonatal, and 100% of the pediatric ICU beds in Abu Dhabi.
Starting in May 2012, the SEHA corporate quality team and ICU CUSP teams attended 14 weekly live webinars on CLABSI prevention conducted by Armstrong Institute faculty, followed by content and coaching webinars every 2 weeks for 24 months. The webinars were recorded by SEHA and posted on a local, shared computer drive, along with educational and training materials.
Armstrong faculty also conducted 4 site visits in Abu Dhabi at the beginning of the study, visiting each ICU to meet the CUSP team and tour the units. A year later, they conducted a 3-day patient safety workshop for participating hospitals.
CUSP teams implemented 3 interventions as part of the program: an effort to prevent CLABSIs that targeted clinicians’ use of evidence-based infection prevention recommendations from the Centers for Disease Control and Prevention, a CUSP process to improve safety culture and teamwork, and measurement of monthly CLABSI data and feedback to safety teams, senior leaders, and ICU staff.
The overall mean crude CLABSI rate for participating ICUs decreased from 2.56 infections per 1000 catheter days to 1.79 per 1,000 catheter days by the end of the study, corresponding to a 30% reduction.
By unit type, CLABSI rates decreased by 16% among adult ICUs, 48% among pediatric ICUs, and 47% in neonatal ICUs. The percentage of ICUs that achieved a quarterly CLABSI rate of less than 1 infection per 1000 catheter days increased from 44% to 61% after the interventions.
“Despite growing awareness, many hospitals around the world continue to struggle in their efforts to meaningfully reduce their CLABSI rates in a sustained manner,” said Sean M. Berenholtz, MD, of the Johns Hopkins University School of Medicine.
“In addition, hospitals and healthcare systems in the Middle East have unique barriers to implementing quality improvement programs, such as challenges with staff recruitment and retention, and personnel fearful of punitive repercussions from speaking up regarding patient safety concerns. In our study, bringing all stakeholders to the same table allowed everyone to share their concerns and ensure their voices were heard.” ![]()
Fake malaria drugs may be less common than we thought
Photo courtesy of the
London School of Hygiene
& Tropical Medicine
Investigations into antimalarial drug quality conducted in Cambodia and Tanzania uncovered no evidence of fake medicines.
Previous reports had suggested that up to a third of antimalarials might be falsified, or do not contain the stated active pharmaceutical ingredient.
The new research revealed no falsified drugs in either country, but it did unearth substandard antimalarial drugs, or genuine medicines that do not have the correct amount of the active ingredient.
These findings were published in 2 articles in the American Journal of Tropical Medicine and Hygiene.
“Although there have been alarming reports about the prevalence of fake antimalarials, our study provides ample data showing that the quality of drugs is not so bad based on comprehensive sampling and analysis presented here,” said Harparkash Kaur, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The lack of falsified medicines in Cambodia and Tanzania are reassuring, but the presence of substandard medicines is definitely a concern.”
The researchers analyzed 2028 antimalarials from Tanzania and Cambodia at 3 independent laboratories in the UK and US. They classified drugs as acceptable, falsified, or substandard.
In Tanzania, the researchers used an “overt sampling” system, telling vendors they were going to analyze the quality of their medicines.
In Cambodia, the researchers used overt sampling as well as a “mystery client” approach, where actors pretended to be patients with malaria, or their relatives, and bought the medicines offered to them.
Both studies used a randomized approach to sampling of drug outlets, which differs from previous studies that mostly used non-representative methods for selecting drugs for analysis.
Neither study unearthed falsified drugs, but substandard drugs were found in 31% of samples in Cambodia and 12% of samples in Tanzania.
“Falsified medicines have received much attention globally, but substandard drugs are far more prevalent and of great concern,” said Shunmay Yeung, MBBS, PhD, also of the London School of Hygiene & Tropical Medicine.
“Not only do they leave patients with malaria undertreated, which could be fatal, but they may also contribute to the development of resistance to [artemisinin-based combination therapies], the most effective drugs for malaria. Generally, the fact that no falsified antimalarials were identified reflects the positive impact of the [countries’ efforts] to control drug quality.” ![]()

Photo courtesy of the
London School of Hygiene
& Tropical Medicine
Investigations into antimalarial drug quality conducted in Cambodia and Tanzania uncovered no evidence of fake medicines.
Previous reports had suggested that up to a third of antimalarials might be falsified, or do not contain the stated active pharmaceutical ingredient.
The new research revealed no falsified drugs in either country, but it did unearth substandard antimalarial drugs, or genuine medicines that do not have the correct amount of the active ingredient.
These findings were published in 2 articles in the American Journal of Tropical Medicine and Hygiene.
“Although there have been alarming reports about the prevalence of fake antimalarials, our study provides ample data showing that the quality of drugs is not so bad based on comprehensive sampling and analysis presented here,” said Harparkash Kaur, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The lack of falsified medicines in Cambodia and Tanzania are reassuring, but the presence of substandard medicines is definitely a concern.”
The researchers analyzed 2028 antimalarials from Tanzania and Cambodia at 3 independent laboratories in the UK and US. They classified drugs as acceptable, falsified, or substandard.
In Tanzania, the researchers used an “overt sampling” system, telling vendors they were going to analyze the quality of their medicines.
In Cambodia, the researchers used overt sampling as well as a “mystery client” approach, where actors pretended to be patients with malaria, or their relatives, and bought the medicines offered to them.
Both studies used a randomized approach to sampling of drug outlets, which differs from previous studies that mostly used non-representative methods for selecting drugs for analysis.
Neither study unearthed falsified drugs, but substandard drugs were found in 31% of samples in Cambodia and 12% of samples in Tanzania.
“Falsified medicines have received much attention globally, but substandard drugs are far more prevalent and of great concern,” said Shunmay Yeung, MBBS, PhD, also of the London School of Hygiene & Tropical Medicine.
“Not only do they leave patients with malaria undertreated, which could be fatal, but they may also contribute to the development of resistance to [artemisinin-based combination therapies], the most effective drugs for malaria. Generally, the fact that no falsified antimalarials were identified reflects the positive impact of the [countries’ efforts] to control drug quality.” ![]()

Photo courtesy of the
London School of Hygiene
& Tropical Medicine
Investigations into antimalarial drug quality conducted in Cambodia and Tanzania uncovered no evidence of fake medicines.
Previous reports had suggested that up to a third of antimalarials might be falsified, or do not contain the stated active pharmaceutical ingredient.
The new research revealed no falsified drugs in either country, but it did unearth substandard antimalarial drugs, or genuine medicines that do not have the correct amount of the active ingredient.
These findings were published in 2 articles in the American Journal of Tropical Medicine and Hygiene.
“Although there have been alarming reports about the prevalence of fake antimalarials, our study provides ample data showing that the quality of drugs is not so bad based on comprehensive sampling and analysis presented here,” said Harparkash Kaur, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The lack of falsified medicines in Cambodia and Tanzania are reassuring, but the presence of substandard medicines is definitely a concern.”
The researchers analyzed 2028 antimalarials from Tanzania and Cambodia at 3 independent laboratories in the UK and US. They classified drugs as acceptable, falsified, or substandard.
In Tanzania, the researchers used an “overt sampling” system, telling vendors they were going to analyze the quality of their medicines.
In Cambodia, the researchers used overt sampling as well as a “mystery client” approach, where actors pretended to be patients with malaria, or their relatives, and bought the medicines offered to them.
Both studies used a randomized approach to sampling of drug outlets, which differs from previous studies that mostly used non-representative methods for selecting drugs for analysis.
Neither study unearthed falsified drugs, but substandard drugs were found in 31% of samples in Cambodia and 12% of samples in Tanzania.
“Falsified medicines have received much attention globally, but substandard drugs are far more prevalent and of great concern,” said Shunmay Yeung, MBBS, PhD, also of the London School of Hygiene & Tropical Medicine.
“Not only do they leave patients with malaria undertreated, which could be fatal, but they may also contribute to the development of resistance to [artemisinin-based combination therapies], the most effective drugs for malaria. Generally, the fact that no falsified antimalarials were identified reflects the positive impact of the [countries’ efforts] to control drug quality.” ![]()
Mutation linked to drug-resistant malaria in Africa
Image by Ute Frevert
and Margaret Shear
New research provides genetic evidence that malaria parasites in Africa are developing resistance to antimalarial drugs.
Researchers found that Plasmodium falciparum parasites with a mutation in the gene ap2mu were less sensitive to both artemisinin and quinine.
A study published in 2013 suggested a link between a mutation in ap2mu and low levels of malaria parasites remaining in the blood of Kenyan children after treatment with artemisinin.
However, further research was needed to confirm that these genetic characteristics represented an early step toward resistance.
In the new study, published in Antimicrobial Agents and Chemotherapy, researchers genetically altered the malaria parasite to mutate ap2mu in the same way that had been observed in Kenya.
The team found the altered parasite was significantly less susceptible to treatment, requiring 32% more drug to be killed by artemisinin. The genetically altered parasite was also 42.4% less susceptible to quinine.
Earlier this year, a different research group discovered mutations in the gene kelch13, which were linked to reduced susceptibility to artemisinin combination treatment in South East Asia.
Historically, resistance to antimalarial medicines has emerged in South East Asia and then spread to Africa. But these new findings suggest a different route to drug resistance may be developing independently in Africa.
“Our findings could be a sign of much worse things to come for malaria in Africa,” said study author Colin Sutherland, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The malaria parasite is constantly evolving to evade our control efforts. We’ve already moved away from using quinine to treat cases as the malaria parasite has become more resistant to it, but if further drug resistance were to develop against our most valuable malaria drug, artemisinin, we would be facing a grave situation.”
“We now know that the gene ap2mu is an important factor in determining how well our drugs kill malaria parasites. We will be conducting laboratory and field studies to more accurately measure the impact of mutations in the ap2mu gene. We hope our findings will help [us] understand resistance of malaria to drugs and potentially be an important tool for monitoring malaria treatment in the future.” ![]()

Image by Ute Frevert
and Margaret Shear
New research provides genetic evidence that malaria parasites in Africa are developing resistance to antimalarial drugs.
Researchers found that Plasmodium falciparum parasites with a mutation in the gene ap2mu were less sensitive to both artemisinin and quinine.
A study published in 2013 suggested a link between a mutation in ap2mu and low levels of malaria parasites remaining in the blood of Kenyan children after treatment with artemisinin.
However, further research was needed to confirm that these genetic characteristics represented an early step toward resistance.
In the new study, published in Antimicrobial Agents and Chemotherapy, researchers genetically altered the malaria parasite to mutate ap2mu in the same way that had been observed in Kenya.
The team found the altered parasite was significantly less susceptible to treatment, requiring 32% more drug to be killed by artemisinin. The genetically altered parasite was also 42.4% less susceptible to quinine.
Earlier this year, a different research group discovered mutations in the gene kelch13, which were linked to reduced susceptibility to artemisinin combination treatment in South East Asia.
Historically, resistance to antimalarial medicines has emerged in South East Asia and then spread to Africa. But these new findings suggest a different route to drug resistance may be developing independently in Africa.
“Our findings could be a sign of much worse things to come for malaria in Africa,” said study author Colin Sutherland, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The malaria parasite is constantly evolving to evade our control efforts. We’ve already moved away from using quinine to treat cases as the malaria parasite has become more resistant to it, but if further drug resistance were to develop against our most valuable malaria drug, artemisinin, we would be facing a grave situation.”
“We now know that the gene ap2mu is an important factor in determining how well our drugs kill malaria parasites. We will be conducting laboratory and field studies to more accurately measure the impact of mutations in the ap2mu gene. We hope our findings will help [us] understand resistance of malaria to drugs and potentially be an important tool for monitoring malaria treatment in the future.” ![]()

Image by Ute Frevert
and Margaret Shear
New research provides genetic evidence that malaria parasites in Africa are developing resistance to antimalarial drugs.
Researchers found that Plasmodium falciparum parasites with a mutation in the gene ap2mu were less sensitive to both artemisinin and quinine.
A study published in 2013 suggested a link between a mutation in ap2mu and low levels of malaria parasites remaining in the blood of Kenyan children after treatment with artemisinin.
However, further research was needed to confirm that these genetic characteristics represented an early step toward resistance.
In the new study, published in Antimicrobial Agents and Chemotherapy, researchers genetically altered the malaria parasite to mutate ap2mu in the same way that had been observed in Kenya.
The team found the altered parasite was significantly less susceptible to treatment, requiring 32% more drug to be killed by artemisinin. The genetically altered parasite was also 42.4% less susceptible to quinine.
Earlier this year, a different research group discovered mutations in the gene kelch13, which were linked to reduced susceptibility to artemisinin combination treatment in South East Asia.
Historically, resistance to antimalarial medicines has emerged in South East Asia and then spread to Africa. But these new findings suggest a different route to drug resistance may be developing independently in Africa.
“Our findings could be a sign of much worse things to come for malaria in Africa,” said study author Colin Sutherland, PhD, of the London School of Hygiene & Tropical Medicine in the UK.
“The malaria parasite is constantly evolving to evade our control efforts. We’ve already moved away from using quinine to treat cases as the malaria parasite has become more resistant to it, but if further drug resistance were to develop against our most valuable malaria drug, artemisinin, we would be facing a grave situation.”
“We now know that the gene ap2mu is an important factor in determining how well our drugs kill malaria parasites. We will be conducting laboratory and field studies to more accurately measure the impact of mutations in the ap2mu gene. We hope our findings will help [us] understand resistance of malaria to drugs and potentially be an important tool for monitoring malaria treatment in the future.” ![]()
Explaining drug-resistant malaria
infecting a red blood cell
Image courtesy of St. Jude
Children’s Research Hospital
Researchers say they have identified a molecular mechanism responsible for making malaria parasites resistant to artemisinins, the leading class of antimalarial drugs.
The team found that a kinase, Plasmodium falciparum phosphatidylinositol-3-kinase (PfPI3K), and its lipid product, phosphatidylinositol-3-phosphate (PI3P), play key roles in artemisinin resistance.
So targeting PfPI3K or PI3P could potentially treat resistant Plasmodium falciparum malaria.
Alassane Mbengue, PhD, of the University of Notre Dame in Indiana, and his colleagues described this research in a letter to Nature.
“We observed that levels of [PI3P] were higher in artemisinin-resistant P falciparum than artemisinin-sensitive strains,” Dr Mbengue said. “This lipid is produced by an enzyme called PfPI3K. We found that artemisinins block this kinase from producing PI3P lipids. We also discovered that the amount of the kinase present in the parasite is controlled by the gene PfKelch13.”
“Mutation in the gene increases the kinase levels, which, in turn, increases PI3P lipid levels. The higher the level of PI3P lipids present in the parasite, the greater the level of artemisinin resistance. We also studied the lipid levels in parasites without the gene mutation and observed that when PI3P lipid levels were increased artificially, the parasites still became proportionately resistant.”
Specifically, the researchers found that increased PfPI3K was associated with the C580Y mutation in PfKelch13. The mutation reduced polyubiquitination of PfPI3K and its binding to PfKelch13, which limited proteolysis of PfPI3K and led to increased levels of both PfPI3K and PI3P.
The team found that PI3P levels were predictive of artemisinin resistance in clinical and engineered parasites. And although increases in PI3P levels induced artemisinin resistance in the absence of PfKelch13 mutations, PI3P levels were still responsive to regulation by PfKelch13.
Dr Mbengue and his colleagues said the next step for this research is to identify drugs that can kill P falciparum by preventing PfPI3K from making PI3P or disrupting the production of the kinase itself. ![]()

infecting a red blood cell
Image courtesy of St. Jude
Children’s Research Hospital
Researchers say they have identified a molecular mechanism responsible for making malaria parasites resistant to artemisinins, the leading class of antimalarial drugs.
The team found that a kinase, Plasmodium falciparum phosphatidylinositol-3-kinase (PfPI3K), and its lipid product, phosphatidylinositol-3-phosphate (PI3P), play key roles in artemisinin resistance.
So targeting PfPI3K or PI3P could potentially treat resistant Plasmodium falciparum malaria.
Alassane Mbengue, PhD, of the University of Notre Dame in Indiana, and his colleagues described this research in a letter to Nature.
“We observed that levels of [PI3P] were higher in artemisinin-resistant P falciparum than artemisinin-sensitive strains,” Dr Mbengue said. “This lipid is produced by an enzyme called PfPI3K. We found that artemisinins block this kinase from producing PI3P lipids. We also discovered that the amount of the kinase present in the parasite is controlled by the gene PfKelch13.”
“Mutation in the gene increases the kinase levels, which, in turn, increases PI3P lipid levels. The higher the level of PI3P lipids present in the parasite, the greater the level of artemisinin resistance. We also studied the lipid levels in parasites without the gene mutation and observed that when PI3P lipid levels were increased artificially, the parasites still became proportionately resistant.”
Specifically, the researchers found that increased PfPI3K was associated with the C580Y mutation in PfKelch13. The mutation reduced polyubiquitination of PfPI3K and its binding to PfKelch13, which limited proteolysis of PfPI3K and led to increased levels of both PfPI3K and PI3P.
The team found that PI3P levels were predictive of artemisinin resistance in clinical and engineered parasites. And although increases in PI3P levels induced artemisinin resistance in the absence of PfKelch13 mutations, PI3P levels were still responsive to regulation by PfKelch13.
Dr Mbengue and his colleagues said the next step for this research is to identify drugs that can kill P falciparum by preventing PfPI3K from making PI3P or disrupting the production of the kinase itself. ![]()

infecting a red blood cell
Image courtesy of St. Jude
Children’s Research Hospital
Researchers say they have identified a molecular mechanism responsible for making malaria parasites resistant to artemisinins, the leading class of antimalarial drugs.
The team found that a kinase, Plasmodium falciparum phosphatidylinositol-3-kinase (PfPI3K), and its lipid product, phosphatidylinositol-3-phosphate (PI3P), play key roles in artemisinin resistance.
So targeting PfPI3K or PI3P could potentially treat resistant Plasmodium falciparum malaria.
Alassane Mbengue, PhD, of the University of Notre Dame in Indiana, and his colleagues described this research in a letter to Nature.
“We observed that levels of [PI3P] were higher in artemisinin-resistant P falciparum than artemisinin-sensitive strains,” Dr Mbengue said. “This lipid is produced by an enzyme called PfPI3K. We found that artemisinins block this kinase from producing PI3P lipids. We also discovered that the amount of the kinase present in the parasite is controlled by the gene PfKelch13.”
“Mutation in the gene increases the kinase levels, which, in turn, increases PI3P lipid levels. The higher the level of PI3P lipids present in the parasite, the greater the level of artemisinin resistance. We also studied the lipid levels in parasites without the gene mutation and observed that when PI3P lipid levels were increased artificially, the parasites still became proportionately resistant.”
Specifically, the researchers found that increased PfPI3K was associated with the C580Y mutation in PfKelch13. The mutation reduced polyubiquitination of PfPI3K and its binding to PfKelch13, which limited proteolysis of PfPI3K and led to increased levels of both PfPI3K and PI3P.
The team found that PI3P levels were predictive of artemisinin resistance in clinical and engineered parasites. And although increases in PI3P levels induced artemisinin resistance in the absence of PfKelch13 mutations, PI3P levels were still responsive to regulation by PfKelch13.
Dr Mbengue and his colleagues said the next step for this research is to identify drugs that can kill P falciparum by preventing PfPI3K from making PI3P or disrupting the production of the kinase itself. ![]()
Team uncovers new role for macrophages
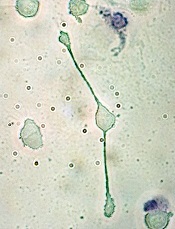
pseudopodia to engulf particles
Macrophages can substitute for dendritic cells as primers of T-cell-dependent immune responses, according to research published in Proceedings of the National Academy of Sciences.
The study showed that macrophages that function as a first line of defense in the innate immune system can also present antigens to T cells, a previously unknown role for macrophages in the induction of adaptive immune responses.
“It has been assumed until now that the dendritic cells are considered to be essentially the only cell type responsible for antigen presentation in the immune system,” said Dr Thomas Brocker, of Ludwig-Maximilians-Universität München in Germany.
“We have now discovered that macrophages can also do this job. Not only that, in certain situations, they can be more effective than dendritic cells.”
Dendritic cells present antigens to cytotoxic T lymphocytes (CTLs) if they have been directly infected, but they can also capture and display antigens from other cells. This type of indirect antigen presentation is referred to as cross-presentation.
“So, theoretically, dendritic cells could be responsible for the induction of all CTL-based responses, regardless of whether they are themselves infected or not,” Dr Brocker said. “But the significance of cross-presentation is hotly debated in the literature.”
Dr Brocker and his team used several antigens that were specifically targeted to and processed by macrophages but could not be taken up directly by dendritic cells. They were able to demonstrate that each antigen induced a normal immune response in a mouse model system, and even in a mouse strain that lacked dendritic cells altogether.
Further experiments showed the targeted macrophages were actually able to prime a more comprehensive immune reaction than cross-presenting dendritic cells. They activated T cells specific for all antigen-binding sites (epitopes) presented, whereas cross-presentation by dendritic cells stimulates only those T cells that recognize immunodominant epitopes.
“Macrophages naturally function as filters,” Dr Brocker noted. “They gobble everything up that might be harmful to the organism. And our study shows that, in contrast to cross-priming dendritic cells, they are capable of producing and presenting all T-cell-priming epitopes we tested. Macrophages therefore induce a complete immune response. These observations indicate that the significance of cross-presentation by dendritic cells has been overrated.”
He added that these findings are relevant for the development of immunization strategies.
“Preclinical trials are already underway with vaccines that are designed to activate specific sets of dendritic cells,” Dr Brocker said. “But the weak epitopes are important for a broadly directed immune response, because they can potentially recognize mutant variants of viruses, for instance.”
“Cross-priming dendritic cells fail to induce weakly antigenic epitopes, as our study shows. Our results indicate that it may make more sense to manipulate macrophages directly because they stimulate a wider-ranging immune response.” ![]()

pseudopodia to engulf particles
Macrophages can substitute for dendritic cells as primers of T-cell-dependent immune responses, according to research published in Proceedings of the National Academy of Sciences.
The study showed that macrophages that function as a first line of defense in the innate immune system can also present antigens to T cells, a previously unknown role for macrophages in the induction of adaptive immune responses.
“It has been assumed until now that the dendritic cells are considered to be essentially the only cell type responsible for antigen presentation in the immune system,” said Dr Thomas Brocker, of Ludwig-Maximilians-Universität München in Germany.
“We have now discovered that macrophages can also do this job. Not only that, in certain situations, they can be more effective than dendritic cells.”
Dendritic cells present antigens to cytotoxic T lymphocytes (CTLs) if they have been directly infected, but they can also capture and display antigens from other cells. This type of indirect antigen presentation is referred to as cross-presentation.
“So, theoretically, dendritic cells could be responsible for the induction of all CTL-based responses, regardless of whether they are themselves infected or not,” Dr Brocker said. “But the significance of cross-presentation is hotly debated in the literature.”
Dr Brocker and his team used several antigens that were specifically targeted to and processed by macrophages but could not be taken up directly by dendritic cells. They were able to demonstrate that each antigen induced a normal immune response in a mouse model system, and even in a mouse strain that lacked dendritic cells altogether.
Further experiments showed the targeted macrophages were actually able to prime a more comprehensive immune reaction than cross-presenting dendritic cells. They activated T cells specific for all antigen-binding sites (epitopes) presented, whereas cross-presentation by dendritic cells stimulates only those T cells that recognize immunodominant epitopes.
“Macrophages naturally function as filters,” Dr Brocker noted. “They gobble everything up that might be harmful to the organism. And our study shows that, in contrast to cross-priming dendritic cells, they are capable of producing and presenting all T-cell-priming epitopes we tested. Macrophages therefore induce a complete immune response. These observations indicate that the significance of cross-presentation by dendritic cells has been overrated.”
He added that these findings are relevant for the development of immunization strategies.
“Preclinical trials are already underway with vaccines that are designed to activate specific sets of dendritic cells,” Dr Brocker said. “But the weak epitopes are important for a broadly directed immune response, because they can potentially recognize mutant variants of viruses, for instance.”
“Cross-priming dendritic cells fail to induce weakly antigenic epitopes, as our study shows. Our results indicate that it may make more sense to manipulate macrophages directly because they stimulate a wider-ranging immune response.” ![]()

pseudopodia to engulf particles
Macrophages can substitute for dendritic cells as primers of T-cell-dependent immune responses, according to research published in Proceedings of the National Academy of Sciences.
The study showed that macrophages that function as a first line of defense in the innate immune system can also present antigens to T cells, a previously unknown role for macrophages in the induction of adaptive immune responses.
“It has been assumed until now that the dendritic cells are considered to be essentially the only cell type responsible for antigen presentation in the immune system,” said Dr Thomas Brocker, of Ludwig-Maximilians-Universität München in Germany.
“We have now discovered that macrophages can also do this job. Not only that, in certain situations, they can be more effective than dendritic cells.”
Dendritic cells present antigens to cytotoxic T lymphocytes (CTLs) if they have been directly infected, but they can also capture and display antigens from other cells. This type of indirect antigen presentation is referred to as cross-presentation.
“So, theoretically, dendritic cells could be responsible for the induction of all CTL-based responses, regardless of whether they are themselves infected or not,” Dr Brocker said. “But the significance of cross-presentation is hotly debated in the literature.”
Dr Brocker and his team used several antigens that were specifically targeted to and processed by macrophages but could not be taken up directly by dendritic cells. They were able to demonstrate that each antigen induced a normal immune response in a mouse model system, and even in a mouse strain that lacked dendritic cells altogether.
Further experiments showed the targeted macrophages were actually able to prime a more comprehensive immune reaction than cross-presenting dendritic cells. They activated T cells specific for all antigen-binding sites (epitopes) presented, whereas cross-presentation by dendritic cells stimulates only those T cells that recognize immunodominant epitopes.
“Macrophages naturally function as filters,” Dr Brocker noted. “They gobble everything up that might be harmful to the organism. And our study shows that, in contrast to cross-priming dendritic cells, they are capable of producing and presenting all T-cell-priming epitopes we tested. Macrophages therefore induce a complete immune response. These observations indicate that the significance of cross-presentation by dendritic cells has been overrated.”
He added that these findings are relevant for the development of immunization strategies.
“Preclinical trials are already underway with vaccines that are designed to activate specific sets of dendritic cells,” Dr Brocker said. “But the weak epitopes are important for a broadly directed immune response, because they can potentially recognize mutant variants of viruses, for instance.”
“Cross-priming dendritic cells fail to induce weakly antigenic epitopes, as our study shows. Our results indicate that it may make more sense to manipulate macrophages directly because they stimulate a wider-ranging immune response.” ![]()
SGR repeal dubbed ‘victory’ for cancer patients
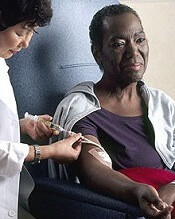
chemotherapy
Photo by Rhoda Baer
A bill that repeals the sustainable growth rate (SGR) formula for physician reimbursement under Medicare is a victory for cancer patients, according to oncologist groups.
The bill—known as H.R.2—passed both the US House of Representatives and the Senate with an overwhelming majority. It must still be signed into law by President Obama, but he has indicated he will sign it.
By repealing the SGR payment methodology, the bill will prevent a 21.2% reduction in physician reimbursement rates.
“Today’s courageous vote by the US Senate to finally end the sustainable growth rate formula is a vote for the millions of patients with cancer who depend on Medicare to help them fight their disease,” said Peter Paul Yu, MD, president of the American Society of Clinical Oncology.
“With Congress passing this historic legislation to finally end the 13-year SGR roller coaster ride, Medicare beneficiaries and their physicians can breathe easier knowing that they will no longer face the perennial threat of payment cuts that risk disruption of care and cause anxiety among patients.”
Under H.R. 2, Medicare’s physician reimbursements will increase by 0.5% in the second half of 2015, then an additional 0.5% annually from 2016 through the end of 2019. The 2019 rates will be maintained through 2025 with no additional increases.
The bill also includes comprehensive structural changes to Medicare’s reimbursement model that aim to promote physician participation in clinical quality improvement activities and value-based care that will take full effect in 2019.
Current Medicare programs that reward electronic health records, quality reporting, the value-based modifier, and meaningful use will be merged by 2019 to encourage participation and to reduce the administrative burden.
The bill also ensures the Children’s Health Insurance Program will receive funding for 2 more years and allocates $7.2 billion for community health centers.
“[P]assage of this legislation represents a long-awaited, historic victory for our patients,” said Bruce G. Haffty, MD, chair of the American Society for Radiation Oncology’s board of directors.
“Permanently repealing the SGR and replacing it with a stabilized reimbursement plan focused on quality will strengthen Medicare and allow us to enhance cancer care for the more than 1 million patients treated with radiation therapy each year.” ![]()

chemotherapy
Photo by Rhoda Baer
A bill that repeals the sustainable growth rate (SGR) formula for physician reimbursement under Medicare is a victory for cancer patients, according to oncologist groups.
The bill—known as H.R.2—passed both the US House of Representatives and the Senate with an overwhelming majority. It must still be signed into law by President Obama, but he has indicated he will sign it.
By repealing the SGR payment methodology, the bill will prevent a 21.2% reduction in physician reimbursement rates.
“Today’s courageous vote by the US Senate to finally end the sustainable growth rate formula is a vote for the millions of patients with cancer who depend on Medicare to help them fight their disease,” said Peter Paul Yu, MD, president of the American Society of Clinical Oncology.
“With Congress passing this historic legislation to finally end the 13-year SGR roller coaster ride, Medicare beneficiaries and their physicians can breathe easier knowing that they will no longer face the perennial threat of payment cuts that risk disruption of care and cause anxiety among patients.”
Under H.R. 2, Medicare’s physician reimbursements will increase by 0.5% in the second half of 2015, then an additional 0.5% annually from 2016 through the end of 2019. The 2019 rates will be maintained through 2025 with no additional increases.
The bill also includes comprehensive structural changes to Medicare’s reimbursement model that aim to promote physician participation in clinical quality improvement activities and value-based care that will take full effect in 2019.
Current Medicare programs that reward electronic health records, quality reporting, the value-based modifier, and meaningful use will be merged by 2019 to encourage participation and to reduce the administrative burden.
The bill also ensures the Children’s Health Insurance Program will receive funding for 2 more years and allocates $7.2 billion for community health centers.
“[P]assage of this legislation represents a long-awaited, historic victory for our patients,” said Bruce G. Haffty, MD, chair of the American Society for Radiation Oncology’s board of directors.
“Permanently repealing the SGR and replacing it with a stabilized reimbursement plan focused on quality will strengthen Medicare and allow us to enhance cancer care for the more than 1 million patients treated with radiation therapy each year.” ![]()

chemotherapy
Photo by Rhoda Baer
A bill that repeals the sustainable growth rate (SGR) formula for physician reimbursement under Medicare is a victory for cancer patients, according to oncologist groups.
The bill—known as H.R.2—passed both the US House of Representatives and the Senate with an overwhelming majority. It must still be signed into law by President Obama, but he has indicated he will sign it.
By repealing the SGR payment methodology, the bill will prevent a 21.2% reduction in physician reimbursement rates.
“Today’s courageous vote by the US Senate to finally end the sustainable growth rate formula is a vote for the millions of patients with cancer who depend on Medicare to help them fight their disease,” said Peter Paul Yu, MD, president of the American Society of Clinical Oncology.
“With Congress passing this historic legislation to finally end the 13-year SGR roller coaster ride, Medicare beneficiaries and their physicians can breathe easier knowing that they will no longer face the perennial threat of payment cuts that risk disruption of care and cause anxiety among patients.”
Under H.R. 2, Medicare’s physician reimbursements will increase by 0.5% in the second half of 2015, then an additional 0.5% annually from 2016 through the end of 2019. The 2019 rates will be maintained through 2025 with no additional increases.
The bill also includes comprehensive structural changes to Medicare’s reimbursement model that aim to promote physician participation in clinical quality improvement activities and value-based care that will take full effect in 2019.
Current Medicare programs that reward electronic health records, quality reporting, the value-based modifier, and meaningful use will be merged by 2019 to encourage participation and to reduce the administrative burden.
The bill also ensures the Children’s Health Insurance Program will receive funding for 2 more years and allocates $7.2 billion for community health centers.
“[P]assage of this legislation represents a long-awaited, historic victory for our patients,” said Bruce G. Haffty, MD, chair of the American Society for Radiation Oncology’s board of directors.
“Permanently repealing the SGR and replacing it with a stabilized reimbursement plan focused on quality will strengthen Medicare and allow us to enhance cancer care for the more than 1 million patients treated with radiation therapy each year.” ![]()
Data breaches of health information on the rise

Photo courtesy of NIH
A new study suggests data breaches of protected health information are on the rise in the US.
Researchers found that, between 2010 and 2013, there were data breaches affecting approximately 29 million records of health information covered
under the Health Insurance Portability and Accountability Act (HIPAA).
Breaches were reported in every state, tended to occur via electronic media, and largely resulted from overt criminal activity.
Vincent Liu, MD, of the Kaiser Permanente Division of Research in Oakland, California, and his colleagues published these findings in JAMA.
The researchers evaluated an online database maintained by the US Department of Health and Human Services that describes data breaches of unencrypted, protected health information (ie, individually identifiable information) reported by entities (health plans and clinicians) covered under HIPAA.
The team included breaches affecting 500 individuals or more that were reported as occurring from 2010 through 2013, accounting for 82% of all reports.
The research revealed 949 breaches affecting 29.1 million records. Six breaches involved more than 1 million records each.
The number of reported breaches increased over time, from 214 in 2010 to 265 in 2013.
Breaches were reported in every state, the District of Columbia, and Puerto Rico. Five states (California, Texas, Florida, New York, and Illinois) accounted for 34% of all breaches. However, when adjusted by population estimates, the states with the highest adjusted number of breaches and affected records varied.
Most breaches occurred via electronic media (67%), frequently involving laptop computers or portable electronic devices (33%). Most breaches also occurred via theft (58%).
The combined frequency of breaches resulting from hacking and unauthorized access or disclosure increased during the study period, from 12% in 2010 to 27% in 2013. Breaches involved external vendors in 29% of reports.
The researchers noted that this study was limited to breaches that were already recognized, reported, and affected at least 500 individuals. Therefore, the team likely underestimated the true number of healthcare data breaches occurring in the US each year. ![]()

Photo courtesy of NIH
A new study suggests data breaches of protected health information are on the rise in the US.
Researchers found that, between 2010 and 2013, there were data breaches affecting approximately 29 million records of health information covered
under the Health Insurance Portability and Accountability Act (HIPAA).
Breaches were reported in every state, tended to occur via electronic media, and largely resulted from overt criminal activity.
Vincent Liu, MD, of the Kaiser Permanente Division of Research in Oakland, California, and his colleagues published these findings in JAMA.
The researchers evaluated an online database maintained by the US Department of Health and Human Services that describes data breaches of unencrypted, protected health information (ie, individually identifiable information) reported by entities (health plans and clinicians) covered under HIPAA.
The team included breaches affecting 500 individuals or more that were reported as occurring from 2010 through 2013, accounting for 82% of all reports.
The research revealed 949 breaches affecting 29.1 million records. Six breaches involved more than 1 million records each.
The number of reported breaches increased over time, from 214 in 2010 to 265 in 2013.
Breaches were reported in every state, the District of Columbia, and Puerto Rico. Five states (California, Texas, Florida, New York, and Illinois) accounted for 34% of all breaches. However, when adjusted by population estimates, the states with the highest adjusted number of breaches and affected records varied.
Most breaches occurred via electronic media (67%), frequently involving laptop computers or portable electronic devices (33%). Most breaches also occurred via theft (58%).
The combined frequency of breaches resulting from hacking and unauthorized access or disclosure increased during the study period, from 12% in 2010 to 27% in 2013. Breaches involved external vendors in 29% of reports.
The researchers noted that this study was limited to breaches that were already recognized, reported, and affected at least 500 individuals. Therefore, the team likely underestimated the true number of healthcare data breaches occurring in the US each year. ![]()

Photo courtesy of NIH
A new study suggests data breaches of protected health information are on the rise in the US.
Researchers found that, between 2010 and 2013, there were data breaches affecting approximately 29 million records of health information covered
under the Health Insurance Portability and Accountability Act (HIPAA).
Breaches were reported in every state, tended to occur via electronic media, and largely resulted from overt criminal activity.
Vincent Liu, MD, of the Kaiser Permanente Division of Research in Oakland, California, and his colleagues published these findings in JAMA.
The researchers evaluated an online database maintained by the US Department of Health and Human Services that describes data breaches of unencrypted, protected health information (ie, individually identifiable information) reported by entities (health plans and clinicians) covered under HIPAA.
The team included breaches affecting 500 individuals or more that were reported as occurring from 2010 through 2013, accounting for 82% of all reports.
The research revealed 949 breaches affecting 29.1 million records. Six breaches involved more than 1 million records each.
The number of reported breaches increased over time, from 214 in 2010 to 265 in 2013.
Breaches were reported in every state, the District of Columbia, and Puerto Rico. Five states (California, Texas, Florida, New York, and Illinois) accounted for 34% of all breaches. However, when adjusted by population estimates, the states with the highest adjusted number of breaches and affected records varied.
Most breaches occurred via electronic media (67%), frequently involving laptop computers or portable electronic devices (33%). Most breaches also occurred via theft (58%).
The combined frequency of breaches resulting from hacking and unauthorized access or disclosure increased during the study period, from 12% in 2010 to 27% in 2013. Breaches involved external vendors in 29% of reports.
The researchers noted that this study was limited to breaches that were already recognized, reported, and affected at least 500 individuals. Therefore, the team likely underestimated the true number of healthcare data breaches occurring in the US each year.
System can diagnose lymphoma, other diseases

Photo by Daniel Sone
Scientists say a smartphone-based system could bring rapid, accurate molecular diagnosis of cancers and other diseases to locations lacking the latest medical technology.
In PNAS, the group explained how the digital diffraction diagnosis (D3) system collects detailed microscopic images for digital analysis of the molecular composition of cells and tissues.
In pilot experiments, the system enabled accurate diagnoses of lymphoma and cervical cancer.
“The emerging genomic and biological data for various cancers, which can be essential to choosing the most appropriate therapy, supports the need for molecular profiling strategies that are more accessible to providers, clinical investigators, and patients,” said study author Cesar Castro, MD, of Massachusetts General Hospital in Boston.
“And we believe the platform we have developed provides essential features at an extraordinarily low cost.”
The D3 system features an imaging module with a battery-powered LED light clipped onto a standard smartphone that records high-resolution imaging data with its camera.
With a greater field of view than traditional microscopy, the D3 system is capable of recording data on more than 100,000 cells from a blood or tissue sample in a single image. The data can then be transmitted for analysis to a remote graphic-processing server via a secure, encrypted cloud service, and the results returned to the point of care.
For molecular analysis of tumors, a sample of blood or tissue is labeled with microbeads that bind to known cancer-related molecules and loaded into the D3 imaging module.
After the image is recorded and data transmitted to the server, the presence of specific molecules is detected by analyzing the diffraction patterns generated by the microbeads. The use of variously sized or coated beads may offer unique diffraction signatures to facilitate detection.
A numerical algorithm the researchers developed can distinguish cells from beads and analyze as much as 10 MB of data in less than nine hundredths of a second.
In a pilot test with cancer cell lines, the D3 system detected the presence of tumor proteins with an accuracy matching that of the current gold standard for molecular profiling. And the system’s larger field of view enabled simultaneous analysis of more than 100,000 cells at a time.
The researchers also conducted analyses of cervical biopsy samples from 25 women with abnormal PAP smears—samples collected along with those used for clinical diagnosis—using microbeads tagged with antibodies against 3 published markers of cervical cancer.
Based on the number of antibody-tagged microbeads binding to cells, D3 analysis promptly and reliably categorized biopsy samples as high-risk, low-risk, or benign. Results matched those of conventional pathologic analysis.
In addition, D3 analysis of fine-needle lymph node biopsy samples was accurately able to differentiate 4 patients whose lymphoma diagnosis was confirmed by conventional pathology from another 4 patients with benign lymph node enlargement.
Along with protein analyses, the D3 system was enhanced to successfully detect DNA—in this instance, from human papilloma virus—with great sensitivity.
In all of these tests, results were available in under an hour and at a cost of $1.80 per assay, a price that would be expected to drop with further refinement of the D3 system.
“We expect that the D3 platform will enhance the breadth and depth of cancer screening in a way that is feasible and sustainable for resource limited-settings,” said Ralph Weissleder, MD, PhD, also of Massachusetts General Hospital.
“By taking advantage of the increased penetration of mobile phone technology worldwide, the system should allow the prompt triaging of suspicious or high-risk cases that could help to offset delays caused by limited pathology services in those regions and reduce the need for patients to return for follow-up care, which is often challenging for them.”
The researchers’ next steps are to investigate D3’s ability to analyze protein and DNA markers of other disease catalysts, integrate the software with larger databases, and conduct clinical studies in settings such as care-delivery sites in developing countries or rural areas.
Massachusetts General Hospital has filed a patent application covering the D3 technology.

Photo by Daniel Sone
Scientists say a smartphone-based system could bring rapid, accurate molecular diagnosis of cancers and other diseases to locations lacking the latest medical technology.
In PNAS, the group explained how the digital diffraction diagnosis (D3) system collects detailed microscopic images for digital analysis of the molecular composition of cells and tissues.
In pilot experiments, the system enabled accurate diagnoses of lymphoma and cervical cancer.
“The emerging genomic and biological data for various cancers, which can be essential to choosing the most appropriate therapy, supports the need for molecular profiling strategies that are more accessible to providers, clinical investigators, and patients,” said study author Cesar Castro, MD, of Massachusetts General Hospital in Boston.
“And we believe the platform we have developed provides essential features at an extraordinarily low cost.”
The D3 system features an imaging module with a battery-powered LED light clipped onto a standard smartphone that records high-resolution imaging data with its camera.
With a greater field of view than traditional microscopy, the D3 system is capable of recording data on more than 100,000 cells from a blood or tissue sample in a single image. The data can then be transmitted for analysis to a remote graphic-processing server via a secure, encrypted cloud service, and the results returned to the point of care.
For molecular analysis of tumors, a sample of blood or tissue is labeled with microbeads that bind to known cancer-related molecules and loaded into the D3 imaging module.
After the image is recorded and data transmitted to the server, the presence of specific molecules is detected by analyzing the diffraction patterns generated by the microbeads. The use of variously sized or coated beads may offer unique diffraction signatures to facilitate detection.
A numerical algorithm the researchers developed can distinguish cells from beads and analyze as much as 10 MB of data in less than nine hundredths of a second.
In a pilot test with cancer cell lines, the D3 system detected the presence of tumor proteins with an accuracy matching that of the current gold standard for molecular profiling. And the system’s larger field of view enabled simultaneous analysis of more than 100,000 cells at a time.
The researchers also conducted analyses of cervical biopsy samples from 25 women with abnormal PAP smears—samples collected along with those used for clinical diagnosis—using microbeads tagged with antibodies against 3 published markers of cervical cancer.
Based on the number of antibody-tagged microbeads binding to cells, D3 analysis promptly and reliably categorized biopsy samples as high-risk, low-risk, or benign. Results matched those of conventional pathologic analysis.
In addition, D3 analysis of fine-needle lymph node biopsy samples was accurately able to differentiate 4 patients whose lymphoma diagnosis was confirmed by conventional pathology from another 4 patients with benign lymph node enlargement.
Along with protein analyses, the D3 system was enhanced to successfully detect DNA—in this instance, from human papilloma virus—with great sensitivity.
In all of these tests, results were available in under an hour and at a cost of $1.80 per assay, a price that would be expected to drop with further refinement of the D3 system.
“We expect that the D3 platform will enhance the breadth and depth of cancer screening in a way that is feasible and sustainable for resource limited-settings,” said Ralph Weissleder, MD, PhD, also of Massachusetts General Hospital.
“By taking advantage of the increased penetration of mobile phone technology worldwide, the system should allow the prompt triaging of suspicious or high-risk cases that could help to offset delays caused by limited pathology services in those regions and reduce the need for patients to return for follow-up care, which is often challenging for them.”
The researchers’ next steps are to investigate D3’s ability to analyze protein and DNA markers of other disease catalysts, integrate the software with larger databases, and conduct clinical studies in settings such as care-delivery sites in developing countries or rural areas.
Massachusetts General Hospital has filed a patent application covering the D3 technology.

Photo by Daniel Sone
Scientists say a smartphone-based system could bring rapid, accurate molecular diagnosis of cancers and other diseases to locations lacking the latest medical technology.
In PNAS, the group explained how the digital diffraction diagnosis (D3) system collects detailed microscopic images for digital analysis of the molecular composition of cells and tissues.
In pilot experiments, the system enabled accurate diagnoses of lymphoma and cervical cancer.
“The emerging genomic and biological data for various cancers, which can be essential to choosing the most appropriate therapy, supports the need for molecular profiling strategies that are more accessible to providers, clinical investigators, and patients,” said study author Cesar Castro, MD, of Massachusetts General Hospital in Boston.
“And we believe the platform we have developed provides essential features at an extraordinarily low cost.”
The D3 system features an imaging module with a battery-powered LED light clipped onto a standard smartphone that records high-resolution imaging data with its camera.
With a greater field of view than traditional microscopy, the D3 system is capable of recording data on more than 100,000 cells from a blood or tissue sample in a single image. The data can then be transmitted for analysis to a remote graphic-processing server via a secure, encrypted cloud service, and the results returned to the point of care.
For molecular analysis of tumors, a sample of blood or tissue is labeled with microbeads that bind to known cancer-related molecules and loaded into the D3 imaging module.
After the image is recorded and data transmitted to the server, the presence of specific molecules is detected by analyzing the diffraction patterns generated by the microbeads. The use of variously sized or coated beads may offer unique diffraction signatures to facilitate detection.
A numerical algorithm the researchers developed can distinguish cells from beads and analyze as much as 10 MB of data in less than nine hundredths of a second.
In a pilot test with cancer cell lines, the D3 system detected the presence of tumor proteins with an accuracy matching that of the current gold standard for molecular profiling. And the system’s larger field of view enabled simultaneous analysis of more than 100,000 cells at a time.
The researchers also conducted analyses of cervical biopsy samples from 25 women with abnormal PAP smears—samples collected along with those used for clinical diagnosis—using microbeads tagged with antibodies against 3 published markers of cervical cancer.
Based on the number of antibody-tagged microbeads binding to cells, D3 analysis promptly and reliably categorized biopsy samples as high-risk, low-risk, or benign. Results matched those of conventional pathologic analysis.
In addition, D3 analysis of fine-needle lymph node biopsy samples was accurately able to differentiate 4 patients whose lymphoma diagnosis was confirmed by conventional pathology from another 4 patients with benign lymph node enlargement.
Along with protein analyses, the D3 system was enhanced to successfully detect DNA—in this instance, from human papilloma virus—with great sensitivity.
In all of these tests, results were available in under an hour and at a cost of $1.80 per assay, a price that would be expected to drop with further refinement of the D3 system.
“We expect that the D3 platform will enhance the breadth and depth of cancer screening in a way that is feasible and sustainable for resource limited-settings,” said Ralph Weissleder, MD, PhD, also of Massachusetts General Hospital.
“By taking advantage of the increased penetration of mobile phone technology worldwide, the system should allow the prompt triaging of suspicious or high-risk cases that could help to offset delays caused by limited pathology services in those regions and reduce the need for patients to return for follow-up care, which is often challenging for them.”
The researchers’ next steps are to investigate D3’s ability to analyze protein and DNA markers of other disease catalysts, integrate the software with larger databases, and conduct clinical studies in settings such as care-delivery sites in developing countries or rural areas.
Massachusetts General Hospital has filed a patent application covering the D3 technology.
Gene appears key to HSC regulation

in the bone marrow
The gene Ash1l plays a key role in regulating the maintenance and self-renewal of hematopoietic stem cells (HSCs), according to a study published in The Journal of Clinical Investigation.
The research provides new insight into how the body creates and maintains a healthy blood supply and immune system. It also opens new lines of inquiry about Ash1l’s potential role in cancers—like leukemia—that involve other members of the same gene family.
“If we find that Ash1l plays a role [in leukemia], that would open up avenues to try to block or slow down its activity pharmacologically,” said study author Ivan Maillard, MD, of the University of Michigan Medical School in Ann Arbor.
The Ash1l gene regulates the expression of multiple downstream homeotic genes, which help ensure the correct anatomical structure of a developing organism. And Ash1l is part of a family of genes that includes MLL1.
The researchers found that both Ash1l and MLL1 contribute to blood renewal. They observed mild defects in mice missing one gene or the other, but lacking both genes led to catastrophic deficiencies.
“We now have clear evidence that these genes cooperate to develop a healthy blood system,” Dr Maillard said.
He and his colleagues also found that Ash1l-deficient mice had normal numbers of HSCs during early development but a lack of HSCs in maturity—an indication the cells were not able to properly maintain themselves in the bone marrow.
Ash1l-deficient HSCs were unable to establish normal blood renewal after an HSC transplant. Moreover, Ash1l-deficient stem cells competed poorly with normal HSCs in the bone marrow and could easily be dislodged.
“By continuing to investigate the basic, underlying mechanisms [of blood renewal], we are helping to untangle the complex machinery . . . that may lay the foundation for new human treatments 5, 10, or 20 years from now,” Dr Maillard said.

in the bone marrow
The gene Ash1l plays a key role in regulating the maintenance and self-renewal of hematopoietic stem cells (HSCs), according to a study published in The Journal of Clinical Investigation.
The research provides new insight into how the body creates and maintains a healthy blood supply and immune system. It also opens new lines of inquiry about Ash1l’s potential role in cancers—like leukemia—that involve other members of the same gene family.
“If we find that Ash1l plays a role [in leukemia], that would open up avenues to try to block or slow down its activity pharmacologically,” said study author Ivan Maillard, MD, of the University of Michigan Medical School in Ann Arbor.
The Ash1l gene regulates the expression of multiple downstream homeotic genes, which help ensure the correct anatomical structure of a developing organism. And Ash1l is part of a family of genes that includes MLL1.
The researchers found that both Ash1l and MLL1 contribute to blood renewal. They observed mild defects in mice missing one gene or the other, but lacking both genes led to catastrophic deficiencies.
“We now have clear evidence that these genes cooperate to develop a healthy blood system,” Dr Maillard said.
He and his colleagues also found that Ash1l-deficient mice had normal numbers of HSCs during early development but a lack of HSCs in maturity—an indication the cells were not able to properly maintain themselves in the bone marrow.
Ash1l-deficient HSCs were unable to establish normal blood renewal after an HSC transplant. Moreover, Ash1l-deficient stem cells competed poorly with normal HSCs in the bone marrow and could easily be dislodged.
“By continuing to investigate the basic, underlying mechanisms [of blood renewal], we are helping to untangle the complex machinery . . . that may lay the foundation for new human treatments 5, 10, or 20 years from now,” Dr Maillard said.

in the bone marrow
The gene Ash1l plays a key role in regulating the maintenance and self-renewal of hematopoietic stem cells (HSCs), according to a study published in The Journal of Clinical Investigation.
The research provides new insight into how the body creates and maintains a healthy blood supply and immune system. It also opens new lines of inquiry about Ash1l’s potential role in cancers—like leukemia—that involve other members of the same gene family.
“If we find that Ash1l plays a role [in leukemia], that would open up avenues to try to block or slow down its activity pharmacologically,” said study author Ivan Maillard, MD, of the University of Michigan Medical School in Ann Arbor.
The Ash1l gene regulates the expression of multiple downstream homeotic genes, which help ensure the correct anatomical structure of a developing organism. And Ash1l is part of a family of genes that includes MLL1.
The researchers found that both Ash1l and MLL1 contribute to blood renewal. They observed mild defects in mice missing one gene or the other, but lacking both genes led to catastrophic deficiencies.
“We now have clear evidence that these genes cooperate to develop a healthy blood system,” Dr Maillard said.
He and his colleagues also found that Ash1l-deficient mice had normal numbers of HSCs during early development but a lack of HSCs in maturity—an indication the cells were not able to properly maintain themselves in the bone marrow.
Ash1l-deficient HSCs were unable to establish normal blood renewal after an HSC transplant. Moreover, Ash1l-deficient stem cells competed poorly with normal HSCs in the bone marrow and could easily be dislodged.
“By continuing to investigate the basic, underlying mechanisms [of blood renewal], we are helping to untangle the complex machinery . . . that may lay the foundation for new human treatments 5, 10, or 20 years from now,” Dr Maillard said.
How telomere length affects cancer mortality
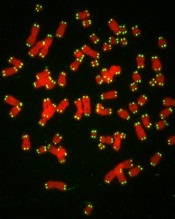
with telomeres in green
Image by Claus Azzalin
New research suggests telomere length is associated with cancer mortality—but not in the way researchers expected.
The study showed that short telomeres in peripheral blood leukocytes were associated with high mortality from all causes.
But genetically determined short telomeres were associated with low cancer mortality.
Line Rode, MD, PhD, of Herlev Hospital in Denmark, and colleagues reported these findings in JNCI: Journal of the National Cancer Institute.
Some previous studies have suggested an association between short telomeres and high mortality, including cancer mortality, while others have not. A possible explanation for the conflicting evidence may be that the association was correlational, but other factors that were not adjusted for (such as age and lifestyle) were the real causes.
Genetic variation in genes associated with telomere length (TERC, TERT, and OBFC1) is independent of age and lifestyle factors. So researchers speculated that a genetic analysis called a Mendelian randomization could eliminate some of the confounding and allow them to confirm the association between telomere length and cancer mortality.
To perform this analysis, the team used data from 2 prospective cohort studies. The Copenhagen City Heart Study and the Copenhagen General Population Study included 64,637 individuals who were followed from 1991 to 2011.
Participants completed a questionnaire, underwent a physical examination, and had blood drawn for biochemistry, genotyping, and telomere length assays.
For each subject, the researchers had information on physical characteristics such as body mass index (BMI), blood pressure, and cholesterol measurements, as well as smoking status, alcohol consumption, physical activity, and socioeconomic variables.
In addition to measuring telomere length for each subject, the researchers used 3 single nucleotide polymorphisms of TERC, TERT, and OBFC1 to construct a score for the presence of telomere-shortening alleles.
A total of 7607 individuals died during the study period, 2420 of cancer. Overall, decreasing telomere length was associated with age, variables such as BMI and smoking, and death from all causes, including cancer.
In contrast, a higher genetic score for telomere shortening was associated with decreased cancer mortality but not with any other causes of death.
The researchers said this suggests the slightly shorter telomeres in cancer patients with the higher genetic score for telomere shortening might be beneficial because uncontrolled cancer cell replication is reduced.
And long telomeres may confer a survival advantage for cancer cells, as they allow for multiple cell divisions that lead to high cancer mortality.

with telomeres in green
Image by Claus Azzalin
New research suggests telomere length is associated with cancer mortality—but not in the way researchers expected.
The study showed that short telomeres in peripheral blood leukocytes were associated with high mortality from all causes.
But genetically determined short telomeres were associated with low cancer mortality.
Line Rode, MD, PhD, of Herlev Hospital in Denmark, and colleagues reported these findings in JNCI: Journal of the National Cancer Institute.
Some previous studies have suggested an association between short telomeres and high mortality, including cancer mortality, while others have not. A possible explanation for the conflicting evidence may be that the association was correlational, but other factors that were not adjusted for (such as age and lifestyle) were the real causes.
Genetic variation in genes associated with telomere length (TERC, TERT, and OBFC1) is independent of age and lifestyle factors. So researchers speculated that a genetic analysis called a Mendelian randomization could eliminate some of the confounding and allow them to confirm the association between telomere length and cancer mortality.
To perform this analysis, the team used data from 2 prospective cohort studies. The Copenhagen City Heart Study and the Copenhagen General Population Study included 64,637 individuals who were followed from 1991 to 2011.
Participants completed a questionnaire, underwent a physical examination, and had blood drawn for biochemistry, genotyping, and telomere length assays.
For each subject, the researchers had information on physical characteristics such as body mass index (BMI), blood pressure, and cholesterol measurements, as well as smoking status, alcohol consumption, physical activity, and socioeconomic variables.
In addition to measuring telomere length for each subject, the researchers used 3 single nucleotide polymorphisms of TERC, TERT, and OBFC1 to construct a score for the presence of telomere-shortening alleles.
A total of 7607 individuals died during the study period, 2420 of cancer. Overall, decreasing telomere length was associated with age, variables such as BMI and smoking, and death from all causes, including cancer.
In contrast, a higher genetic score for telomere shortening was associated with decreased cancer mortality but not with any other causes of death.
The researchers said this suggests the slightly shorter telomeres in cancer patients with the higher genetic score for telomere shortening might be beneficial because uncontrolled cancer cell replication is reduced.
And long telomeres may confer a survival advantage for cancer cells, as they allow for multiple cell divisions that lead to high cancer mortality.

with telomeres in green
Image by Claus Azzalin
New research suggests telomere length is associated with cancer mortality—but not in the way researchers expected.
The study showed that short telomeres in peripheral blood leukocytes were associated with high mortality from all causes.
But genetically determined short telomeres were associated with low cancer mortality.
Line Rode, MD, PhD, of Herlev Hospital in Denmark, and colleagues reported these findings in JNCI: Journal of the National Cancer Institute.
Some previous studies have suggested an association between short telomeres and high mortality, including cancer mortality, while others have not. A possible explanation for the conflicting evidence may be that the association was correlational, but other factors that were not adjusted for (such as age and lifestyle) were the real causes.
Genetic variation in genes associated with telomere length (TERC, TERT, and OBFC1) is independent of age and lifestyle factors. So researchers speculated that a genetic analysis called a Mendelian randomization could eliminate some of the confounding and allow them to confirm the association between telomere length and cancer mortality.
To perform this analysis, the team used data from 2 prospective cohort studies. The Copenhagen City Heart Study and the Copenhagen General Population Study included 64,637 individuals who were followed from 1991 to 2011.
Participants completed a questionnaire, underwent a physical examination, and had blood drawn for biochemistry, genotyping, and telomere length assays.
For each subject, the researchers had information on physical characteristics such as body mass index (BMI), blood pressure, and cholesterol measurements, as well as smoking status, alcohol consumption, physical activity, and socioeconomic variables.
In addition to measuring telomere length for each subject, the researchers used 3 single nucleotide polymorphisms of TERC, TERT, and OBFC1 to construct a score for the presence of telomere-shortening alleles.
A total of 7607 individuals died during the study period, 2420 of cancer. Overall, decreasing telomere length was associated with age, variables such as BMI and smoking, and death from all causes, including cancer.
In contrast, a higher genetic score for telomere shortening was associated with decreased cancer mortality but not with any other causes of death.
The researchers said this suggests the slightly shorter telomeres in cancer patients with the higher genetic score for telomere shortening might be beneficial because uncontrolled cancer cell replication is reduced.
And long telomeres may confer a survival advantage for cancer cells, as they allow for multiple cell divisions that lead to high cancer mortality.